Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe p.E294* pathogenic mutation (also known as c.880G>T), located in coding exon 7 of the TP53 gene, results from a G to T substitution at nucleotide position 880. This changes the amino acid from a glutamic acid to a stop codon within coding exon 7. This alteration has been reported in an individual diagnosed with bilateral breast cancers at age 33 and 41 whose son was reported to have an osteosarcoma diagnosed at age 15 (Veschi S et al. Ann Oncol, 2007 Jun;18 Suppl 6:vi86-92). This alteration was also reported in a pediatric patient with a personal history of choroid plexus carcinoma, Spitzoid melanoma, and myelodysplasia (Kollipara R et al. Pediatr Dev Pathol Nov;17:64-9). This alteration has been observed numerous times as a somatic mutation in the cancerhotspots.org database (Chang MT et al. Cancer Discov. 2018 02;8:174-183). This variant is considered to be rare based on population cohorts in the Genome Aggregation Database (gnomAD). In addition to the clinical data presented in the literature, this alteration is expected to result in loss of function by premature protein truncation or nonsense-mediated mRNA decay. As such, this alteration is interpreted as a disease-causing mutation.
"This variant has been reported in ClinVar as Pathogenic (4 clinical laboratories)."
COSMIC Somatic Evidence
Open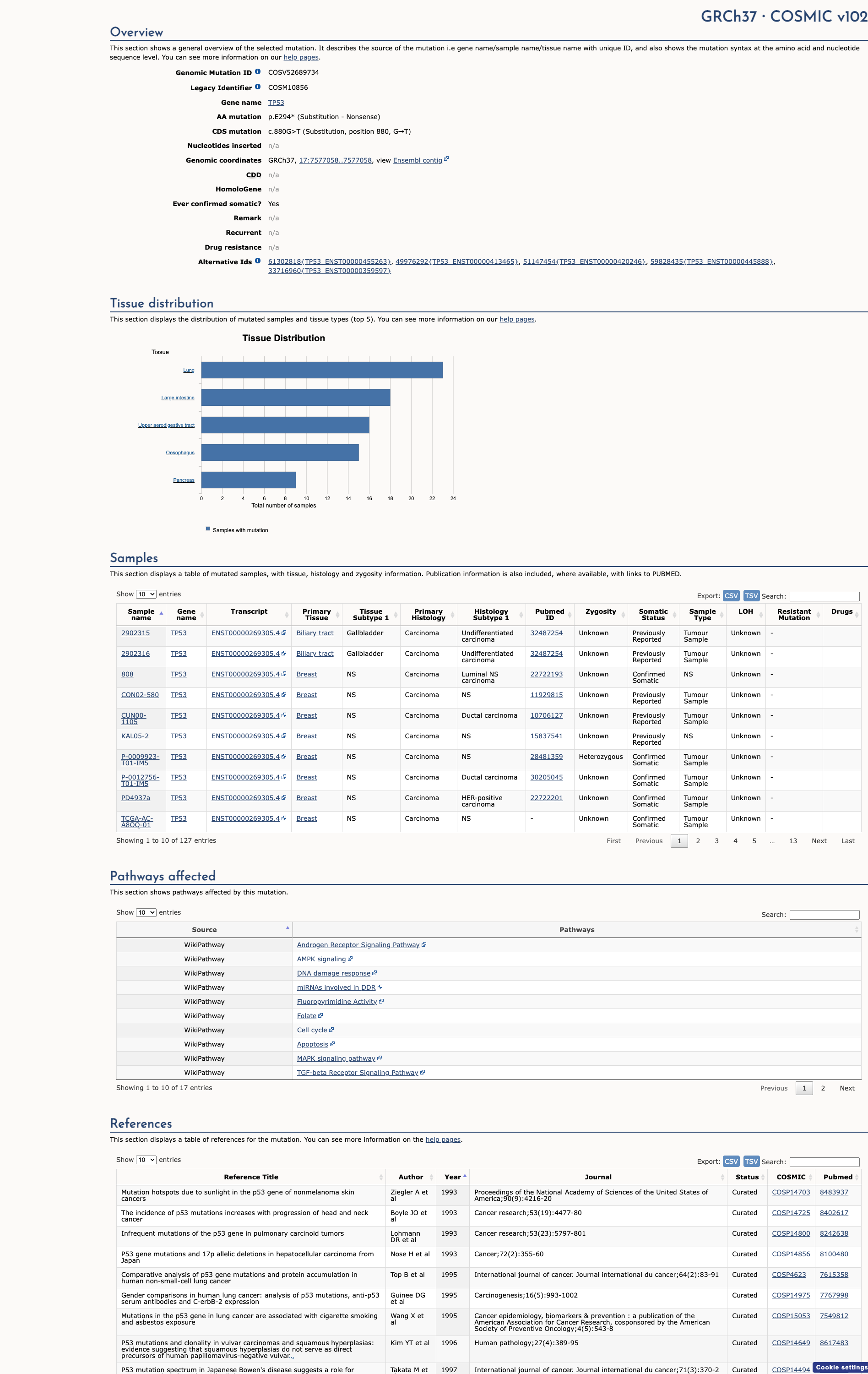
Functional Impact & Domains
Functional Domain
The TP53 E294* variant is a truncating mutation in the tumor suppressor gene TP53. Functional evidence indicates that truncating mutations in TP53 lead to the production of C-terminally truncated protein forms, which are predicted to be inactivating. Experimental studies have shown that these mutations promote cancer cell proliferation, survival, and metastasis, partly due to aberrant localization of truncated proteins to the mitochondria, affecting genes involved in cell survival. Therefore, the TP53 E294* variant is functionally characterized as likely damaging.
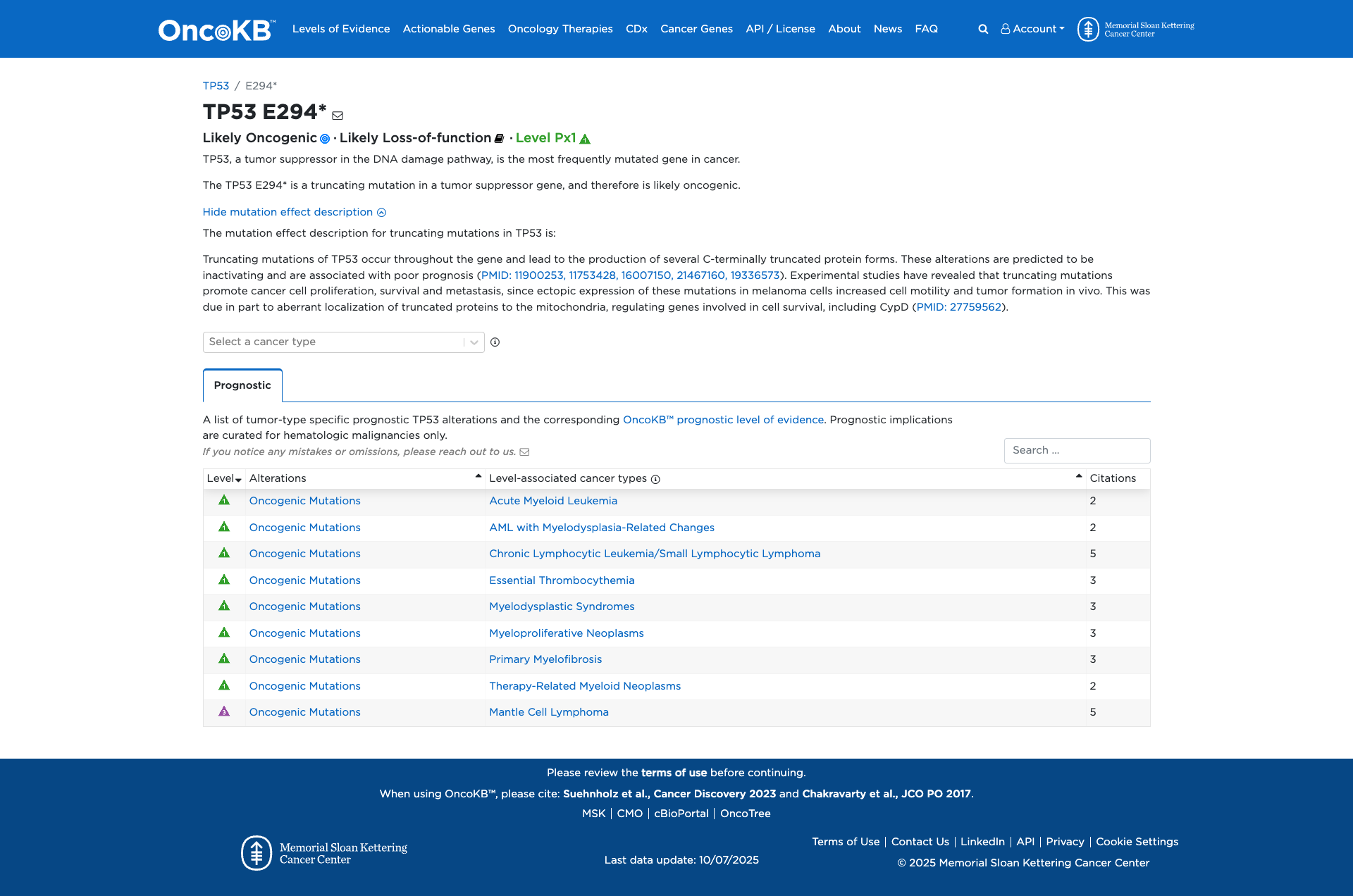

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.01 | 100 bp |
| Donor Loss (DL) | 0.0 | -39 bp |
| Acceptor Gain (AG) | 0.02 | -401 bp |
| Donor Gain (DG) | 0.01 | 2 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Very Strong)
According to VCEP guidelines, "PVS1 applies to variants predicted to result in nonsense-mediated decay (NMD) for nonsense variants upstream of p.Lys351". The evidence for this variant shows it introduces a premature stop at p.E294, upstream of p.Lys351 and predicted to undergo NMD. Therefore, PVS1 is applied at Very Strong strength because it is a null variant in a gene where loss of function is a known mechanism of disease.
PS1 (Not Applied)
According to VCEP guidelines, "PS1: Can be applied to variants asserted as Pathogenic following the TP53 VCEP’s specifications." The evidence for this variant shows no other nucleotide change results in p.E294* that has been asserted as pathogenic. Therefore, PS1 is not applied.
PS2 (Not Applied)
According to VCEP guidelines, "PS2_Very Strong: ≥ 8 points." No de novo evidence is available for this variant. Therefore, PS2 is not applied.
PS3 (Not Applied)
According to VCEP guidelines, "PS3 should not be applied at any strength if PVS1 is applied at full strength." PVS1 was applied at Very Strong strength. Therefore, PS3 is not applied.
PS4 (Not Applied)
According to VCEP guidelines, "PS4_Very Strong: ≥ 8 points; PS4_Strong: 4-7.5 points; PS4_Moderate: 2-3.5 points; PS4_Supporting: 1-1.5 points." No case-control or multiple proband data are available. Therefore, PS4 is not applied.
PM1 (Not Applied)
According to VCEP guidelines, "PM1: Moderate Strength for missense variants within codons 175, 245, 248, 249, 273, 282." This variant is a nonsense change. Therefore, PM1 is not applied.
PM2 (Supporting)
According to VCEP guidelines, "PM2_Supporting: allele frequency <0.00003 in gnomAD." The variant is absent from gnomAD. Therefore, PM2 is applied at Supporting strength.
PM3 (Not Applied)
According to standard ACMG guidelines, "PM3: For recessive disorders, detected in trans with a pathogenic variant." TP53 is associated with dominant inheritance and no compound heterozygosity is observed. Therefore, PM3 is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, "PM4: Protein length changes due to in-frame insertions, deletions, or stop-loss variants." This is a stop-gain variant. Therefore, PM4 is not applied.
PM5 (Not Applied)
According to VCEP guidelines, "PM5: Strong Strength for missense variants at residues with ≥2 pathogenic missense variants." This is a nonsense variant. Therefore, PM5 is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines, "PM6: Assumed de novo variant, without confirmation of paternity and maternity." No de novo evidence is available. Therefore, PM6 is not applied.
PP1 (Not Applied)
According to VCEP guidelines, "PP1: Supporting Strength requiring 3-4 meioses of cosegregation." No segregation data are available. Therefore, PP1 is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, "PP2: Missense variant in a gene with low rate of benign missense variation." This is a nonsense variant. Therefore, PP2 is not applied.
PP3 (Not Applied)
According to VCEP guidelines, "PP3: Supporting or Moderate strength for in silico predictions of missense variants." This is a nonsense variant. Therefore, PP3 is not applied.
PP4 (Not Applied)
According to standard ACMG guidelines, "PP4: Specific phenotype/family history for a disease with a single genetic etiology." No clinical phenotype data are provided. Therefore, PP4 is not applied.
PP5 (Supporting)
According to standard ACMG guidelines, "PP5: Reputable source reports variant as pathogenic without available evidence." ClinVar lists this variant as Pathogenic from multiple clinical laboratories. Therefore, PP5 is applied at Supporting strength.
BA1 (Not Applied)
According to VCEP guidelines, "BA1: Filtering allele frequency ≥0.001 in gnomAD continental subpopulations." The variant is absent from gnomAD. Therefore, BA1 is not applied.
BS1 (Not Applied)
According to VCEP guidelines, "BS1: Filtering allele frequency ≥0.0003 but <0.001." The variant is absent from gnomAD. Therefore, BS1 is not applied.
BS2 (Not Applied)
According to VCEP guidelines, "BS2: Strong requires ≥8 unaffected older females without cancer." No such data are available. Therefore, BS2 is not applied.
BS3 (Not Applied)
According to VCEP guidelines, "BS3: Strong for functional data showing no loss of function on assays." Functional studies indicate loss of function. Therefore, BS3 is not applied.
BS4 (Not Applied)
According to VCEP guidelines, "BS4: Strong requires lack of segregation among affected family members." No segregation data are available. Therefore, BS4 is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, "BP1: Missense variant in gene where truncating variants are known mechanism." This is a nonsense variant. Therefore, BP1 is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, "BP2: Observed in trans with a pathogenic variant in a dominant disorder." No such data are available. Therefore, BP2 is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, "BP3: In-frame deletions/insertions in repetitive region without known function." This is a nonsense variant. Therefore, BP3 is not applied.
BP4 (Not Applied)
According to VCEP guidelines, "BP4: Supporting strength for benign in silico predictions for missense variants." This is a nonsense variant. Therefore, BP4 is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, "BP5: Variant found with an alternate molecular basis for disease." No such data are available. Therefore, BP5 is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, "BP6: Reputable source reports variant as benign." No such reports exist. Therefore, BP6 is not applied.
BP7 (Not Applied)
According to VCEP guidelines, "BP7: Supporting strength for synonymous or intronic variants with no splicing impact." This is a nonsense variant. Therefore, BP7 is not applied.