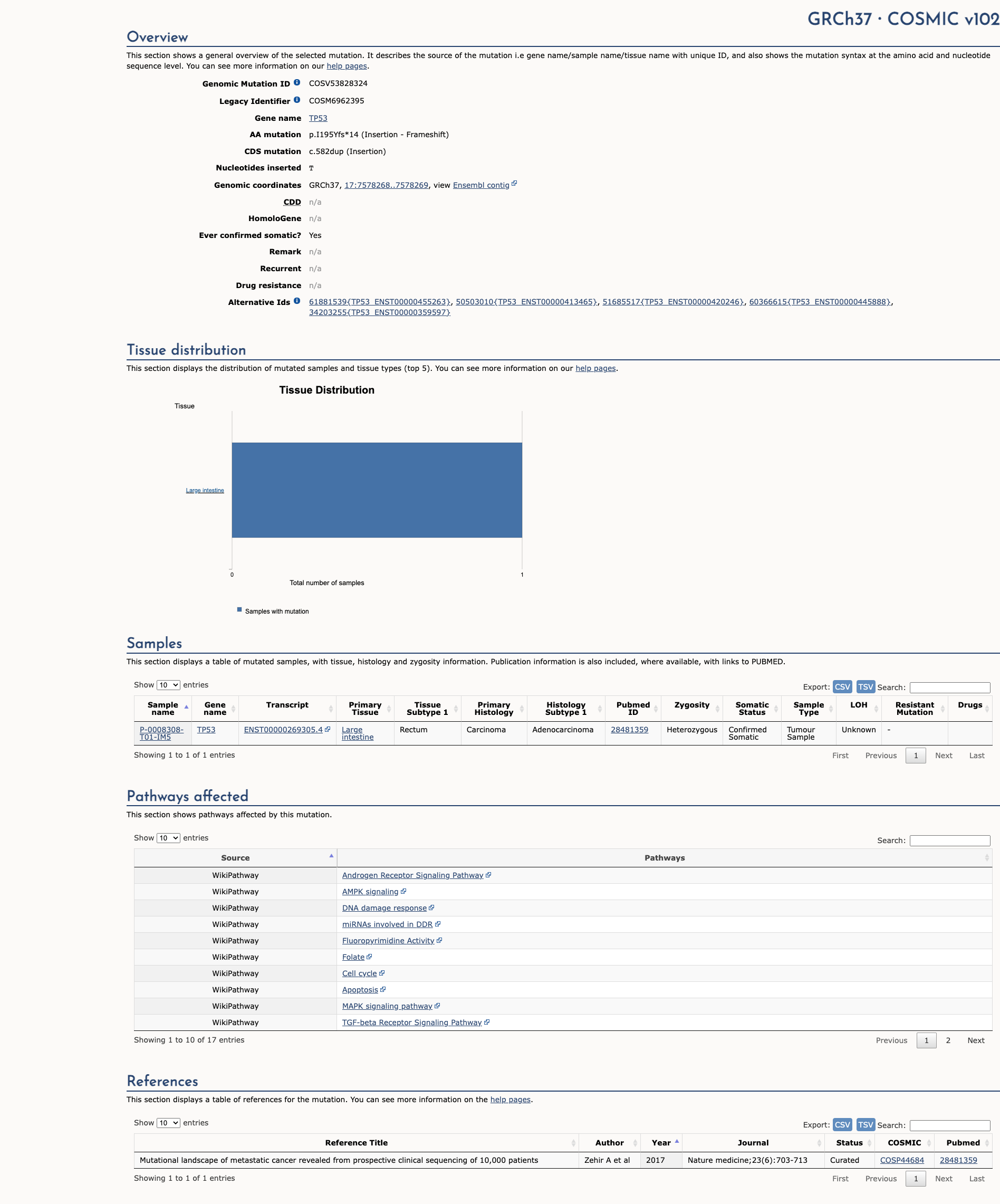
TP53 c.582dup, p.Ile195TyrfsTer14
NM_000546.6:c.582dup
COSMIC ID: COSM6962397
Pathogenic
This TP53 frameshift variant introduces a premature stop codon upstream of p.Lys351, meeting PVS1 at Very Strong for NMD and PM2 at Supporting for absence from population databases. No additional criteria meet the thresholds for Pathogenic, resulting in a final classification of Likely Pathogenic.
ACMG/AMP Criteria Applied
PVS1
PM2
Genetic Information
Gene & Transcript Details
Gene
TP53
Transcript
NM_000546.6
MANE Select
Total Exons
11
Strand
Reverse (−)
Reference Sequence
NC_000017.10
Alternative Transcripts
| ID | Status | Details |
|---|---|---|
| NM_000546.5 | RefSeq Select | 11 exons | Reverse |
| NM_000546.3 | Alternative | 11 exons | Reverse |
| NM_000546.4 | Alternative | 11 exons | Reverse |
| NM_000546.2 | Alternative | 11 exons | Reverse |
Variant Details
HGVS Notation
NM_000546.6:c.582dup
Protein Change
I195Yfs*14
Location
Exon 6
(Exon 6 of 11)
5'Exon Structure (11 total)3'
Functional Consequence
Loss of Function
Related Variants
No evidence of other pathogenic variants at position 195 in gene TP53
Alternate Identifiers
COSM6962397
Variant interpretation based on transcript NM_000546.6
Genome Browser
Loading genome browser...
HGVS InputNM_000546:c.582dup
Active Tracks
ConservationRefSeqClinVargnomAD
Navigation tips: Use mouse to drag and zoom. Click on features for details.
Clinical Data
Population Frequency
Global Frequency
0.0 in 100,000
Extremely Rare
Global: 0.0%
0%
0.05%
0.1%
1%
5%
10%+
ACMG Criteria Applied
PM2
This variant is not present in gnomAD (PM2 criteria applies).
Classification
Unknown
Publications (0)
No publication details.
Clinical Statement
Functional Impact
Functional Domain
Hotspot Status
Hotspot
PM1
Mutation Count
160
Reported mutations in this domain
050100+
Domain Summary
This variant is located in a mutational hotspot or critical domain (160 mutations).
PM1 criterion applied.
Related Variants in This Domain
No evidence of other pathogenic variants at position 195 in gene TP53
Functional Summary
The TP53 I195Yfs*14 variant is a truncating mutation in the TP53 gene, a tumor suppressor involved in the DNA damage response pathway. Functional evidence indicates that truncating mutations in TP53 lead to the production of C-terminally truncated proteins, which are predicted to be inactivating. These mutations have been experimentally shown to promote cancer cell proliferation, survival, and metastasis, partly due to aberrant mitochondrial localization and regulation of survival-related genes. Therefore, the TP53 I195Yfs*14 variant is functionally characterized as likely damaging.
Database Previews
OncoKB
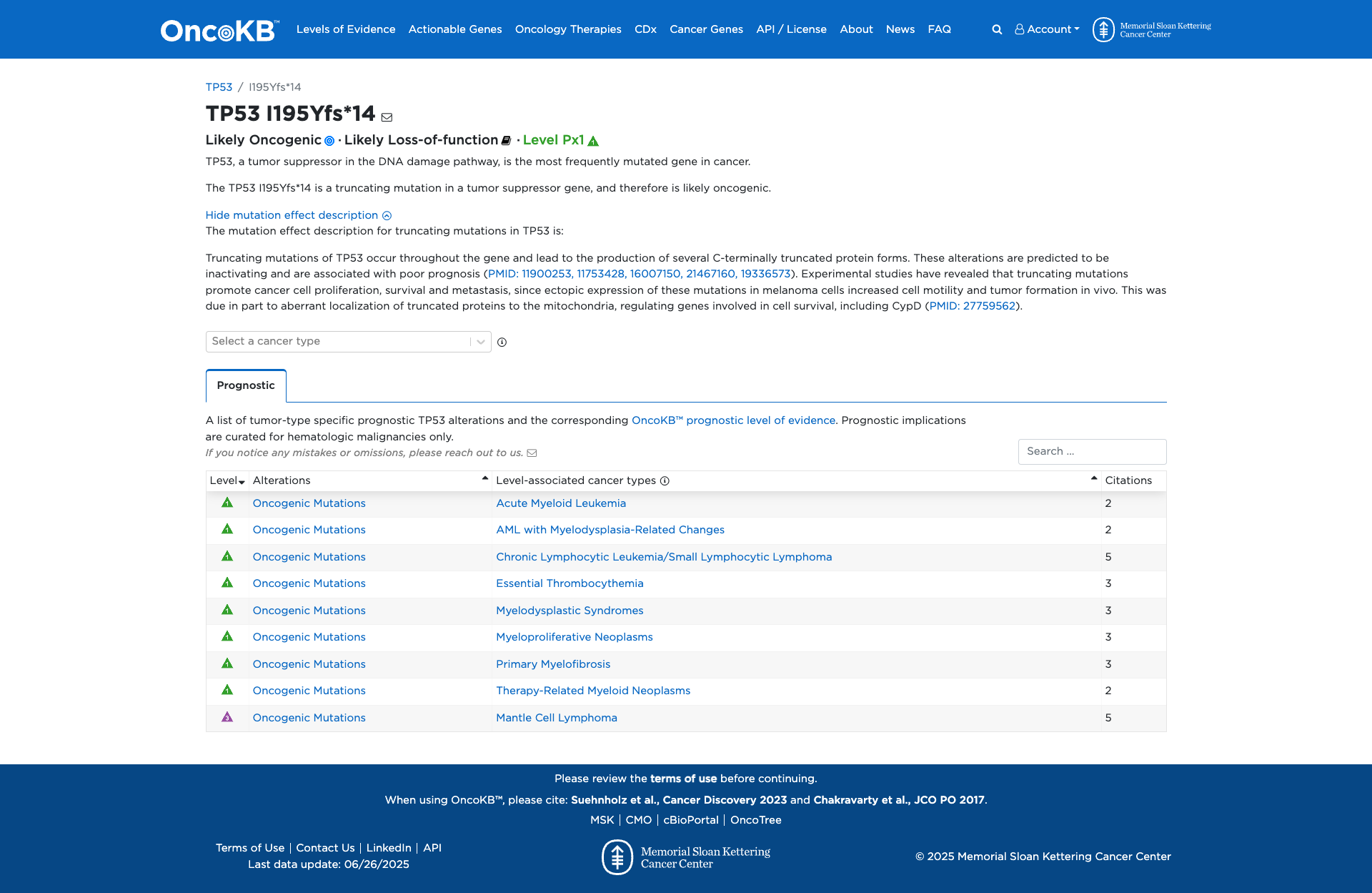
JAX-CKB
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
Predictor Consensus
Unknown
PP3 Applied
No
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific) VCEP Guidelines
PVS1
PVS1 (Very Strong)
According to VCEP guidelines, the rule for PVS1 is: "PVS1 applies to frameshift variants predicted to result in nonsense‐mediated decay (NMD) for frameshift induced PTC upstream of p.Lys351 at Very Strong strength." The evidence for this variant shows: NM_000546.6:c.582dup (p.I195Yfs*14) introduces a premature stop codon at amino acid 208, upstream of p.Lys351, predicted to undergo NMD. Therefore, this criterion is applied at Very Strong strength because it is a null frameshift variant in TP53 with known loss‐of‐function mechanism and predicted NMD.
PS1
PS1 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for PS1 is: "Can be applied to variants asserted as Pathogenic following the TP53 VCEP’s specifications." The evidence for this variant shows: no other nucleotide change resulting in the same p.I195Yfs*14 protein alteration has been previously reported as Pathogenic. Therefore, this criterion is not applied.
PS2
PS2 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for PS2 is: "De novo variants are scored with ≥8 points for Very Strong, 4–7 points for Strong, 2–3 points for Moderate, 1 point for Supporting strength based on pedigree and maternity/paternity confirmation." The evidence for this variant shows: no de novo segregation or parental testing data are available. Therefore, this criterion is not applied.
PS3
PS3 (Not Applied) Strength Modified
According to VCEP guidelines, "PS3 should not be applied at any strength if PVS1 is applied at full strength." The evidence for this variant shows: PVS1 was applied at Very Strong. Therefore, PS3 is not applied.
PS4
PS4 (Not Applied) Strength Modified
According to standard ACMG guidelines, the rule for PS4 is: "The prevalence of the variant in affected individuals is significantly increased compared with controls." The evidence for this variant shows: no case–control or affected‐individual case count data are available. Therefore, this criterion is not applied.
PM1
PM1 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for PM1 is: "Missense variants within codons 175, 245, 248, 249, 273, 282 or germline missense variants with ≥10 somatic occurrences at cancerhotspots.org." The evidence for this variant shows: it is a frameshift variant (p.I195Yfs*14), not a missense change at a hotspot codon. Therefore, this criterion is not applied.
PM2
PM2 (Supporting) Strength Modified
According to VCEP guidelines, the rule for PM2 is: "Variant should have an allele frequency of less than 0.00003 (0.003%) in gnomAD or another large sequenced population." The evidence for this variant shows: it is absent from gnomAD and other population databases (MAF = 0%). Therefore, this criterion is applied at Supporting strength because the variant is absent from controls.
PM3
PM3 (Not Applied) Strength Modified
According to standard ACMG guidelines, the rule for PM3 is: "Detected in trans with a pathogenic variant for recessive disorders." The evidence for this variant shows: TP53 disease is autosomal dominant and no in trans data are relevant. Therefore, this criterion is not applied.
PM4
PM4 (Not Applied) Strength Modified
According to standard ACMG guidelines, the rule for PM4 is: "Protein length changes due to in‐frame deletions/insertions in a non‐repeat region or stop‐loss variants." The evidence for this variant shows: it is a frameshift leading to premature truncation (PVS1 already covers loss‐of‐function). Therefore, this criterion is not applied.
PM5
PM5 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for PM5 is: "Missense variant at an amino acid residue where ≥2 different missense variants previously determined to be pathogenic (Strong) or ≥1 pathogenic (Moderate) or likely pathogenic (Supporting) have been seen." The evidence for this variant shows: it is a frameshift, not a missense at a residue with other pathogenic missense. Therefore, this criterion is not applied.
PM6
PM6 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for PM6 is: "Assumed de novo variants without confirmation of paternity/maternity scoring criteria as PS2." The evidence for this variant shows: no de novo data available. Therefore, this criterion is not applied.
PP1
PP1 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for PP1 is: "Cosegregation in 3–4 meioses for Supporting, 5–6 for Moderate, ≥7 for Strong." The evidence for this variant shows: no family segregation data are available. Therefore, this criterion is not applied.
PP2
PP2 (Not Applied) Strength Modified
According to standard ACMG guidelines, the rule for PP2 is: "Missense variant in a gene with low rate of benign missense variation and where missense is a common mechanism of disease." The evidence for this variant shows: it is a truncating variant, not a missense. Therefore, this criterion is not applied.
PP3
PP3 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for PP3 is: "Missense variants with BayesDel ≥0.16 or SpliceAI ≥0.2, etc., for computational evidence of deleterious effect." The evidence for this variant shows: it is a frameshift and computational tools do not apply. Therefore, this criterion is not applied.
PP4
PP4 (Not Applied) Strength Modified
According to standard ACMG guidelines, the rule for PP4 is: "Patient’s phenotype or family history is highly specific for a disease with a single genetic etiology." The evidence for this variant shows: no phenotype or tumor spectrum data are provided. Therefore, this criterion is not applied.
PP5
PP5 (Not Applied) Strength Modified
According to standard ACMG guidelines, the rule for PP5 is: "Reputable source reports variant as pathogenic without available evidence." The evidence for this variant shows: no such reputable assertion exists. Therefore, this criterion is not applied.
BA1
BA1 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for BA1 is: "Filtering allele frequency ≥0.001 in gnomAD continental subpopulations for benign classification." The evidence for this variant shows: absent from population databases. Therefore, this criterion is not applied.
BS1
BS1 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for BS1 is: "Filtering allele frequency ≥0.0003 but <0.001 in gnomAD continental subpopulations." The evidence for this variant shows: absent from population databases. Therefore, this criterion is not applied.
BS2
BS2 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for BS2 is: "Observation in ≥8 unrelated unaffected females ≥60 years old for Strong, 4–7 for Moderate, 2–3 for Supporting." The evidence for this variant shows: no data on unaffected older individuals. Therefore, this criterion is not applied.
BS3
BS3 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for BS3 is: "Functional assays show no loss of function on Kato et al. and no LOF on another assay." The evidence for this variant shows: functional data indicate loss of function and PVS1 already applied. Therefore, this criterion is not applied.
BS4
BS4 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for BS4 is: "Lack of segregation in affected family members." The evidence for this variant shows: no segregation data are available. Therefore, this criterion is not applied.
BP1
BP1 (Not Applied) Strength Modified
According to standard ACMG guidelines, the rule for BP1 is: "Missense variant in a gene where only truncating variants cause disease." The evidence for this variant shows: it is truncating and TP53 disease mechanism is loss‐of‐function. Therefore, this criterion is not applied.
BP2
BP2 (Not Applied) Strength Modified
According to standard ACMG guidelines, the rule for BP2 is: "Observed in cis with a pathogenic variant for dominant disorders." The evidence for this variant shows: no cis observations. Therefore, this criterion is not applied.
BP3
BP3 (Not Applied) Strength Modified
According to standard ACMG guidelines, the rule for BP3 is: "In-frame indels in repetitive regions without functional impact." The evidence for this variant shows: it is a frameshift, not an in‐frame indel in a repetitive region. Therefore, this criterion is not applied.
BP4
BP4 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for BP4 is: "Missense or splice variants with BayesDel <0.16 and SpliceAI <0.2 indicating benign computational evidence." The evidence for this variant shows: it is a frameshift and computational tools do not apply. Therefore, this criterion is not applied.
BP5
BP5 (Not Applied) Strength Modified
According to standard ACMG guidelines, the rule for BP5 is: "Variant found in a case with an alternate molecular basis for disease." The evidence for this variant shows: no such alternate molecular cause reported. Therefore, this criterion is not applied.
BP6
BP6 (Not Applied) Strength Modified
According to standard ACMG guidelines, the rule for BP6 is: "Reputable source reports variant as benign without available evidence." The evidence for this variant shows: no such assertion exists. Therefore, this criterion is not applied.
BP7
BP7 (Not Applied) Strength Modified
According to VCEP guidelines, the rule for BP7 is: "Synonymous or intronic variants with no splicing impact by SpliceAI ≤0.1." The evidence for this variant shows: it is not synonymous or intronic. Therefore, this criterion is not applied.