Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis sequence change replaces glycine, which is neutral and non-polar, with cysteine, which is neutral and slightly polar, at codon 244 of the TP53 protein (p.Gly244Cys). This variant is not present in population databases (gnomAD no frequency). This variant has not been reported in the literature in individuals affected with TP53-related conditions. ClinVar contains an entry for this variant (Variation ID: 376599). Advanced modeling performed at Invitae incorporating data from internal and/or published experimental studies (PMID: 12826609, 29979965, 30224644) indicates that this missense variant is expected to disrupt TP53 function with a positive predictive value of 97.5%. Experimental studies have shown that this missense change affects TP53 function (PMID: 12826609, 29979965). In summary, the available evidence is currently insufficient to determine the role of this variant in disease. Therefore, it has been classified as a Variant of Uncertain Significance.
"This variant has been reported in ClinVar as Pathogenic (2 clinical laboratories) and as Uncertain significance (1 clinical laboratories) and as Likely pathogenic (1 clinical laboratories)."
COSMIC Somatic Evidence
Open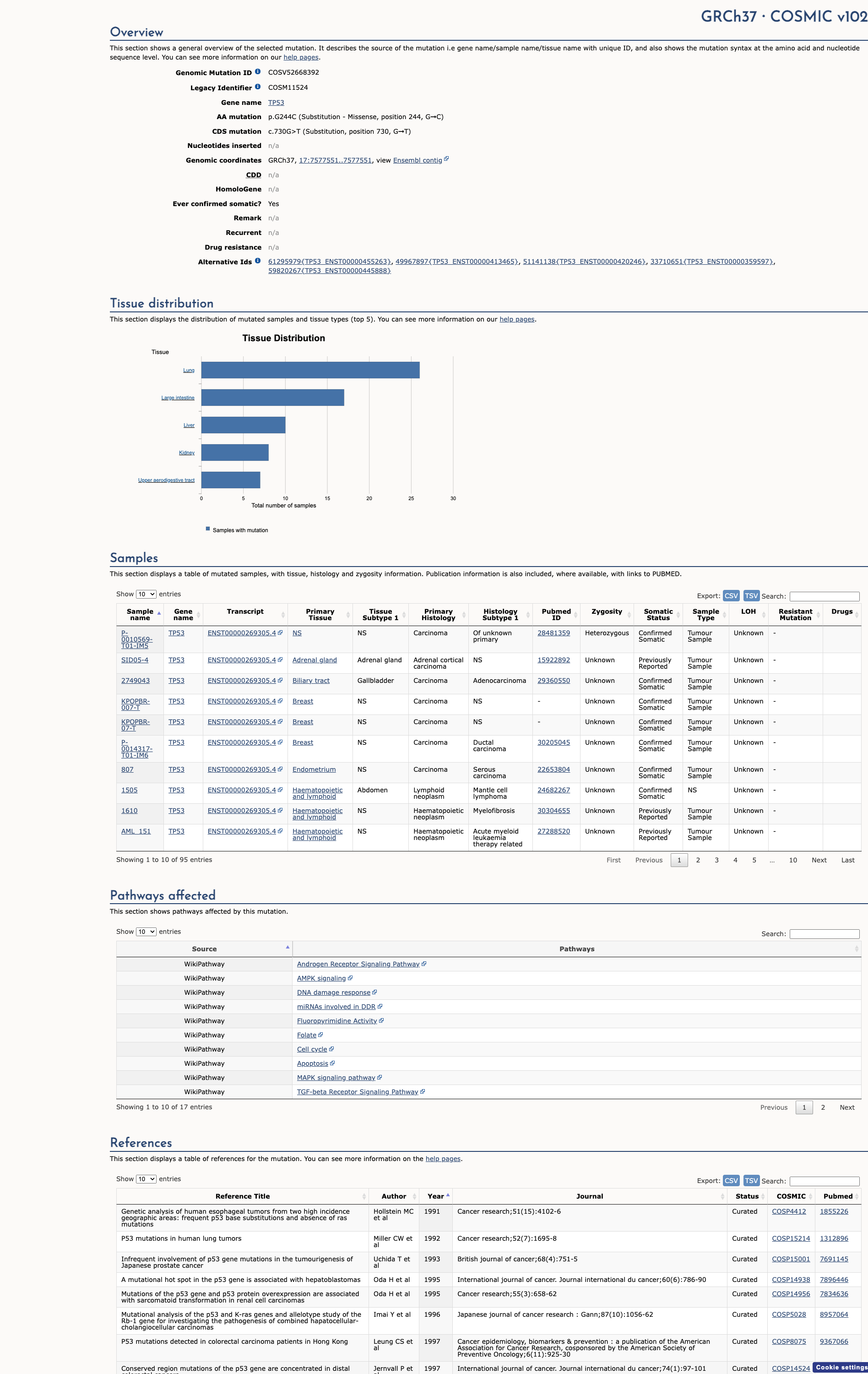
Functional Impact & Domains
Functional Domain
The TP53 G244C variant has been functionally characterized as inactivating, as demonstrated by in vivo studies in yeast showing a loss of transactivational activity compared to the wildtype. This suggests a likely loss-of-function effect.
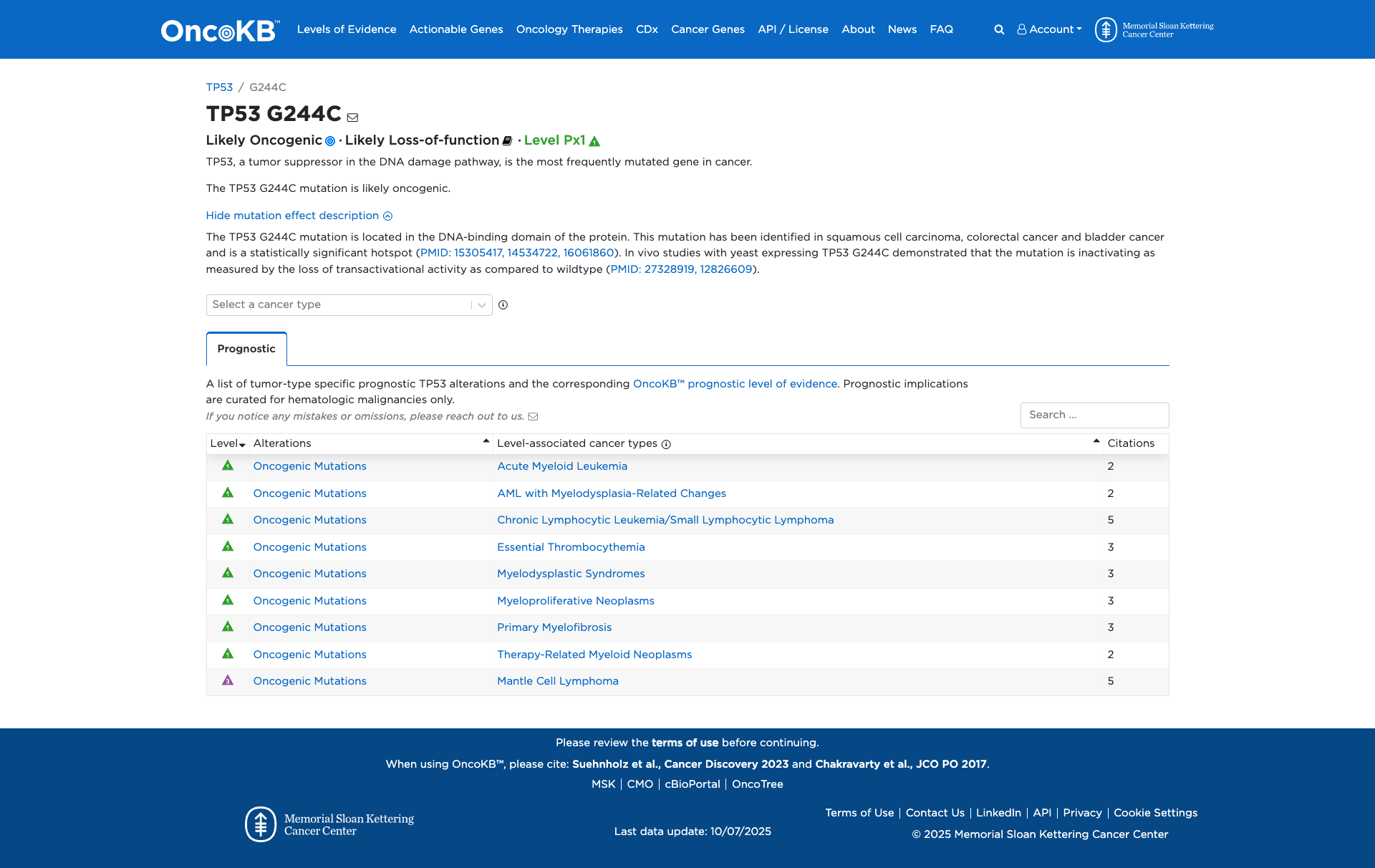
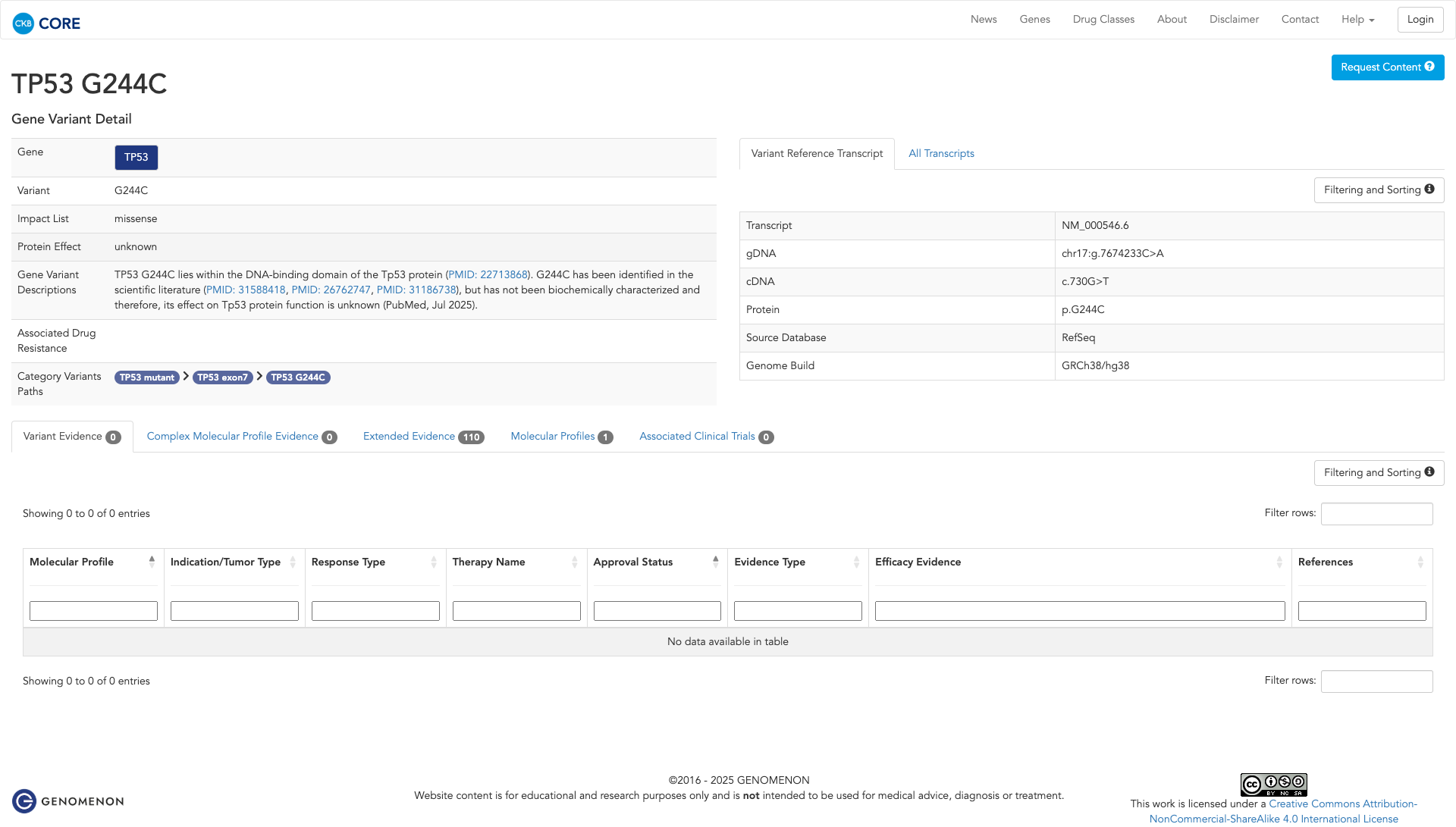
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -393 bp |
| Donor Loss (DL) | 0.0 | -52 bp |
| Acceptor Gain (AG) | 0.0 | 10 bp |
| Donor Gain (DG) | 0.0 | -342 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP TP53-specific guidelines, PVS1 applies to null variants (nonsense, frameshift, canonical splice) leading to NMD. The evidence for this variant shows: a missense change (G244C). Therefore, this criterion is not applied because the variant type does not meet PVS1 requirements.
PS1 (Not Applied)
According to VCEP TP53-specific guidelines, PS1 applies when a different nucleotide change results in the same amino acid change as a previously established pathogenic variant. The evidence for this variant shows: no other nucleotide change at codon 244 known to be pathogenic. Therefore, this criterion is not applied.
PS2 (Not Applied)
According to VCEP TP53-specific guidelines, PS2 requires confirmed de novo occurrence with points. The evidence for this variant shows: no de novo data available. Therefore, this criterion is not applied.
PS3 (Not Applied)
According to VCEP TP53-specific guidelines, PS3 requires non-functional on Kato et al. assay AND loss of function on a second assay for Strong, or specified combinations for Moderate/Supporting. The evidence for this variant shows: loss of transactivation in a yeast assay (Kato) but no second assay data. Therefore, this criterion is not applied because the evidence does not meet the multi-assay requirement.
PS4 (Not Applied)
According to VCEP TP53-specific guidelines, PS4 requires statistical enrichment with proband points. The evidence for this variant shows: no case–control or proband point data. Therefore, this criterion is not applied.
PM1 (Not Applied)
According to VCEP TP53-specific guidelines, PM1 applies at Moderate strength for missense variants in codons 175, 245, 248, 249, 273, or 282. The evidence for this variant shows: codon 244, not listed. Therefore, this criterion is not applied.
PM2 (Supporting)
According to VCEP TP53-specific guidelines, the rule for PM2_Supporting is: 'This rule should be applied at supporting level for allele frequency <0.00003 in gnomAD.' The evidence for this variant shows: absent from gnomAD. Therefore, PM2 is applied at Supporting strength because the variant is absent in population databases.
PM3 (Not Applied)
According to standard ACMG guidelines, PM3 applies to recessive disorders with variants in trans with a pathogenic variant. The evidence for this variant shows: TP53 is dominant and no in trans data. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, PM4 applies to protein length changes (in-frame indels). The evidence for this variant shows: a missense change with no length alteration. Therefore, this criterion is not applied.
PM5 (Not Applied)
According to VCEP TP53-specific guidelines, PM5 applies when ≥2 different missense changes at the same residue are pathogenic (Strong) or 1 is pathogenic (Moderate). The evidence for this variant shows: no other pathogenic missense reported at codon 244. Therefore, this criterion is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines, PM6 applies to presumed de novo variants without confirmation. The evidence for this variant shows: no de novo data. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to VCEP TP53-specific guidelines, PP1 requires cosegregation data across meiotic events. The evidence for this variant shows: no family segregation data. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, PP2 applies when missense variants are common in a gene with low benign missense variation. TP53 does not meet this criterion. Therefore, this criterion is not applied.
PP3 (Not Applied)
According to VCEP TP53-specific guidelines, PP3 requires BayesDel or aGVGD scores per flowchart. The evidence for this variant shows: only a REVEL score, no BayesDel/aGVGD. Therefore, this criterion is not applied.
PP4 (Not Applied)
According to VCEP TP53-specific guidelines, PP4 applies when specific LFS-associated phenotype observations are present. The evidence for this variant shows: no phenotype data. Therefore, this criterion is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, PP5 is not recommended and applies to assertions without available evidence. The evidence for this variant shows conflicting ClinVar assertions. Therefore, this criterion is not applied.
BA1 (Not Applied)
According to VCEP TP53-specific guidelines, BA1 applies at Stand Alone strength for allele frequency ≥0.001. The evidence for this variant shows: absent in gnomAD. Therefore, this criterion is not applied.
BS1 (Not Applied)
According to VCEP TP53-specific guidelines, BS1 applies at Strong strength for allele frequency ≥0.0003. The evidence for this variant shows: absent in gnomAD. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to VCEP TP53-specific guidelines, BS2 requires ≥2 unrelated elderly unaffected females. The evidence for this variant shows: no such data. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to VCEP TP53-specific guidelines, BS3 requires functional evidence of no loss of function across assays. The evidence for this variant shows: loss of function in yeast. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to VCEP TP53-specific guidelines, BS4 requires lack of segregation in affected members. The evidence for this variant shows: no segregation data. Therefore, this criterion is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, BP1 applies to missense variants in genes where only truncating causes disease. TP53 has pathogenic missense. Therefore, this criterion is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, BP2 applies to variants seen in cis/trans with pathogenic. The evidence for this variant shows: no such data. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, BP3 applies to in-frame indels in repetitive regions. The evidence for this variant shows: missense change. Therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP TP53-specific guidelines, BP4 requires BayesDel <0.16 and no splicing impact. The evidence for this variant shows: only REVEL and no predicted splice effect. Therefore, this criterion is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, BP5 applies when variant found with alternative molecular cause. The evidence for this variant shows: no alternative cause. Therefore, this criterion is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, BP6 applies to benign assertions without evidence. The evidence for this variant shows: no such reliable benign assertion. Therefore, this criterion is not applied.
BP7 (Not Applied)
According to VCEP TP53-specific guidelines, BP7 applies to synonymous variants with no splice impact. The evidence for this variant shows: missense change. Therefore, this criterion is not applied.