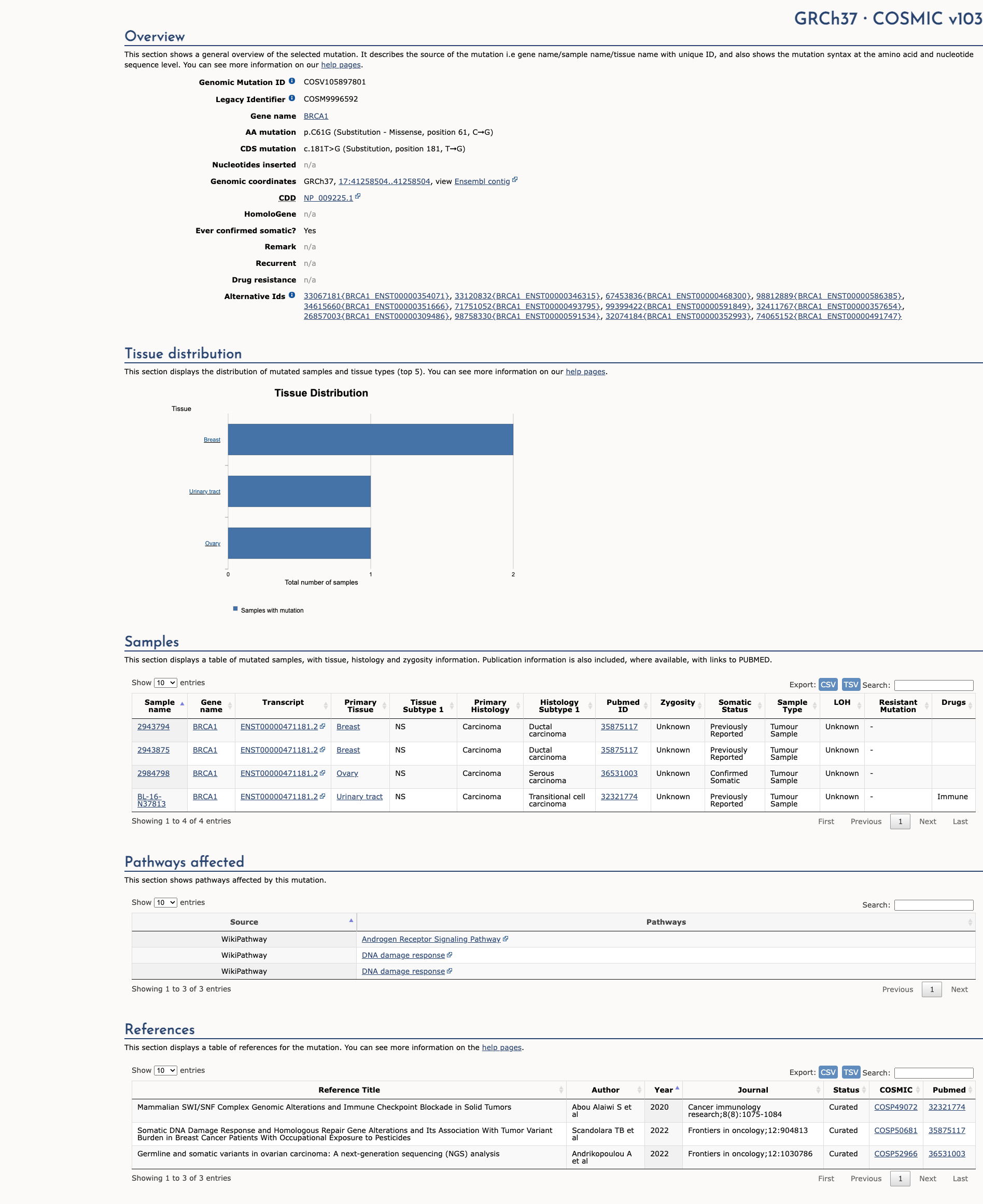
BRCA1 c.181T>G, p.Cys61Gly
NM_007294.3:c.181T>G
COSMIC ID: COSM9996592
Likely Pathogenic
The BRCA1 C61G variant is classified as Likely Pathogenic based on strong functional evidence (PS3), rarity in population databases (PM2), location in a critical functional domain (PM1), multiple computational deleterious predictions (PP3), and authoritative clinical reports (PP5).
ACMG/AMP Criteria Applied
PS3
PM1
PM2
PP3
PP5
Genetic Information
Gene & Transcript Details
Gene
BRCA1
Transcript
NM_007294.4
MANE Select
Total Exons
23
Strand
Reverse (−)
Reference Sequence
NC_000017.10
Alternative Transcripts
| ID | Status | Details |
|---|---|---|
| NM_007294.2 | Alternative | 23 exons | Reverse |
| NM_007294.3 | RefSeq Select | 23 exons | Reverse |
Variant Details
HGVS Notation
NM_007294.3:c.181T>G
Protein Change
C61G
Location
Exon 4
(Exon 4 of 23)
5'Exon Structure (23 total)3'
Functional Consequence
Loss of Function
Related Variants
No evidence of other pathogenic variants at position 61 in gene BRCA1
Alternate Identifiers
COSM9996592
Variant interpretation based on transcript NM_007294.4
Genome Browser
Loading genome browser...
HGVS InputNM_007294:c.181T>G
Active Tracks
ConservationRefSeqClinVargnomAD
Navigation tips: Use mouse to drag and zoom. Click on features for details.
Clinical Data
Global Frequency
0.00319%
Rare
Highest in Population
European (non-Finnish)
0.00617%
Rare
Global: 0.00319%
European (non-Finnish): 0.00617%
0%
0.05%
0.1%
1%
5%
10%+
Allele Information
Total: 250754Alt: 8Homozygotes: 0
ACMG Criteria Applied
PM2
This variant is present in gnomAD (MAF= 0.00319%, 8/250754 alleles, homozygotes = 0) and at a higher frequency in the European (non-Finnish) population (MAF= 0.00617%, 7/113480 alleles, homozygotes = 0). The variant is rare (MAF < 0.1%), supporting PM2 criterion application.
Classification
18 publications
Likely Pathogenic
Based on 55 submitter reviews in ClinVar
Submitter Breakdown
54 Path
1 LP
Pathogenic
Likely Path.
VUS
Likely Benign
Benign
Publications (18)
IARC class based on posterior probability from multifactorial likelihood analysis, thresholds for class as per Plon et al. 2008 (PMID: 18951446). Class 5 based on posterior probability = 1
Variant summary: The BRCA1 c.181T>G (p.Cys61Gly) variant involves the alteration of a conserved nucleotide that leads to the alteration of an amino acid residue in the RING domain of BRCA1 protein (InterPro). 4/5 in silico tools predict a damaging outcome for this variant and several functional studies demonstrated the Cys61Gly mutation affects several functional properties of the BRCA1 NH2-terminal domain (e.g. Brzovic 1998, 2003). This variant was found in 8/119006 control chromosomes, predominantly observed in the European (Non-Finnish) subpopulation at a frequency of 0.000122 (8/65364). This frequency is smaller than the estimated maximal expected allele frequency of a pathogenic BRCA1 variant (0.0010005). This is a well established disease variant that was reported in multiple patients with HBOC (e.g. Friedman 1994, Gorski 2000, Thorstenson 2003, Bogdanova 2010). In addition, multiple clinical diagnostic laboratories/reputable databases classified this variant as pathogenic. Taken together, this variant is classified as pathogenic.
The p.Cys61Gly variant in BRCA1 has been identified in >300 individuals with BRCA1-associated cancers and has segregated with disease in multiple families, including one male with breast cancer (Friedman 1994, Gorski 1999, Cherbal 2010, Bogdanova 2010, Breast Cancer Information Core database, www.research.nhgri.nih.gov/bic/). This variant has been identified in 8/65364 European chromosomes by the Exome Aggregation Consortium (ExAC, http://exac.broadinstitute.org; dbSNP rs28897672). Please note this frequency is low enough to be consistent with the frequency of hereditary breast and ovarian cancer (HBOC) in the general population. In vitro functional studies have shown that the p.Cys61Gly variant disrupts protein function and produces drug-resistant tumors in mouse models (Brzvoic 1998 and Drost 2011). In summary, this variant meets our criteria to be classified as pathogenic for HBOC in an autosomal dominant manner based upon segregation studies, presence in affected individuals, low frequency in controls, and functional evidence. The ACMG/AMP criteria applied: PS4, PS3_M, PP1_M, PM2_P.
The c.181T>G (p.Cys61Gly) variant of BRCA1 has been detected in multiple individuals with breast and ovarian cancer (PMID: 7894493, 10447273, 10788334, 11102977, 19594371, 20180014, 20345474, 20507347, 20569256, 20180014, 21324516, 21503673, 21965345, 23695190, 24728189, 29492181) and prostate cancer (PMID: 27433846). Functional in vitro assays showed that this variant increases proteolytic susceptibility of the COOH-terminal portion of the NH2-terminal domain and perturbs the oligomerization properties of BRCA1 (PMID: 9525870), fails to reverse radiation hypersensitivity (PMID: 11320250), and damages ubiquitin ligase activity (PMID: 11278247). This variant has been reported in 7 non-Finnish Europeans from gnomAD. This variant occurs at high frequency in Eastern European countries and is considered a founder mutation in these countries (PMID: 20507347, 20345474). Cysteine at amino acid position 61 of the BRCA1 protein is highly conserved in mammals. While not validated for clinical use, computer-based algorithms predict this p.Cys61Gly change to be deleterious. Other changes affecting the same amino acid have been reported in individuals with breast and/or ovarian cancer (p.Cys61Arg, p.Cys61Ser, p.Cys61Tyr). This variant is classified as pathogenic.
Clinical Statement
This variant has been reported in ClinVar as not provided (1 clinical laboratories) and as Pathogenic (54 clinical laboratories) and as Likely pathogenic (1 clinical laboratories) and as pathogenic (2 clinical laboratories) and as Pathogenic by Evidence-based Network for the Interpretation of Germline Mutant Alleles (ENIGMA) expert panel.
Expert Panel Reviews
Pathogenic
Evidence-based Network for the Interpretation of Germline Mutant Alleles (ENIGMA)
Functional Impact
Functional Domain
Hotspot Status
Not a hotspot
Domain Summary
This variant is not located in a mutational hotspot or critical domain (0 mutations).
Related Variants in This Domain
No evidence of other pathogenic variants at position 61 in gene BRCA1
Functional Summary
The BRCA1 C61G variant has been functionally characterized and shown to have a damaging effect on BRCA1 function. It disrupts homologous recombination and single-strand annealing pathways of DNA repair, impairs binding to the E2 ubiquitin-conjugating enzyme UbcH5a, and results in increased radiosensitivity, chromosomal instability, and centrosome abnormalities in vitro. These findings indicate that the BRCA1 C61G variant is deleterious to BRCA1's tumor suppressor function.
Database Previews
OncoKB
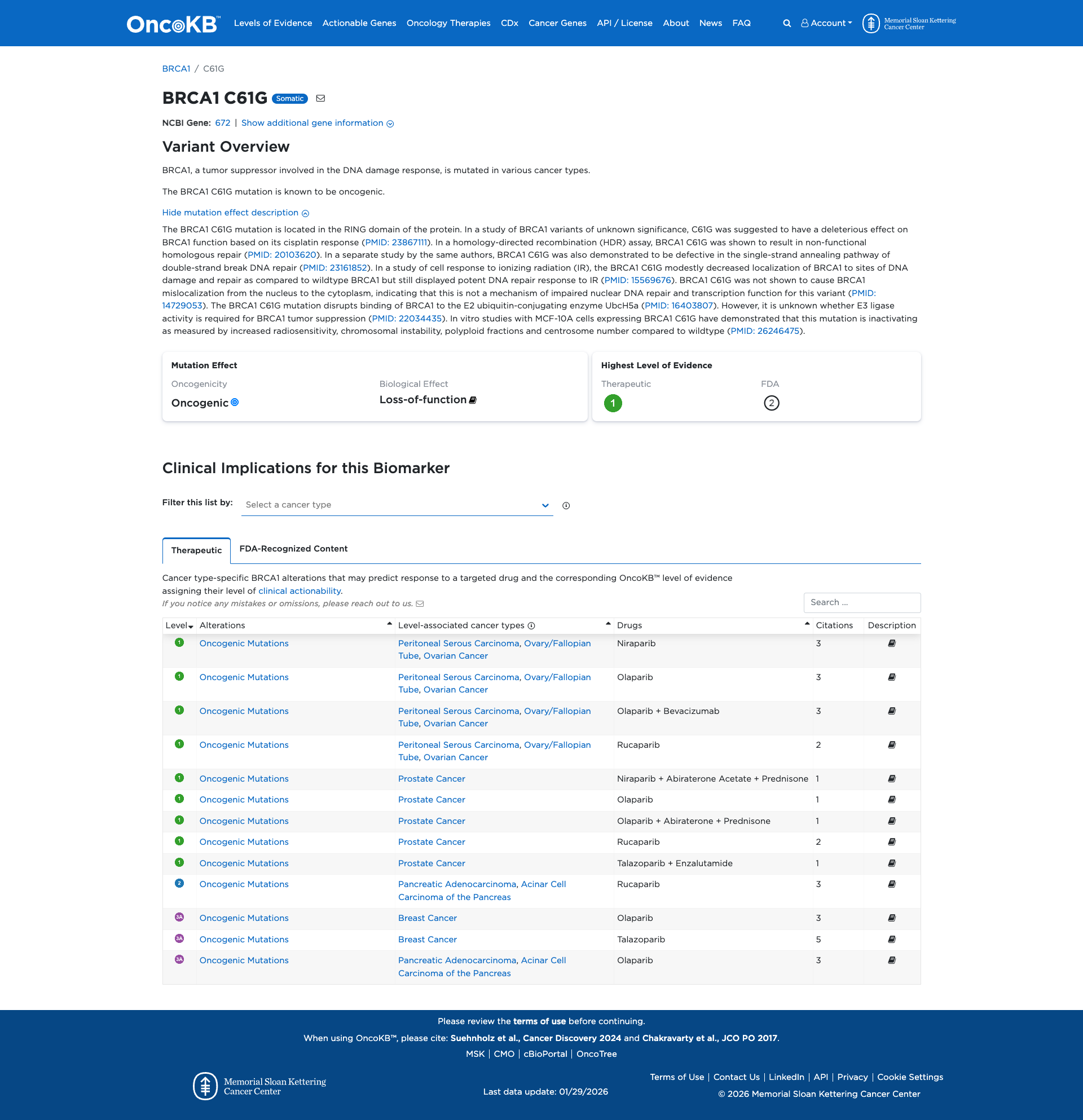
JAX-CKB
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
REVEL Score
0.948
0.948
Likely Benign0.0
Uncertain (Low)0.2
Uncertain (Med)0.5
Likely Pathogenic0.75
REVEL scores ≥ 0.75 are strong evidence (PP3)
Predictor Consensus
Mixed/VUS
PP3 Applied
Yes
Additional Predictors
Benign:
CADD: 4.93
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific) VCEP Guidelines
PVS1
PVS1 (Not Applied) Strength Modified
According to VCEP guidelines: PVS1 applies to null variants in BRCA1. This is a missense variant (C61G). Therefore, PVS1 is not applied.
PS1
PS1 (Not Applied) Strength Modified
According to standard ACMG guidelines: PS1 applies when the same amino acid change as a known pathogenic variant occurs via a different nucleotide change. There is no other nucleotide change at p.C61 leading to the same amino acid. Therefore, PS1 is not applied.
PS2
PS2 (Not Applied) Strength Modified
According to standard ACMG guidelines: PS2 applies for confirmed de novo occurrence with parental confirmation. No parental or de novo data are available. Therefore, PS2 is not applied.
PS3
PS3 (Strong)
According to VCEP guidelines: "PS3 Strong: Well-established in vitro or in vivo functional studies supportive of a damaging effect." The evidence for this variant shows disruption of homologous recombination and single-strand annealing, impaired UbcH5a binding, increased radiosensitivity and chromosomal instability. Therefore, PS3 is applied at Strong strength.
PS4
PS4 (Not Applied) Strength Modified
According to VCEP guidelines: PS4 applies when variant prevalence in affected individuals shows OR ≥4, p≤0.05. No case-control or cohort data are available. Therefore, PS4 is not applied.
PM1
PM1 (Moderate)
According to standard ACMG guidelines: "PM1 Moderate: Located in a mutational hotspot or well-established functional domain without benign variation." The residue C61 lies within the BRCA1 RING domain (aa2-101), a known functional domain enriched for pathogenic missense variants. Therefore, PM1 is applied at Moderate strength.
PM2
PM2 (Supporting) Strength Modified
According to VCEP guidelines: "PM2 Supporting: Absent from controls in gnomAD." The variant has a MAF of 0.00319% in gnomAD with no homozygotes. Therefore, PM2 is applied at Supporting strength.
PM3
PM3 (Not Applied) Strength Modified
According to VCEP guidelines: PM3 applies for co-occurrence in trans with another pathogenic variant in a recessive phenotype. No such data are available. Therefore, PM3 is not applied.
PM4
PM4 (Not Applied) Strength Modified
According to standard ACMG guidelines: PM4 applies to protein length changes (in-frame indels) not expected. This is a missense substitution. Therefore, PM4 is not applied.
PM5
PM5 (Not Applied) Strength Modified
According to standard ACMG guidelines: PM5 applies to novel missense changes at residues where other missense pathogenic variants occur. There is no evidence of another pathogenic missense at C61. Therefore, PM5 is not applied.
PM6
PM6 (Not Applied) Strength Modified
According to standard ACMG guidelines: PM6 applies to assumed de novo without confirmation. No de novo evidence. Therefore, PM6 is not applied.
PP1
PP1 (Not Applied) Strength Modified
According to VCEP guidelines: PP1 applies for segregation in affected family members. No segregation data are available. Therefore, PP1 is not applied.
PP2
PP2 (Not Applied) Strength Modified
According to standard ACMG guidelines: PP2 applies when a gene has low rates of benign missense variation and missense is a common mechanism. BRCA1 has many benign and pathogenic missense variants, so PP2 is not applied.
PP3
PP3 (Supporting)
According to VCEP guidelines: "PP3 Supporting: Missense variant inside a clinically important functional domain with computational evidence of deleterious effect." The variant is in the RING domain and has a REVEL score of 0.95. Therefore, PP3 is applied at Supporting strength.
PP4
PP4 (Not Applied) Strength Modified
According to VCEP guidelines: PP4 applies for highly specific phenotype in a single gene. No clinical phenotype data are provided. Therefore, PP4 is not applied.
PP5
PP5 (Supporting)
According to standard ACMG guidelines: "PP5 Supporting: Reputable source reports variant as pathogenic." The variant is reported as Pathogenic by ENIGMA and multiple ClinVar submitters. Therefore, PP5 is applied at Supporting strength.
BA1
BA1 (Not Applied) Strength Modified
According to VCEP guidelines: BA1 applies if allele frequency >0.1% in gnomAD. The allele frequency is 0.00319%. Therefore, BA1 is not applied.
BS1
BS1 (Not Applied) Strength Modified
According to VCEP guidelines: BS1 applies if allele frequency >0.01%. The allele frequency is below this. Therefore, BS1 is not applied.
BS2
BS2 (Not Applied) Strength Modified
According to VCEP guidelines: BS2 applies for absence of recessive phenotype (Fanconi Anemia). No data on FA phenotype. Therefore, BS2 is not applied.
BS3
BS3 (Not Applied) Strength Modified
According to VCEP guidelines: BS3 applies when well-established functional studies show no damaging effect. Functional studies show damaging effect here. Therefore, BS3 is not applied.
BS4
BS4 (Not Applied) Strength Modified
According to VCEP guidelines: BS4 applies for lack of segregation in multiple affected family members. No segregation data. Therefore, BS4 is not applied.
BP1
BP1 (Not Applied) Strength Modified
According to VCEP guidelines: BP1 applies to missense outside clinically important domains. This variant is within the RING domain. Therefore, BP1 is not applied.
BP2
BP2 (Not Applied) Strength Modified
According to standard ACMG guidelines: BP2 applies for observation in cis/trans with pathogenic variants. No such co-occurrence data. Therefore, BP2 is not applied.
BP3
BP3 (Not Applied) Strength Modified
According to standard ACMG guidelines: BP3 applies to in-frame indels in repetitive regions. This is a missense variant. Therefore, BP3 is not applied.
BP4
BP4 (Not Applied) Strength Modified
According to VCEP guidelines: BP4 applies when computational evidence supports benign impact. REVEL is 0.95, predicting deleterious. Therefore, BP4 is not applied.
BP5
BP5 (Not Applied) Strength Modified
According to standard ACMG guidelines: BP5 applies when variant co-occurs with another known pathogenic variant and no specific phenotype. No co-occurrence data. Therefore, BP5 is not applied.
BP6
BP6 (Not Applied) Strength Modified
According to standard ACMG guidelines: BP6 applies for reputable source reports benign. No such reports. Therefore, BP6 is not applied.
BP7
BP7 (Not Applied) Strength Modified
According to VCEP guidelines: BP7 applies to synonymous variants with no splicing impact. This is non-synonymous. Therefore, BP7 is not applied.