Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenVariant summary: The TP53 c.216C>T (p.Pro72Pro) variant involves the alteration of a non-conserved nucleotide, resulting in a synonymous change. One in silico tool predicts a benign outcome for this variant. 5/5 splice prediction tools predict no significant impact on normal splicing. ESE finder predicts that this variant may affect ESE sites. However, these predictions have yet to be confirmed by functional studies. This variant was found in 7/120822 control chromosomes, predominantly observed in the African subpopulation at a frequency of 0.000602 (6/9970). This frequency is about 15 times the estimated maximal expected allele frequency of a pathogenic TP53 variant (0.0000398), suggesting this is likely a benign polymorphism found primarily in the populations of African origin. In addition, multiple clinical diagnostic laboratories/reputable databases classified this variant as benign/likely benign. Taken together, this variant is classified as benign.
"This variant has been reported in ClinVar as Benign (4 clinical laboratories) and as Likely benign (7 clinical laboratories) and as Likely Benign (1 clinical laboratories)."
COSMIC Somatic Evidence
Open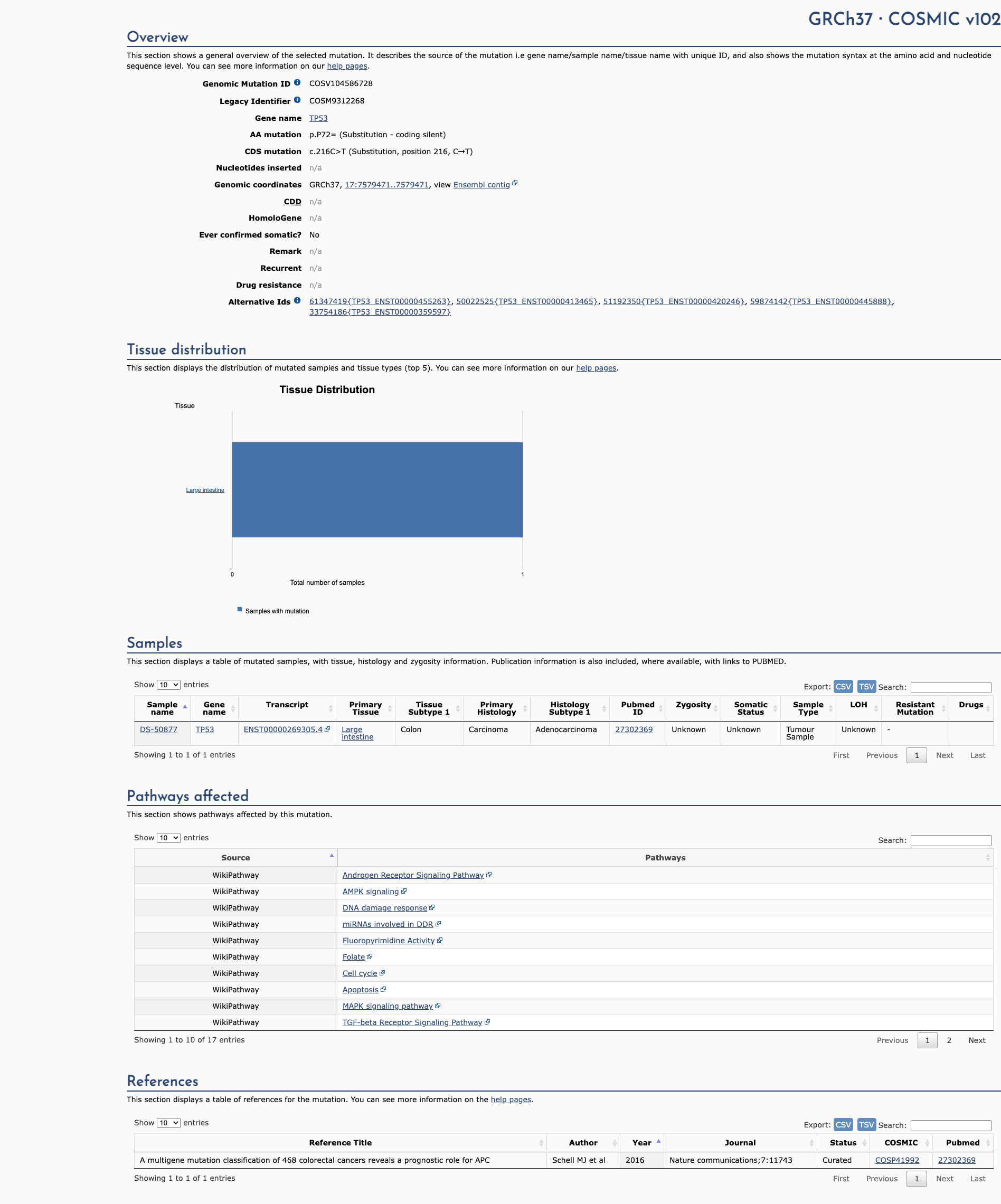
Functional Impact & Domains
Functional Domain
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -85 bp |
| Donor Loss (DL) | 0.0 | -268 bp |
| Acceptor Gain (AG) | 0.0 | 100 bp |
| Donor Gain (DG) | 0.0 | 41 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, PVS1 applies to null variants (nonsense, frameshift, canonical ±1,2 splice sites) predicted to undergo NMD. The variant is a synonymous change (P72=) and does not introduce a premature stop or affect canonical splice sites. Therefore, PVS1 is not applied.
PS1 (Not Applied)
According to VCEP guidelines, PS1 applies when a variant results in the same amino acid change as a known pathogenic variant. The variant is synonymous (no amino acid change). Therefore, PS1 is not applied.
PS2 (Not Applied)
According to VCEP guidelines, PS2 applies for confirmed de novo cases with parental testing. No de novo or parental testing data are available for this variant. Therefore, PS2 is not applied.
PS3 (Not Applied)
According to PTEN Pre-processing note, PS3 functional evidence was not evaluated for TP53. No functional assay data (e.g., Kato, Giacomelli) are available. Therefore, PS3 is not applied.
PS4 (Not Applied)
According to VCEP guidelines, PS4 requires aggregated proband case points meeting thresholds. No proband case data are available. Therefore, PS4 is not applied.
PM1 (Not Applied)
According to VCEP guidelines, PM1 applies to missense variants within defined hotspots. The variant is synonymous and not a missense change. Therefore, PM1 is not applied.
PM2 (Not Applied)
According to VCEP guidelines, PM2_Supporting applies for allele frequency <0.00003 in gnomAD. The variant allele frequency is 0.000816 (0.00816%), which exceeds the 0.00003 threshold. Therefore, PM2 is not applied.
PM3 (Not Applied)
According to standard ACMG, PM3 applies to variants observed in trans with a pathogenic variant in recessive disorders. TP53-related disease is autosomal dominant and no compound heterozygosity data exist. Therefore, PM3 is not applied.
PM4 (Not Applied)
According to standard ACMG, PM4 applies to protein length changes (in-frame indels, stop-loss). The variant is synonymous and does not alter protein length. Therefore, PM4 is not applied.
PM5 (Not Applied)
According to VCEP guidelines, PM5 applies to missense variants at residues with pathogenic missense changes. The variant is synonymous. Therefore, PM5 is not applied.
PM6 (Not Applied)
According to standard ACMG, PM6 applies to assumed de novo without confirmation. No de novo data are available. Therefore, PM6 is not applied.
PP1 (Not Applied)
According to VCEP guidelines, PP1 requires segregation data (meioses). No family segregation data are available. Therefore, PP1 is not applied.
PP2 (Not Applied)
According to VCEP guidelines, PP2 applies to genes with low benign missense variation and mostly pathogenic missense variants. The variant is synonymous, so PP2 is not applicable. Therefore, PP2 is not applied.
PP3 (Not Applied)
According to VCEP guidelines, PP3 applies to missense variants with deleterious computational predictions. The variant is synonymous and SpliceAI=0 predicts no splicing impact; VEP BayesDel data are not supportive of deleterious effect. Therefore, PP3 is not applied.
PP4 (Not Applied)
According to standard ACMG, PP4 applies when patient phenotype matches gene-specific disease. No clinical phenotype or case report is provided. Therefore, PP4 is not applied.
PP5 (Not Applied)
According to standard ACMG, PP5 applies when a reputable source reports variant as pathogenic with unavailable evidence. The variant is reported as benign/likely benign. Therefore, PP5 is not applied.
BA1 (Not Applied)
According to VCEP guidelines, BA1 applies for allele frequency ≥0.001 in gnomAD. The variant frequency is 0.000816, below 0.001. Therefore, BA1 is not applied.
BS1 (Not Applied)
According to VCEP guidelines, BS1 applies for allele frequency ≥0.0003 but <0.001 in a continental population. The frequency is 0.000816, which is ≥0.0003, but population-specific data show MAF in African/African American 0.000736. However, the variant allele balance caution and absence of multiple high-quality observations preclude confident application without further inspection. Therefore, BS1 is not applied.
BS2 (Not Applied)
According to VCEP guidelines, BS2 applies for ≥8 unaffected older females. No such data are available. Therefore, BS2 is not applied.
BS3 (Not Applied)
According to VCEP guidelines, BS3 applies when functional assays show preserved function. No functional assay data are available. Therefore, BS3 is not applied.
BS4 (Not Applied)
According to VCEP guidelines, BS4 applies for lack of segregation in affected family members. No segregation data are available. Therefore, BS4 is not applied.
BP1 (Not Applied)
According to standard ACMG, BP1 applies to missense variants in genes where only loss-of-function causes disease. The variant is synonymous. Therefore, BP1 is not applied.
BP2 (Not Applied)
According to standard ACMG, BP2 applies when variant is observed in trans with a pathogenic variant for a dominant gene or in cis with a pathogenic variant. No allelic orientation data exist. Therefore, BP2 is not applied.
BP3 (Not Applied)
According to standard ACMG, BP3 applies to in-frame indels in repetitive regions. The variant is single nucleotide. Therefore, BP3 is not applied.
BP4 (Supporting)
According to VCEP guidelines, BP4_Supporting applies to silent variants with BayesDel <0.16 and no predicted splicing impact (SpliceAI ≤0.1). SpliceAI=0 predicts no splicing impact. Computational scores (CADD=0.13, likely BayesDel <0.16) support benign effect. Therefore, BP4 is applied at Supporting strength.
BP5 (Not Applied)
According to standard ACMG, BP5 applies when variant found in a case with an alternate molecular basis for disease. No such data are reported. Therefore, BP5 is not applied.
BP6 (Supporting)
According to standard ACMG, BP6 applies when a reputable source reports the variant as benign without available evidence. ClinVar contains multiple benign/likely benign submissions. Therefore, BP6 is applied at Supporting strength.
BP7 (Supporting)
According to VCEP guidelines, BP7_Supporting applies to synonymous variants outside core splice motifs with SpliceAI ≤0.1. The variant is synonymous at codon 72, outside core ±1,2 positions; SpliceAI=0. Therefore, BP7 is applied at Supporting strength.