Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenVariant summary: TP53 c.215_216delinsGT (p.Pro72Arg) results in a non-conservative amino acid change in the encoded protein sequence. Algorithms developed to predict the effect of missense changes on protein structure and function are either unavailable or do not agree on the potential impact of this missense change. This variant was found in 3/120822 control chromosomes at a frequency of 0.0000248, which does not exceed the estimated maximal expected allele frequency of a pathogenic TP53 variant (0.0000398). However, the variant c.215C>G alone, which leads to the same Pro72Arg missense change, is found at a very high frequency in the population (79805/120924 control chromosomes; 27306 homozygotes), which strongly suggests that the Pro72Arg change does not affect protein function. ClinVar contains an entry for this variant (Variation ID: 237944). Based on the evidence outlined above, the variant was classified as likely benign.
"This variant has been reported in ClinVar as Likely benign (6 clinical laboratories) and as Benign (1 clinical laboratories)."
COSMIC Somatic Evidence
Open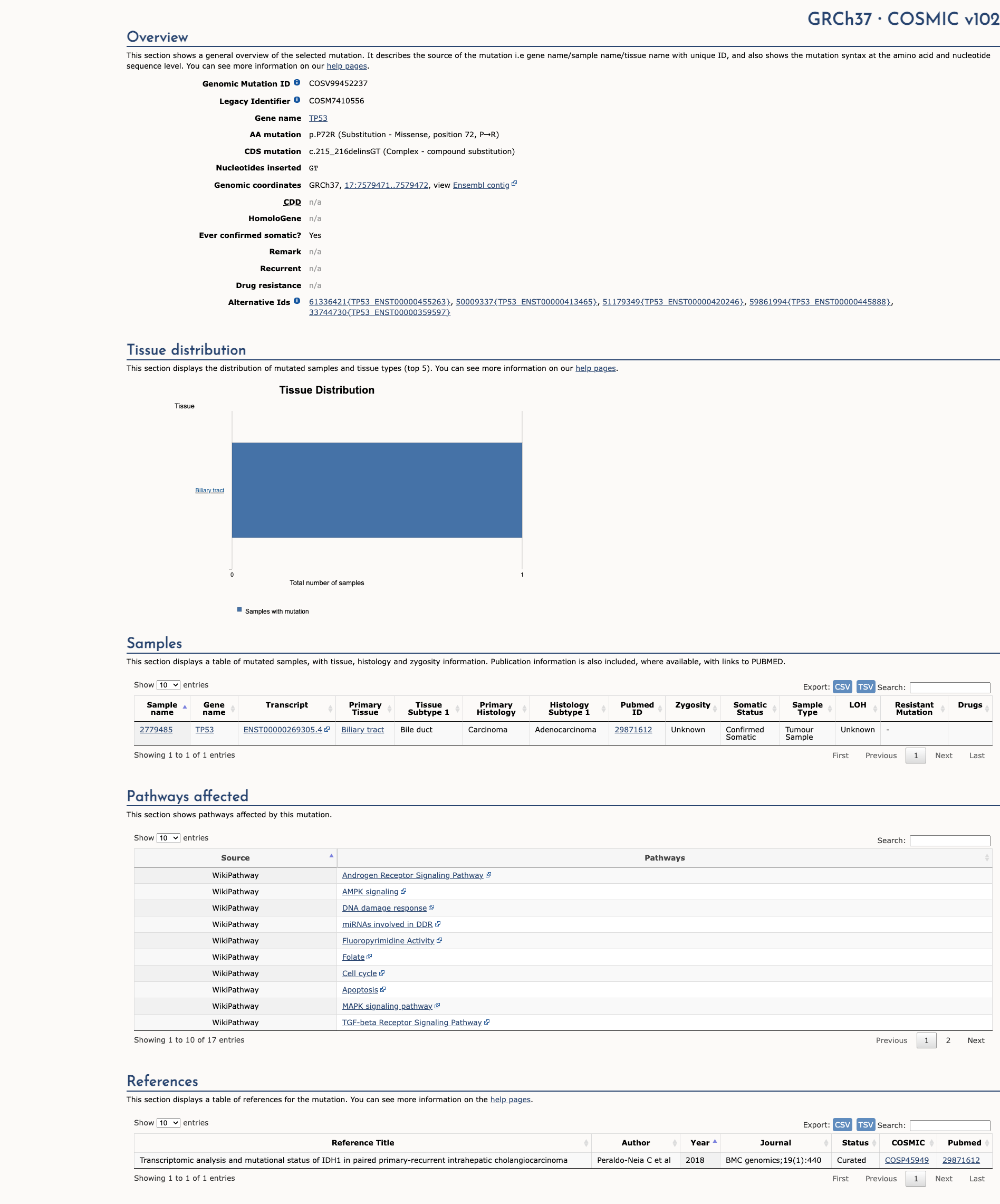
Functional Impact & Domains
Functional Domain
The TP53 P72R variant has been functionally studied and shown to decrease Pgc-1a binding, increase mitochondrial function, and enhance migration, invasion, and metastasis in the presence of mutant TP53 in cell culture and mouse models. However, it also transactivates downstream target genes and inhibits cell growth similarly to wild-type TP53 in culture. Thus, the overall effect of the P72R variant on TP53 protein function remains unclear.
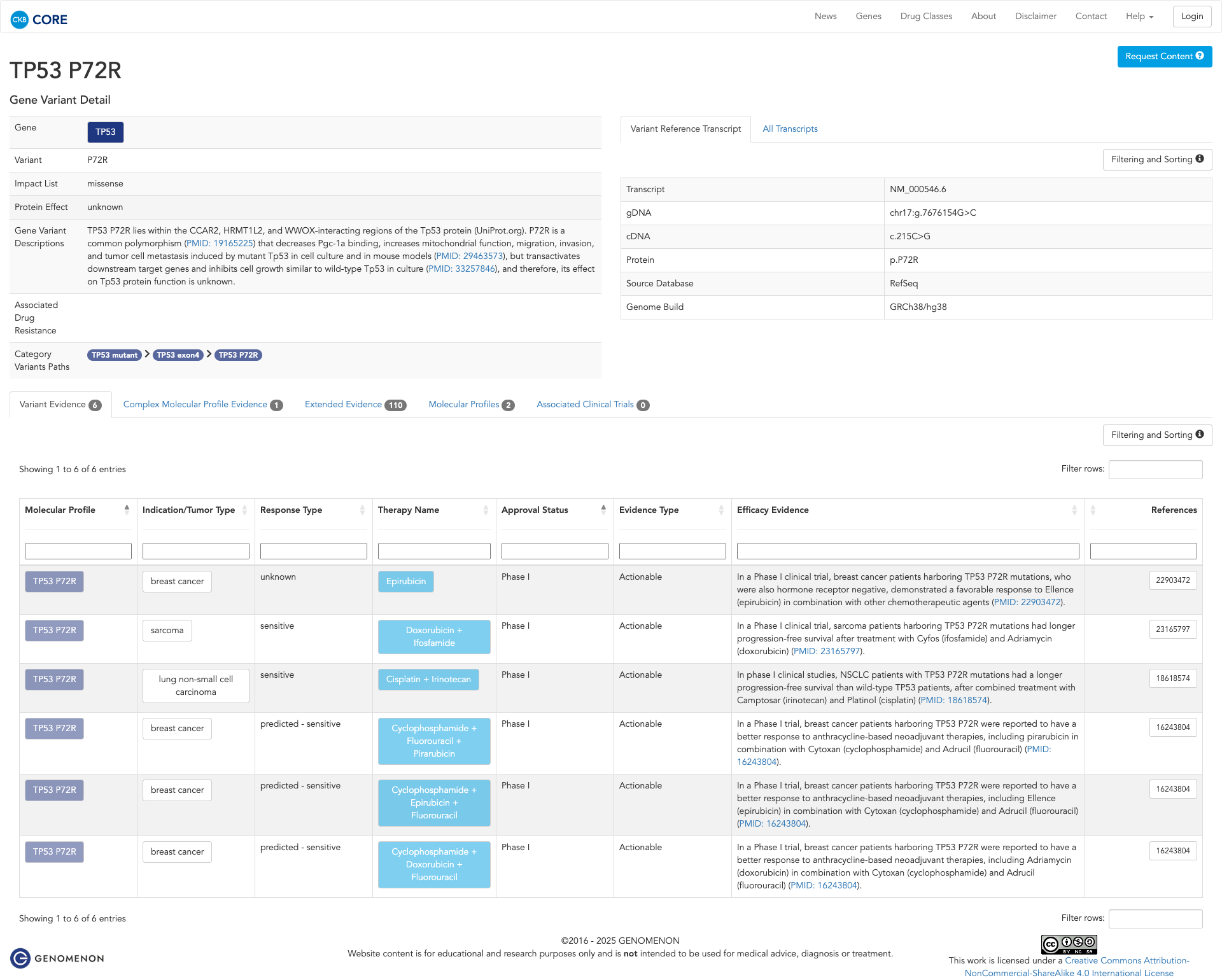
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -85 bp |
| Donor Loss (DL) | 0.04 | 41 bp |
| Acceptor Gain (AG) | 0.0 | 100 bp |
| Donor Gain (DG) | 0.0 | -159 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, PVS1 applies to null variants predicted to undergo NMD or disrupt critical splice sites. The evidence for this variant shows: it is a missense change (p.P72R), not a null or canonical splice variant. Therefore, this criterion is not applied.
PS1 (Not Applied)
According to VCEP guidelines, PS1 applies to a variant causing the same amino acid change as a known pathogenic variant. The evidence for this variant shows: no other pathogenic p.P72R variant reported. Therefore, this criterion is not applied.
PS2 (Not Applied)
According to standard ACMG guidelines, PS2 applies to confirmed de novo variants with parental confirmation. The evidence for this variant shows: no de novo data available. Therefore, this criterion is not applied.
PS3 (Not Applied)
According to VCEP guidelines, PS3 Strong requires non-functional Kato data AND LOF on another assay. The evidence for this variant shows: functional studies are conflicting with overall preserved TP53 transactivation. Therefore, this criterion is not applied.
PS4 (Not Applied)
According to VCEP guidelines, PS4 requires quantitative case–control or proband point scoring. The evidence for this variant shows: no case enrichment data available. Therefore, this criterion is not applied.
PM1 (Not Applied)
According to VCEP guidelines, PM1 Moderate applies to missense variants at TP53 hotspots codons 175, 245, 248, 249, 273, or 282. The evidence for this variant shows: it is at codon 72, outside these hotspots. Therefore, this criterion is not applied.
PM2 (Not Applied)
According to VCEP guidelines, PM2 Supporting applies when the allele frequency is <0.00003 in gnomAD. The evidence for this variant shows: allele frequency ∼0.3 in gnomAD. Therefore, this criterion is not applied.
PM3 (Not Applied)
According to standard ACMG guidelines, PM3 applies to variants observed in trans with a pathogenic variant in recessive disorders. The evidence for this variant shows: TP53 disease is dominant and no trans observations reported. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, PM4 applies to protein length changes via in‐frame indels. The evidence for this variant shows: it is a missense substitution (p.P72R), not an in‐frame indel. Therefore, this criterion is not applied.
PM5 (Not Applied)
According to VCEP guidelines, PM5 applies to missense variants at a residue with ≥2 pathogenic changes (Strong) or 1 pathogenic change (Moderate). The evidence for this variant shows: no other pathogenic missense at codon 72. Therefore, this criterion is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines, PM6 applies to assumed de novo cases without confirmation. The evidence for this variant shows: no de novo data available. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to VCEP guidelines, PP1 Supporting requires cosegregation in 3–4 meioses; Moderate requires 5–6; Strong requires ≥7. The evidence for this variant shows: no family segregation data. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, PP2 applies when missense variants in a gene with low benign variation and pathogenic missense are common. The evidence for this variant shows: TP53 has many tolerated missense variants. Therefore, this criterion is not applied.
PP3 (Not Applied)
According to standard ACMG guidelines, PP3 applies when multiple computational tools predict a deleterious effect. The evidence for this variant shows: computational tools predict benign. Therefore, this criterion is not applied.
PP4 (Not Applied)
According to VCEP guidelines, PP4 applies with phenotype specificity and variant observation. The evidence for this variant shows: no phenotype data. Therefore, this criterion is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, PP5 applies when a reputable source reports a pathogenic assertion. The evidence for this variant shows: no pathogenic assertions. Therefore, this criterion is not applied.
BA1 (Stand Alone)
According to VCEP guidelines, BA1 Stand Alone applies when the filtering allele frequency ≥0.001 in gnomAD. The evidence for this variant shows: allele frequency ∼0.3 in gnomAD, well above the 0.001 threshold. Therefore, this criterion is applied at Stand Alone strength because the variant is a common polymorphism inconsistent with pathogenicity.
BS1 (Not Applied)
According to VCEP guidelines, BS1 Strong applies when the filtering allele frequency is ≥0.0003 but <0.001. The evidence for this variant shows: allele frequency ∼0.3, which exceeds the upper limit. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to VCEP guidelines, BS2 Strong applies when ≥8 unrelated elderly females without cancer carry the variant. The evidence for this variant shows: no such data. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to VCEP guidelines, BS3 Strong applies when functional assays demonstrate no loss of function. The evidence for this variant shows: functional studies are conflicting with some evidence of altered binding. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to VCEP guidelines, BS4 applies to lack of segregation in affected family members. The evidence for this variant shows: no family data. Therefore, this criterion is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, BP1 applies when missense variants occur in a gene where only truncating variants cause disease. The evidence for this variant shows: TP53 pathogenic missense are common. Therefore, this criterion is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, BP2 applies when observed in trans with pathogenic variant for dominant disorders. The evidence for this variant shows: no such observations. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, BP3 applies to in‐frame deletions/insertions in repetitive regions. The evidence for this variant shows: it is a missense substitution. Therefore, this criterion is not applied.
BP4 (Supporting)
According to standard ACMG guidelines, BP4 applies when multiple computational tools predict no impact. The evidence for this variant shows: PolyPhen benign and SpliceAI 0.04 indicating no splicing impact. Therefore, this criterion is applied at Supporting strength because computational evidence supports benign effect.
BP5 (Not Applied)
According to standard ACMG guidelines, BP5 applies when a variant is found in trans with a pathogenic variant in a dominant disorder. The evidence for this variant shows: no such finding. Therefore, this criterion is not applied.
BP6 (Supporting)
According to standard ACMG guidelines, BP6 applies when a reputable source reports a benign assertion without available evidence. The evidence for this variant shows: ClinVar entries by multiple laboratories classify it as Likely benign/Benign. Therefore, this criterion is applied at Supporting strength.
BP7 (Not Applied)
According to VCEP guidelines, BP7 applies to synonymous or intronic variants outside splice motifs. The evidence for this variant shows: it is a missense change. Therefore, this criterion is not applied.