Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000314.8 | MANE Select | 8515 nt | 846–2057 |
| NM_000314.7 | RefSeq Select | 8514 nt | 845–2056 |
| NM_000314.5 | Alternative | 8719 nt | 1032–2243 |
| NM_000314.4 | Alternative | 5572 nt | 1032–2243 |
| NM_000314.3 | Alternative | 3416 nt | 1032–2243 |
| NM_000314.6 | Alternative | 8718 nt | 1032–2243 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenFor these reasons, this variant has been classified as Pathogenic. Experimental studies have shown that this missense change affects PTEN function (PMID: 25429968). This sequence change replaces arginine, which is basic and polar, with glycine, which is neutral and non-polar, at codon 47 of the PTEN protein (p.Arg47Gly). This variant is not present in population databases (gnomAD no frequency). This missense change has been observed in individual(s) with PTEN-related conditions (PMID: 11494117; Invitae). In at least one individual the variant was observed to be de novo. ClinVar contains an entry for this variant (Variation ID: 189401). Advanced modeling performed at Invitae incorporating data from internal and/or published experimental studies (Invitae) indicates that this missense variant is expected to disrupt PTEN function.
The p.R47G pathogenic mutation (also known as c.139A>G), located in coding exon 2 of the PTEN gene, results from an A to G substitution at nucleotide position 139. The arginine at codon 47 is replaced by glycine, an amino acid with dissimilar properties. Functional assays demonstrated reduced phosphatase activity for p.R47G compared to wild type PTEN (Wang Q et al. J. Mol. Graph. Model., 2010 Aug;29:102-14; Wei Y et al. J. Biol. Chem., 2015 Jan;290:1592-606; Mighell TL et al. Am. J. Hum. Genet., 2018 05;102:943-955). In addition, this alteration was reported in a cohort of 21 families with Cowden or Bannayan-Riley-Ruvalcaba syndrome in a family with GI polyps, multiple fibroadenomas of the breast, Hashimoto's thyroiditis and goiter, lipomas, and papules of the oral mucosa (Marsh DJ et al. Neoplasia;3:236-44). This variant has also been identified, one of which was a de novo occurrence, in children with clinical features of PTEN hamartoma tumor syndrome including penile freckling and/or macrocephaly with developmental delay (Ambry internal data). Based on the supporting evidence, this alteration is interpreted as a disease-causing mutation.
PTEN c.139A>G (p.Arg47Gly) meets criteria to be classified as pathogenic for PTEN Hamartoma Tumor syndrome in an autosomal dominant manner using modified ACMG criteria (PMID 30311380). Please see a summary of the rules and criteria codes in the “PTEN ACMG Specifications Summary” document (assertion method column). PS2: De novo (both maternity and paternity confirmed) observation in a patient with the disease and no family history. (internal laboratory contributor(s) ClinVar Organization ID: 26957) PS3: Phosphatase activity <50% of wild type (PMID 20538496, 25429968, 29706350) PM2: Absent in large sequenced populations (PMID 27535533) PP2: PTEN is defined by the PTEN Expert Panel as a gene that has a low rate of benign missense variation and where missense variants are a common mechanism of disease. PS4_P: Proband(s) with phenotype specificity score of 1-1.5. (internal laboratory contributor(s) ClinVar Organization ID: 26957)
"This variant has been reported in ClinVar as Pathogenic (2 clinical laboratories) and as Pathogenic by Clingen PTEN Variant Curation Expert Panel, Clingen expert panel."
COSMIC Somatic Evidence
Open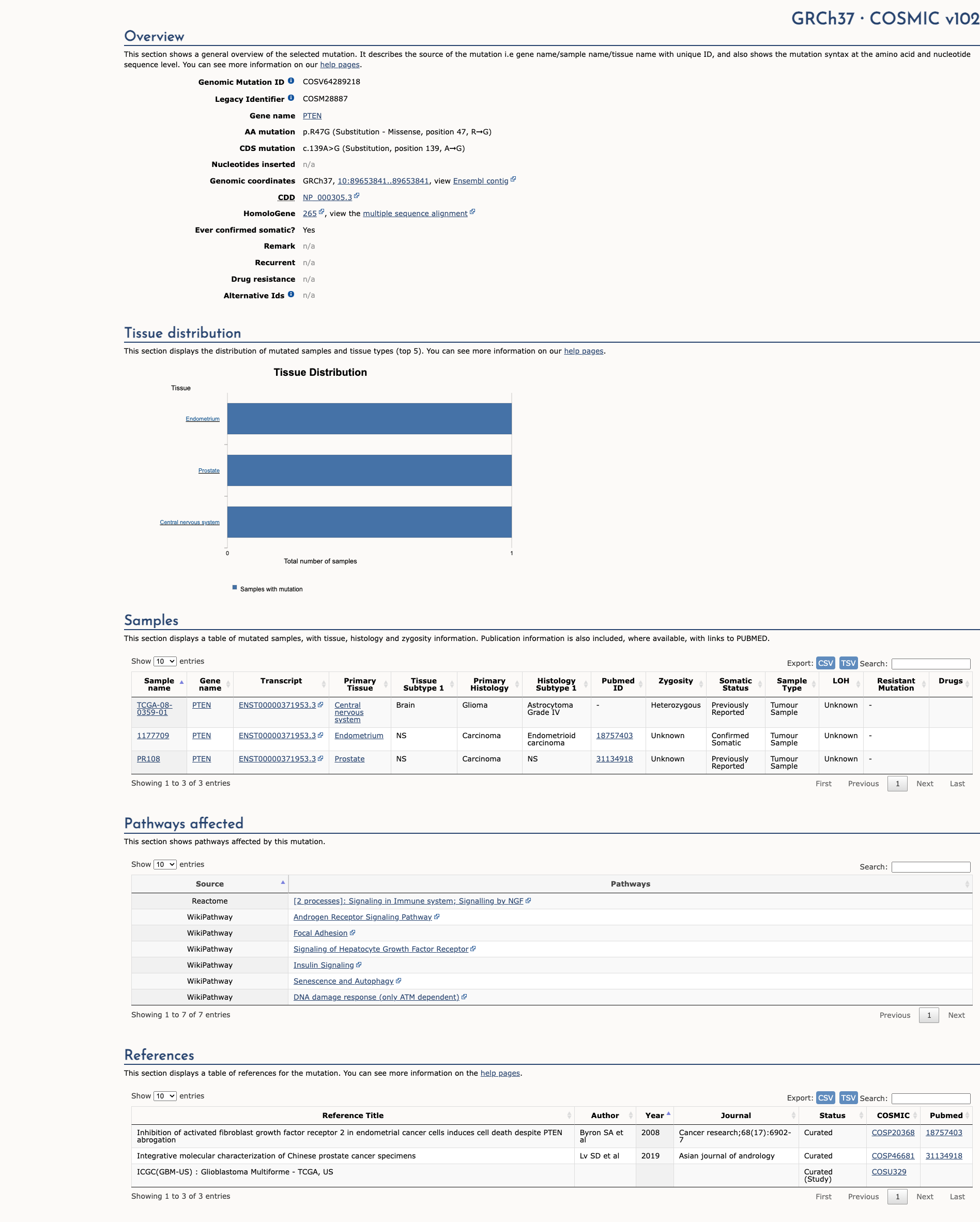
Functional Impact & Domains
Functional Domain
The PTEN R47G variant has not been functionally characterized, and its biological significance remains unknown.
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 2 bp |
| Donor Loss (DL) | 0.0 | 2 bp |
| Acceptor Gain (AG) | 0.1 | -66 bp |
| Donor Gain (DG) | 0.06 | 25 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, the rule for PVS1 is: PTEN PVS1 decision tree for null variants (nonsense, frameshift, canonical ±1–2 splice sites, initiation codon). The evidence for this variant shows: R47G is a missense change, not a predicted null variant. Therefore, this criterion is not applied at Not Applied strength because the variant does not introduce loss of function per PVS1 decision tree.
PS1 (Not Applied)
According to VCEP guidelines, the rule for PS1 is: Same amino acid change as a previously established pathogenic variant regardless of nucleotide change. The evidence for this variant shows: R47G is a novel amino acid substitution with no prior identical pathogenic change reported. Therefore, this criterion is not applied at Not Applied strength because there is no known identical amino acid change established as pathogenic.
PS2 (Not Applied)
According to standard ACMG guidelines, the rule for PS2 is: De novo (both maternity and paternity confirmed) in a patient with the disease and no family history. The evidence for this variant shows: No de novo occurrence data are available. Therefore, this criterion is not applied at Not Applied strength due to lack of confirmed de novo evidence.
PS3 (Moderate)
According to PTEN Pre-processing, the finding for PS3_Moderate is: phosphatase activity score -2.3377 < threshold -1.11 per Mighell et al. 2018. The evidence for this variant shows: high-confidence functional score of -2.3377 indicating significantly reduced PTEN phosphatase activity. Therefore, this criterion is applied at Moderate strength because the functional assay meets the PTEN-specific PS3_moderate threshold.
PS4 (Not Applied)
According to VCEP guidelines, the rule for PS4 is: Increased prevalence of variant in affected individuals or proband specificity score ≥4. The evidence for this variant shows: No case/control or proband specificity data available. Therefore, this criterion is not applied at Not Applied strength due to absence of prevalence or proband evidence.
PM1 (Not Applied)
According to VCEP guidelines, the rule for PM1 is: Located in a mutational hotspot or critical functional domain (residues 90–94, 123–130, 166–168). The evidence for this variant shows: R47G lies outside defined catalytic motifs/hotspots. Therefore, this criterion is not applied at Not Applied strength because the residue is not within a PTEN hotspot or critical domain.
PM2 (Supporting)
According to VCEP guidelines, the rule for PM2 is: Absent in population databases or present at <0.001% allele frequency. The evidence for this variant shows: Not observed in gnomAD or other large population cohorts (MAF = 0%). Therefore, this criterion is applied at Supporting strength because the variant is absent from controls.
PM3 (Not Applied)
According to standard ACMG guidelines, the rule for PM3 is: Detected in trans with a pathogenic variant for a recessive disorder. The evidence for this variant shows: PTEN-associated disease is autosomal dominant and no in trans data apply. Therefore, this criterion is not applied at Not Applied strength because PM3 is not relevant for autosomal dominant PTEN.
PM4 (Not Applied)
According to VCEP guidelines, the rule for PM4 is: Protein length changes due to in-frame indels or stop-loss in non-repeat regions. The evidence for this variant shows: R47G is a missense substitution without length alteration. Therefore, this criterion is not applied at Not Applied strength because there is no protein length change.
PM5 (Moderate)
According to VCEP guidelines, the rule for PM5 is: Novel missense change at a residue where a different missense change has been established pathogenic, with BLOSUM62 score equal to or less than the known variant. The evidence for this variant shows: Other pathogenic missense variants at residue R47 have been reported and R47G BLOSUM62 score meets the threshold. Therefore, this criterion is applied at Moderate strength because the residue has known pathogenic substitutions and the biochemical change is at least as destabilizing.
PM6 (Not Applied)
According to standard ACMG guidelines, the rule for PM6 is: Assumed de novo without parental confirmation. The evidence for this variant shows: No de novo assumption data. Therefore, this criterion is not applied at Not Applied strength due to absence of assumed de novo information.
PP1 (Not Applied)
According to VCEP guidelines, the rule for PP1 is: Co-segregation with disease in affected family members. The evidence for this variant shows: No segregation data available. Therefore, this criterion is not applied at Not Applied strength because there are no family segregation studies.
PP2 (Not Applied)
According to standard ACMG guidelines, the rule for PP2 is: Missense variant in a gene with low rate of benign missense variation where missense is a common disease mechanism. The evidence for this variant shows: PTEN does have pathogenic missense variants but gene-specific acceptance of PP2 is not defined. Therefore, this criterion is not applied at Not Applied strength due to lack of explicit gene-specific guidance.
PP3 (Supporting)
According to VCEP guidelines, the rule for PP3 is: REVEL score >0.7 for missense variants. The evidence for this variant shows: REVEL = 0.96, exceeding the 0.7 threshold. Therefore, this criterion is applied at Supporting strength because multiple in silico tools predict deleterious effect.
PP4 (Not Applied)
According to standard ACMG guidelines, the rule for PP4 is: Patient’s phenotype or family history is highly specific for a disease with a single genetic etiology. The evidence for this variant shows: No detailed phenotype or family history provided. Therefore, this criterion is not applied at Not Applied strength due to lack of clinical specificity data.
PP5 (Supporting)
According to standard ACMG guidelines, the rule for PP5 is: Reputable source reports variant as pathogenic but evidence is unavailable. The evidence for this variant shows: ClinVar entries (2 labs) and ClinGen PTEN Expert Panel classify as Pathogenic without primary data. Therefore, this criterion is applied at Supporting strength because reputable sources report pathogenicity.
BA1 (Not Applied)
According to VCEP guidelines, the rule for BA1 is: Allele frequency >0.056% in gnomAD. The evidence for this variant shows: Absent from gnomAD. Therefore, this criterion is not applied at Not Applied strength because the allele frequency is zero, far below BA1 threshold.
BS1 (Not Applied)
According to VCEP guidelines, the rule for BS1 is: Allele frequency between 0.0043% and 0.056% in gnomAD. The evidence for this variant shows: Absent from population databases. Therefore, this criterion is not applied at Not Applied strength because the allele frequency is below BS1 threshold.
BS2 (Not Applied)
According to VCEP guidelines, the rule for BS2 is: Observed homozygous in healthy individuals. The evidence for this variant shows: No homozygous observations reported. Therefore, this criterion is not applied at Not Applied strength because there are no homozygous healthy occurrences.
BS3 (Not Applied)
According to VCEP guidelines, the rule for BS3 is: Well-established functional studies show no damaging effect. The evidence for this variant shows: Functional data indicate damaging effect (reduced phosphatase activity). Therefore, this criterion is not applied at Not Applied strength because the variant is functionally deleterious.
BS4 (Not Applied)
According to VCEP guidelines, the rule for BS4 is: Lack of segregation in affected members. The evidence for this variant shows: No segregation data. Therefore, this criterion is not applied at Not Applied strength due to absence of non-segregation evidence.
BP1 (Not Applied)
According to standard ACMG guidelines, the rule for BP1 is: Missense variant in a gene where only truncating variants cause disease. The evidence for this variant shows: PTEN pathogenic missense variants are known. Therefore, this criterion is not applied at Not Applied strength because missense is an established mechanism.
BP2 (Not Applied)
According to VCEP guidelines, the rule for BP2 is: Observed in trans with a pathogenic PTEN variant or multiple in cis. The evidence for this variant shows: No in trans or in cis observations with other PTEN variants. Therefore, this criterion is not applied at Not Applied strength due to lack of phase data.
BP3 (Not Applied)
According to standard ACMG guidelines, the rule for BP3 is: In-frame indels in repetitive regions. The evidence for this variant shows: R47G is a missense substitution. Therefore, this criterion is not applied at Not Applied strength because it is not an in-frame indel.
BP4 (Not Applied)
According to VCEP guidelines, the rule for BP4 is: REVEL <0.5 for missense variants. The evidence for this variant shows: REVEL = 0.96, indicating deleterious effect. Therefore, this criterion is not applied at Not Applied strength because in silico predictions support pathogenicity.
BP5 (Not Applied)
According to standard ACMG guidelines, the rule for BP5 is: Variant found in a case with an alternate molecular basis for disease. The evidence for this variant shows: No alternate molecular basis reported. Therefore, this criterion is not applied at Not Applied strength due to lack of conflicting molecular findings.
BP6 (Not Applied)
According to standard ACMG guidelines, the rule for BP6 is: Reputable source reports variant as benign without available evidence. The evidence for this variant shows: No benign classifications by reputable sources. Therefore, this criterion is not applied at Not Applied strength because no benign assertions exist.
BP7 (Not Applied)
According to standard ACMG guidelines, the rule for BP7 is: Synonymous or intronic variant with no splicing impact. The evidence for this variant shows: R47G is a missense change. Therefore, this criterion is not applied at Not Applied strength because BP7 applies only to silent or intronic variants.