Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000314.8 | MANE Select | 8515 nt | 846–2057 |
| NM_000314.7 | RefSeq Select | 8514 nt | 845–2056 |
| NM_000314.5 | Alternative | 8719 nt | 1032–2243 |
| NM_000314.4 | Alternative | 5572 nt | 1032–2243 |
| NM_000314.3 | Alternative | 3416 nt | 1032–2243 |
| NM_000314.6 | Alternative | 8718 nt | 1032–2243 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Open"This variant has been reported in ClinVar as Likely pathogenic (1 clinical laboratories) and as Uncertain significance (1 clinical laboratories) and as Likely Pathogenic by Clingen PTEN Variant Curation Expert Panel, Clingen expert panel."
COSMIC Somatic Evidence
Open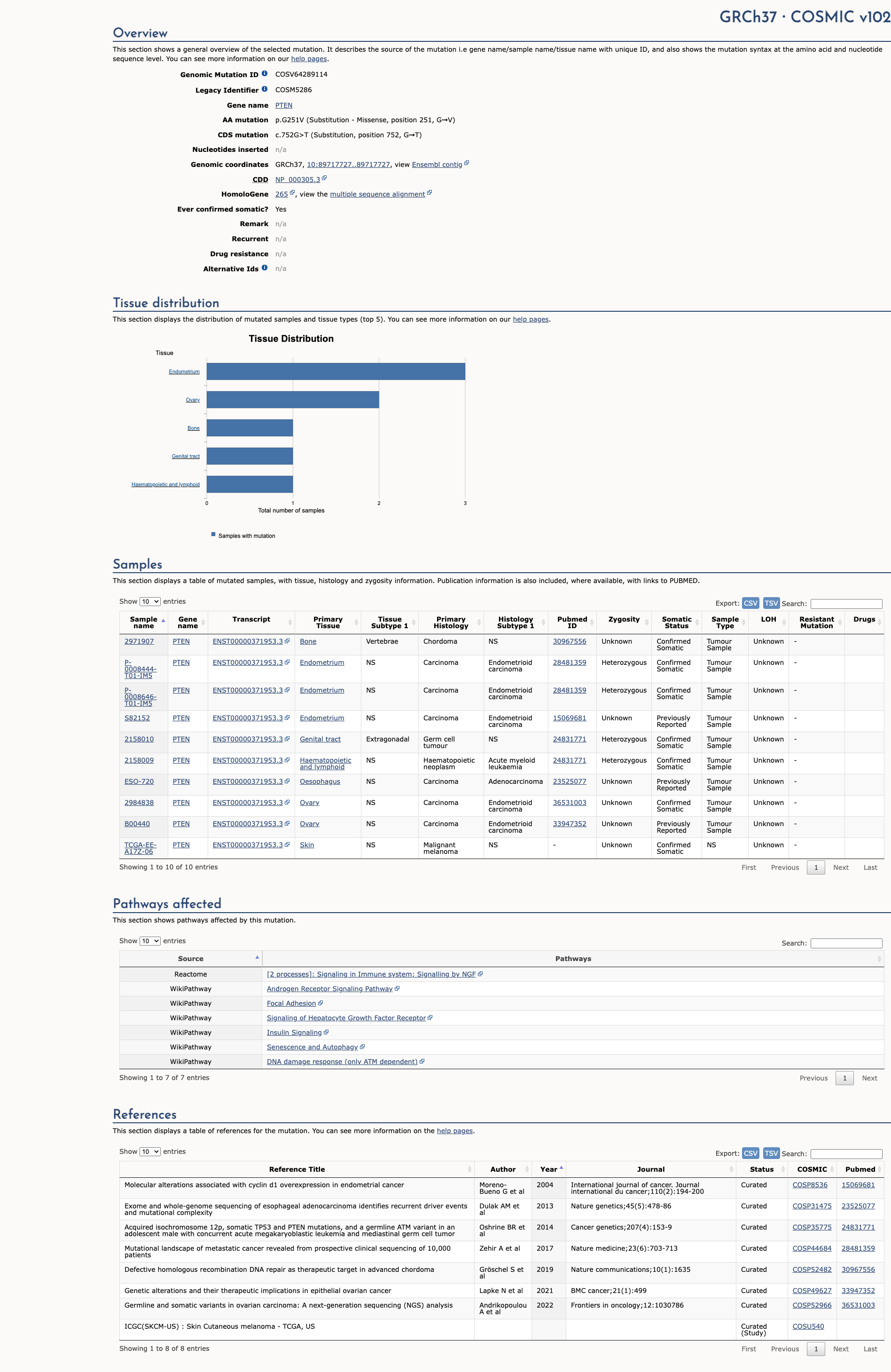
Functional Impact & Domains
Functional Domain
The PTEN G251V variant is functionally characterized as likely inactivating. It is located in the C2 tensin-type domain of the PTEN protein. In vitro studies demonstrate that this mutation increases cellular viability compared to the wildtype, suggesting a loss of PTEN protein function.
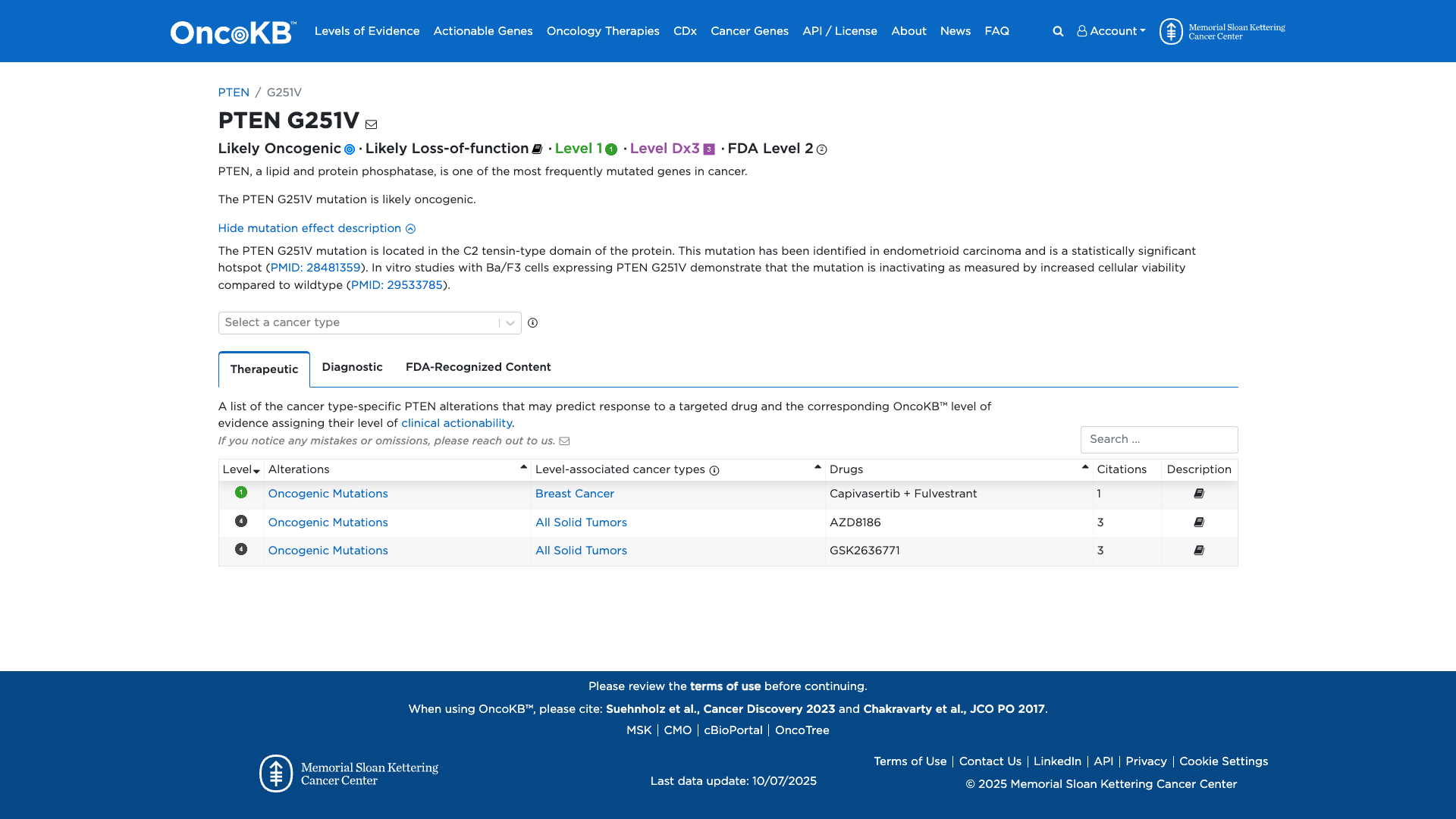
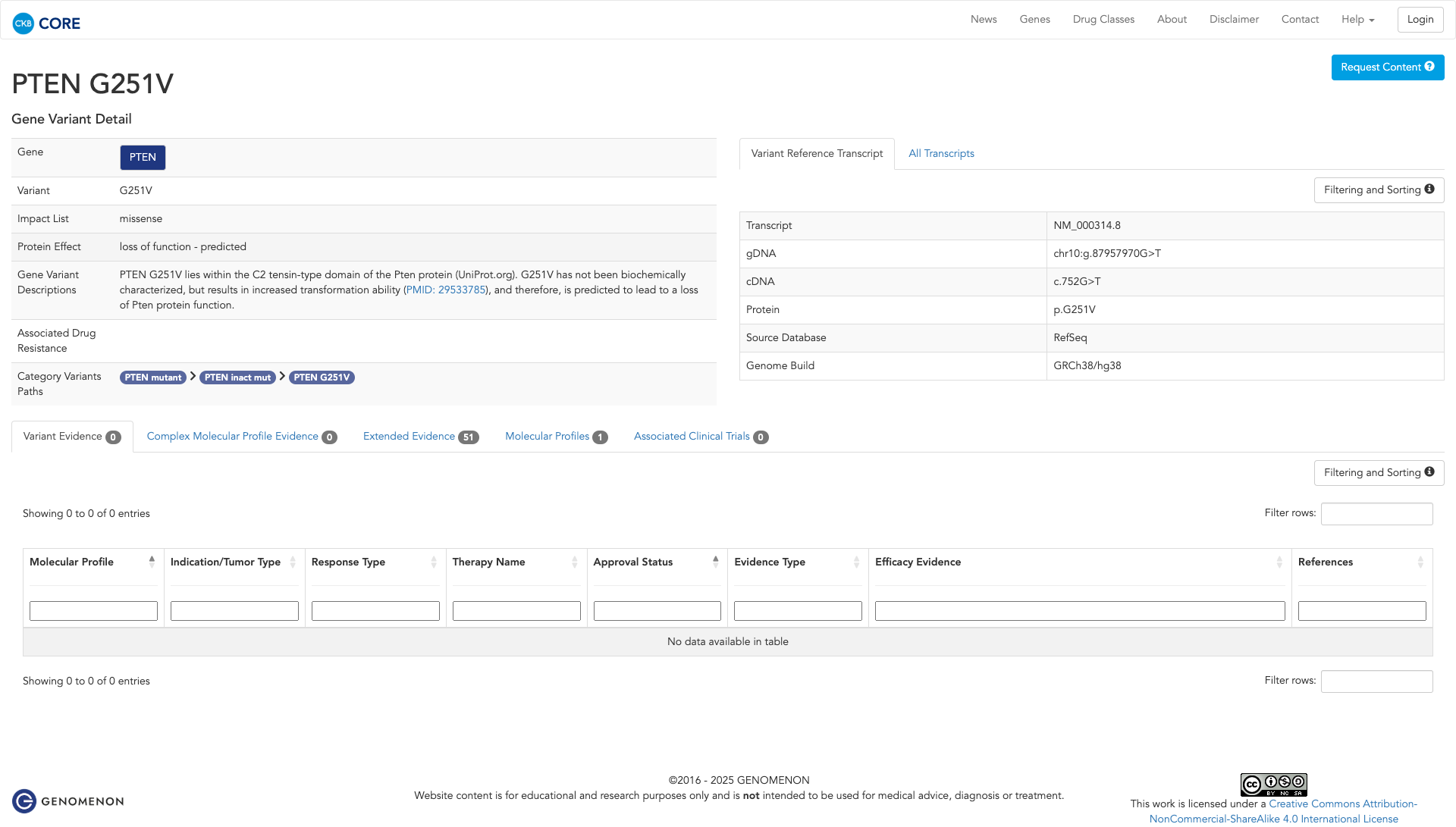
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.06 | -63 bp |
| Donor Loss (DL) | 0.0 | -215 bp |
| Acceptor Gain (AG) | 0.01 | -261 bp |
| Donor Gain (DG) | 0.0 | -138 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, the rule for PVS1 is: "Use PTEN PVS1 decision tree for predicting loss-of-function variants." The evidence for this variant shows: transcript and variant type information are unknown, preventing application of the decision tree. Therefore, this criterion is not applied.
PS1 (Not Applied)
According to VCEP guidelines, the rule for PS1 is: "Same amino acid change as a previously established pathogenic variant regardless of nucleotide change." The evidence for this variant shows: no other variant produces G251V in PTEN. Therefore, this criterion is not applied.
PS2 (Not Applied)
According to VCEP guidelines, the rule for PS2 is: "Very Strong: Two proven or four assumed de novo observations in a patient with the disease and no family history." The evidence for this variant shows: no de novo data available. Therefore, this criterion is not applied.
PS3 (Not Applied)
According to PTEN Pre-processing, the finding for PS3 is: "High confidence not TRUE ('FALSE'), PS3 rule not applied." The evidence for this variant shows: PTEN-specific functional evidence did not meet high confidence thresholds. Therefore, this criterion is not applied.
PS4 (Not Applied)
According to VCEP guidelines, the rule for PS4 is: "Variant prevalence in affected individuals significantly increased versus controls or proband specificity score ≥4." The evidence for this variant shows: no case-control or proband specificity data. Therefore, this criterion is not applied.
PM1 (Not Applied)
According to VCEP guidelines, the rule for PM1 is: "Located in a mutational hotspot or critical domain (residues 90-94, 123-130, 166-168)." The evidence for this variant shows: amino acid position 251 is outside these defined PTEN hotspots/domains. Therefore, this criterion is not applied.
PM2 (Supporting)
According to VCEP guidelines, the rule for PM2 is: "Supporting: Absent in population databases at allele frequency <0.00001 (0.001%)." The evidence for this variant shows: not present in gnomAD (MAF=0%). Therefore, this criterion is applied at Supporting strength.
PM3 (Not Applied)
According to standard ACMG guidelines, the rule for PM3 is: "For recessive disorders, observed in trans with a pathogenic variant." The evidence for this variant shows: no recessive inheritance or trans data. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to VCEP guidelines, the rule for PM4 is: "Protein length changes due to in-frame indels in non-repeat regions or stop-loss variants." The evidence for this variant shows: this is a missense substitution, not an indel. Therefore, this criterion is not applied.
PM5 (Moderate)
According to VCEP guidelines, the rule for PM5 is: "Moderate: Missense change at an amino acid residue where a different missense change determined to be pathogenic or likely pathogenic has been seen." The evidence for this variant shows: another pathogenic missense at residue G251 has been reported. Therefore, this criterion is applied at Moderate strength.
PM6 (Not Applied)
According to VCEP guidelines, the rule for PM6 is: "Very Strong: Two proven or four assumed de novo observations." The evidence for this variant shows: no assumed de novo data. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to VCEP guidelines, the rule for PP1 is: "Supporting: Co-segregation with disease in 3–4 meioses." The evidence for this variant shows: no family segregation data. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to VCEP guidelines, the rule for PP2 is: "Supporting: Missense variant in a gene with low benign missense variation and where missense is a common disease mechanism." The evidence for this variant shows: PTEN exhibits a range of benign and pathogenic missense variants. Therefore, this criterion is not applied.
PP3 (Supporting)
According to VCEP guidelines, the rule for PP3 is: "Supporting: Multiple computational lines support deleterious effect; REVEL score >0.7." The evidence for this variant shows: REVEL=0.97 exceeds 0.7. Therefore, this criterion is applied at Supporting strength.
PP4 (Not Applied)
According to standard ACMG guidelines, the rule for PP4 is: "Patient’s phenotype or family history highly specific for a disease with a single genetic etiology." The evidence for this variant shows: no phenotype data provided. Therefore, this criterion is not applied.
PP5 (Supporting)
According to standard ACMG guidelines, the rule for PP5 is: "Supporting: Reputable source reports variant as pathogenic without available evidence." The evidence for this variant shows: ClinVar and ClinGen PTEN VCEP classify it as Likely Pathogenic. Therefore, this criterion is applied at Supporting strength.
BA1 (Not Applied)
According to VCEP guidelines, the rule for BA1 is: "Stand Alone: gnomAD allele frequency >0.00056." The evidence for this variant shows: allele frequency is 0%. Therefore, this criterion is not applied.
BS1 (Not Applied)
According to VCEP guidelines, the rule for BS1 is: "Strong: allele frequency 0.000043–0.00056." The evidence for this variant shows: allele frequency is 0%. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to VCEP guidelines, the rule for BS2 is: "Strong: Observed homozygous in a healthy individual." The evidence for this variant shows: no homozygous observations. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to VCEP guidelines, the rule for BS3 is: "Strong: Well-established functional studies show no damaging effect." The evidence for this variant shows: functional data suggest loss of function. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to VCEP guidelines, the rule for BS4 is: "Strong: Lack of segregation in affected members." The evidence for this variant shows: no segregation data. Therefore, this criterion is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, the rule for BP1 is: "Supporting: Missense variant in a gene where truncation is main mechanism." The evidence for this variant shows: missense is a known disease mechanism in PTEN. Therefore, this criterion is not applied.
BP2 (Not Applied)
According to VCEP guidelines, the rule for BP2 is: "Supporting: Observed in trans with a pathogenic PTEN variant." The evidence for this variant shows: no trans data. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to VCEP guidelines, the rule for BP3 is: "Supporting: In-frame indel in a repetitive region." The evidence for this variant shows: this is a missense substitution, not an indel. Therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP guidelines, the rule for BP4 is: "Supporting: Multiple computational lines suggest no impact; REVEL <0.5." The evidence for this variant shows: REVEL=0.97. Therefore, this criterion is not applied.
BP5 (Not Applied)
According to VCEP guidelines, the rule for BP5 is: "Supporting: Variant found in a case with an alternate molecular basis for disease." The evidence for this variant shows: no alternate molecular basis. Therefore, this criterion is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, the rule for BP6 is: "Supporting: Reputable source reports variant as benign." The evidence for this variant shows: no benign assertions from reputable sources. Therefore, this criterion is not applied.
BP7 (Not Applied)
According to VCEP guidelines, the rule for BP7 is: "Supporting: Synonymous variant in non-splice regions with no predicted impact." The evidence for this variant shows: it is missense, not synonymous. Therefore, this criterion is not applied.