Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_177438.2 | RefSeq Select | 10323 nt | 239–6007 |
| NM_177438.3 | MANE Select | 10384 nt | 346–6114 |
| NM_177438.1 | Alternative | 10276 nt | 239–6007 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenACMG criteria met: None
"This variant has been reported in ClinVar as Benign (6 clinical laboratories)."
COSMIC Somatic Evidence
Open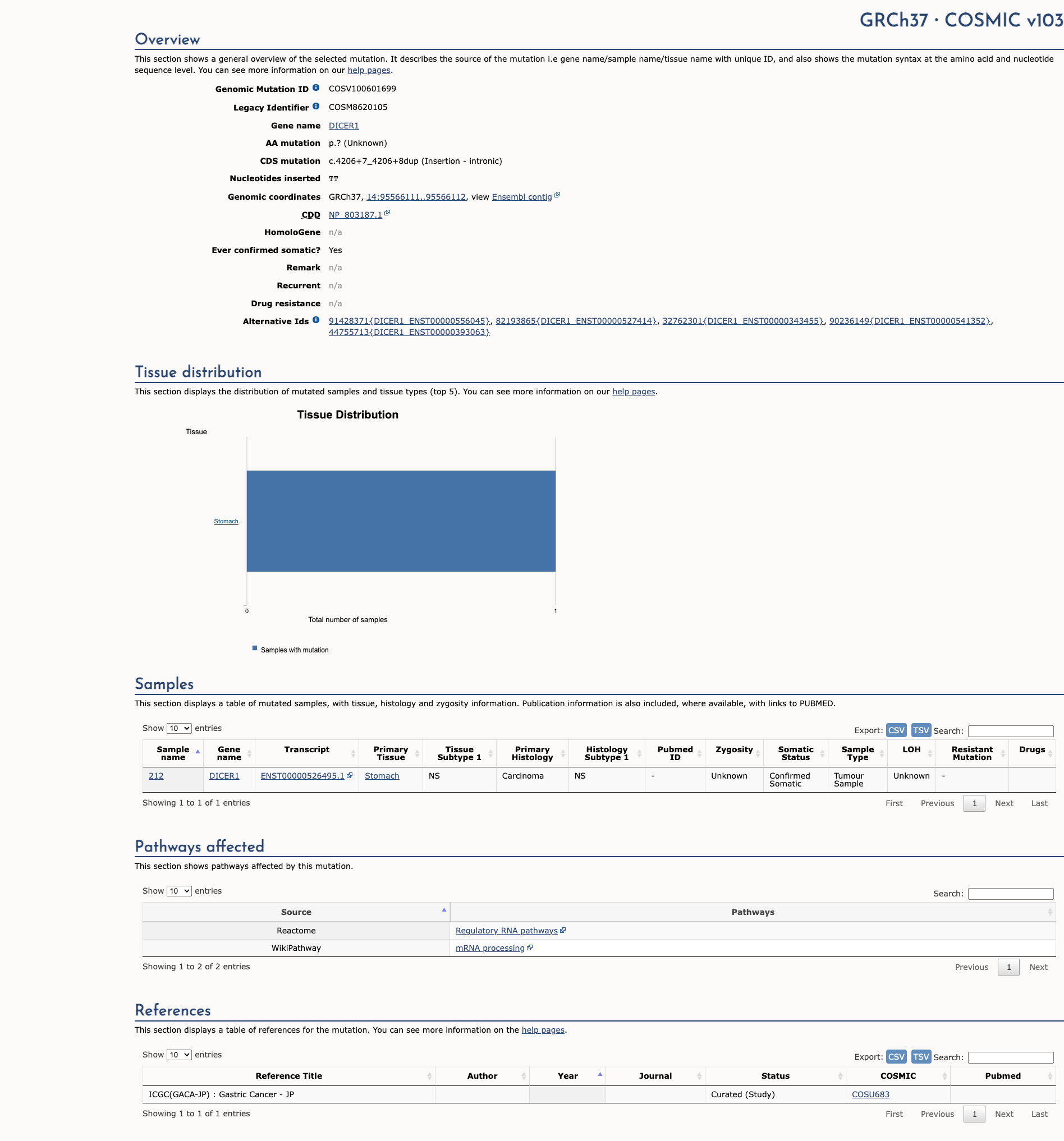
Functional Impact & Domains
Functional Domain
The DICER1 4206+7_4206+8dup variant has not been functionally characterized, and its biological significance remains unknown.
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.01 | 159 bp |
| Donor Loss (DL) | 0.0 | -61 bp |
| Acceptor Gain (AG) | 0.0 | 164 bp |
| Donor Gain (DG) | 0.0 | 9 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, the rule for PVS1 is: "Canonical splice variants (+/- 1,2 intronic positions): PVS1 applies with exceptions..." The evidence for this variant shows it is located at positions +7 and +8, outside the canonical splice donor/acceptor sites. Therefore, this criterion is not applied at Not Applied strength because the variant does not meet the requirement for canonical splice site disruption.
PS1 (Not Applied)
According to standard ACMG guidelines, the rule for PS1 is: "Strong Strength: same amino acid change as a known pathogenic variant with no difference in splicing." The evidence for this variant shows it is intronic with no predicted amino acid change. Therefore, this criterion is not applied at Not Applied strength.
PS2 (Not Applied)
According to standard ACMG guidelines, the rule for PS2 is: "Very Strong Strength: confirmed de novo (both maternity and paternity confirmed) in a patient with the disease and no family history." There is no evidence of de novo occurrence provided. Therefore, this criterion is not applied at Not Applied strength.
PS3 (Not Applied)
According to VCEP guidelines, the rule for PS3 is: "Strong: RNA assay shows splicing impact that is out-of-frame..." There are no functional RNA splicing assay data for this variant. Therefore, this criterion is not applied at Not Applied strength due to lack of functional evidence.
PS4 (Not Applied)
According to standard ACMG guidelines, the rule for PS4 is: "Strong Strength: statistical evidence from case-control data or phenotype points ≥4." No case-control or phenotype point data are available. Therefore, this criterion is not applied at Not Applied strength.
PM1 (Not Applied)
According to VCEP guidelines, the rule for PM1 is: "Moderate Strength: putative missense variants at metal ion-binding residues in RNase IIIb domain." This variant is intronic and not a missense change in the RNase IIIb domain. Therefore, this criterion is not applied at Not Applied strength.
PM2 (Not Applied)
According to VCEP guidelines, the rule for PM2 is: "Supporting Strength: allele frequency <0.000005 across gnomAD." The evidence for this variant shows a global MAF of approximately 0.00945 (0.945%) and up to 2.94% in African/African American populations, which exceeds the threshold. Therefore, this criterion is not applied at Not Applied strength.
PM3 (Not Applied)
According to standard ACMG guidelines, the rule for PM3 is: "Moderate Strength: detected in trans with a pathogenic variant in a recessive disorder." DICER1 disease mechanism is dominant, and no trans observations are reported. Therefore, this criterion is not applied at Not Applied strength.
PM4 (Not Applied)
According to VCEP guidelines, the rule for PM4 is: "Moderate Strength: in-frame indels within RNase IIIb domain; Supporting Strength: in-frame indels outside RNase IIIb domain." This variant is an intronic duplication, not an in-frame indel in coding sequence. Therefore, this criterion is not applied at Not Applied strength.
PM5 (Not Applied)
According to standard ACMG guidelines, the rule for PM5 is: "Moderate Strength: novel missense change at an amino acid residue where a different missense change is pathogenic." This variant is intronic, not a missense change. Therefore, this criterion is not applied at Not Applied strength.
PM6 (Not Applied)
According to standard ACMG guidelines, the rule for PM6 is: "Supporting Strength: assumed de novo, but without confirmation of paternity and maternity." No de novo evidence is provided. Therefore, this criterion is not applied at Not Applied strength.
PP1 (Not Applied)
According to standard ACMG guidelines, the rule for PP1 is: "Supporting Strength: cosegregation with disease in multiple affected family members." No segregation data are available. Therefore, this criterion is not applied at Not Applied strength.
PP2 (Not Applied)
According to standard ACMG guidelines, the rule for PP2 is: "Supporting Strength: missense variant in a gene with low rate of benign missense variation and pathogenic missense variants are common." This is an intronic variant, not missense. Therefore, this criterion is not applied at Not Applied strength.
PP3 (Not Applied)
According to VCEP guidelines, the rule for PP3 is: "Supporting Strength for splicing variants: concordance of MaxEntScan and SpliceAI predicting splicing effects." SpliceAI score is 0.01, predicting no significant splicing impact, and no MaxEntScan evidence of splice alteration. Therefore, this criterion is not applied at Not Applied strength.
PP4 (Not Applied)
According to VCEP guidelines, the rule for PP4 is: "Supporting Strength: somatic tumor testing identifies a second hit in hotspot codon and retention of germline variant." No tumor or somatic data are available. Therefore, this criterion is not applied at Not Applied strength.
PP5 (Not Applied)
According to standard ACMG guidelines, the rule for PP5 is: "Supporting Strength: reputable source reports variant as pathogenic without available evidence." No such pathogenic assertions exist. Therefore, this criterion is not applied at Not Applied strength.
BA1 (Stand Alone)
According to VCEP guidelines, the rule for BA1 is: "Stand Alone Strength: frequency >0.003 (0.3%) in gnomAD subpopulations." The evidence for this variant shows MAF 0.945% overall and 2.94% in African/African American populations, exceeding the threshold. Therefore, this criterion is applied at Stand Alone strength because the allele frequency is above the established benign threshold.
BS1 (Strong)
According to VCEP guidelines, the rule for BS1 is: "Strong Strength: frequency >0.0003 (0.03%) in gnomAD subpopulations." The evidence for this variant shows MAF 0.945% overall and 2.94% in African/African American populations, exceeding the threshold. Therefore, this criterion is applied at Strong strength because the allele frequency is substantially above the threshold.
BS2 (Strong)
According to VCEP guidelines, the rule for BS2 is: "Strong Strength: 2+ observations of homozygosity in healthy individuals." The evidence for this variant shows 578 homozygotes globally and 128 homozygotes in African/African American populations in gnomAD. Therefore, this criterion is applied at Strong strength because multiple homozygous observations in healthy individuals are documented.
BS3 (Not Applied)
According to VCEP guidelines, the rule for BS3 is: "Strong Strength: no splicing impact observed via RNA assay for intronic variants (observed more than once)." No RNA assay data are available for this variant. Therefore, this criterion is not applied at Not Applied strength.
BS4 (Not Applied)
According to standard ACMG guidelines, the rule for BS4 is: "Strong Strength: non-segregation in affected family members." No family segregation data are available. Therefore, this criterion is not applied at Not Applied strength.
BP1 (Not Applied)
According to standard ACMG guidelines, the rule for BP1 is: "Supporting Strength: missense variant in a gene where only loss-of-function causes disease." This is an intronic variant. Therefore, this criterion is not applied at Not Applied strength.
BP2 (Not Applied)
According to VCEP guidelines, the rule for BP2 is: "Supporting Strength: observation in trans with P/LP variant or in cis with multiple P/LP variants." No such observations exist. Therefore, this criterion is not applied at Not Applied strength.
BP3 (Not Applied)
According to standard ACMG guidelines, the rule for BP3 is: "Supporting Strength: in-frame indel in repetitive region without known function." This is an intronic duplication, not an in-frame coding indel. Therefore, this criterion is not applied at Not Applied strength.
BP4 (Supporting)
According to VCEP guidelines, the rule for BP4 is: "Supporting Strength: for intronic/non-coding variants, concordance of MaxEntScan and SpliceAI predicting no splicing effects." The evidence for this variant shows SpliceAI score of 0.01 and no MaxEntScan predicted impact. Therefore, this criterion is applied at Supporting strength because computational evidence indicates no impact on splicing.
BP5 (Not Applied)
According to standard ACMG guidelines, the rule for BP5 is: "Supporting Strength: variant found in a case with an alternate molecular basis for disease." No such case data are available. Therefore, this criterion is not applied at Not Applied strength.
BP6 (Supporting)
According to standard ACMG guidelines, the rule for BP6 is: "Supporting Strength: reputable source reports variant as benign without available evidence." The evidence for this variant shows ClinVar reports it as benign from 6 laboratories. Therefore, this criterion is applied at Supporting strength because a reputable database classifies it as benign.
BP7 (Supporting)
According to VCEP guidelines, the rule for BP7 is: "Supporting Strength: silent or intronic variant at or beyond +7 to -21 positions if BP4 is met." The evidence for this variant shows it is at positions +7/+8 and BP4 is met. Therefore, this criterion is applied at Supporting strength because it meets the intronic position and computational benign criteria.