Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_006218.2 | Alternative | 3724 nt | 158–3364 |
| NM_006218.3 | Alternative | 9104 nt | 158–3364 |
| NM_006218.4 | MANE Select | 9259 nt | 324–3530 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Open""
COSMIC Somatic Evidence
Open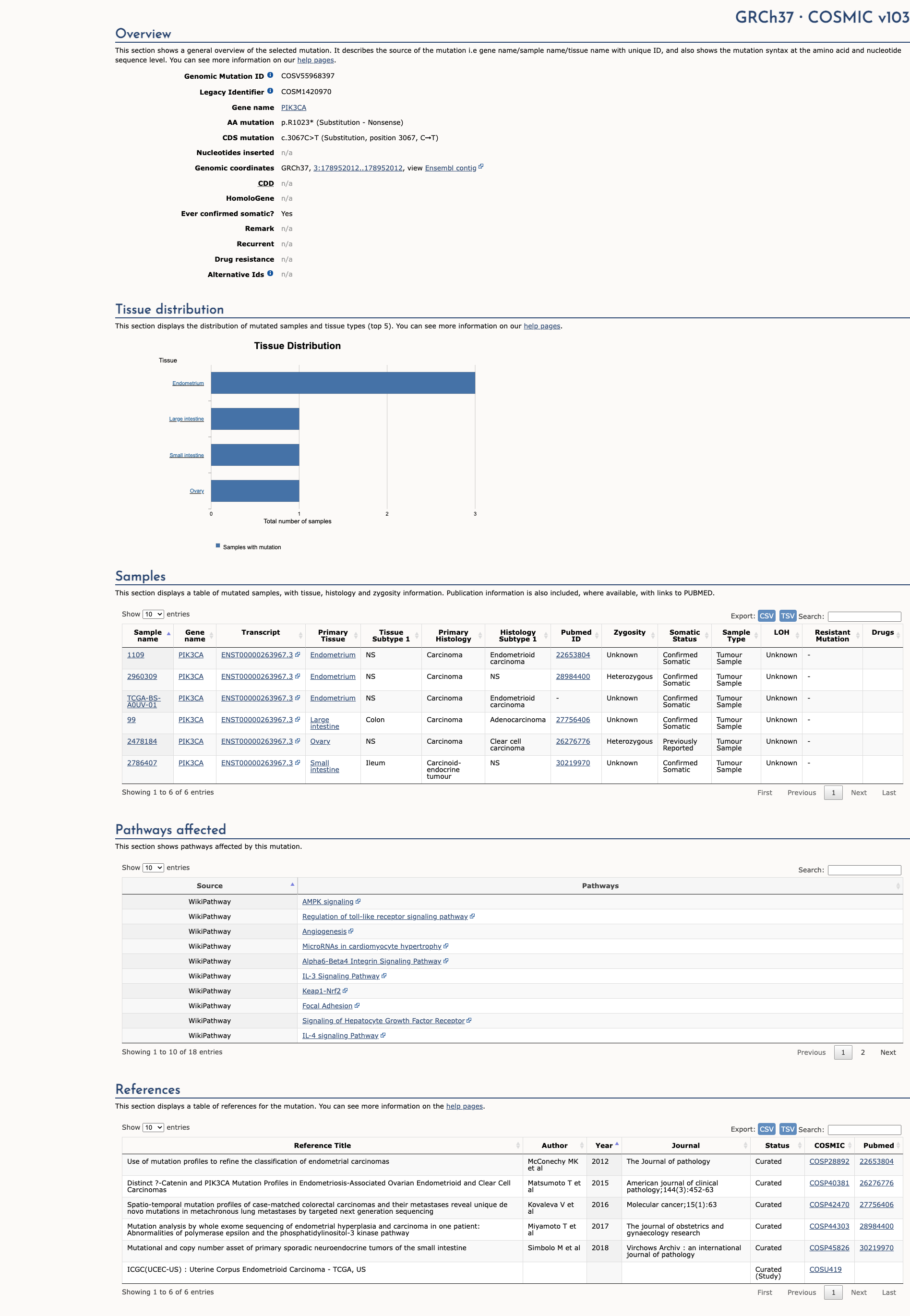
Functional Impact & Domains
Functional Domain
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.21 | -130 bp |
| Donor Loss (DL) | 0.0 | -272 bp |
| Acceptor Gain (AG) | 0.0 | -234 bp |
| Donor Gain (DG) | 0.0 | 398 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to standard ACMG guidelines, the rule for PVS1 is: "Null variant (nonsense, frameshift, canonical ±1 or 2 splice sites, initiation codon, single or multi-exon deletion) in a gene where loss-of-function is a known mechanism of disease"; the evidence for this variant shows that PIK3CA is a proto-oncogene in which gain-of-function, not loss-of-function, is the established disease mechanism; therefore, this criterion is not applied because loss-of-function is not a known mechanism for PIK3CA-related disease.
PS1 (Not Applied)
According to VCEP guidelines, the rule for PS1 is: "Strong Strength: Strong No change. Modification Type: None"; the evidence for this variant shows a nonsense change rather than a missense change identical to a known pathogenic variant; therefore, this criterion is not applied.
PS2 (Not Applied)
According to VCEP guidelines, the rule for PS2 is: "Strong Strength: Strong Award the PS2_Strong point if Criteria 1 AND Criteria 2 are fulfilled..."; no de novo or mosaic segregation data are available for this somatic variant; therefore, this criterion is not applied.
PS3 (Not Applied)
According to VCEP guidelines, the rule for PS3 is: "Strong Strength: Strong Follow recommendations set forth by the SVI... Award PS3 if the functional assay meets the acceptability criteria..."; no functional assay data are available for this variant; therefore, this criterion is not applied.
PS4 (Not Applied)
According to VCEP guidelines, the rule for PS4 is: "Very Strong Strength: Very Strong Points are assigned for phenotype according to (Table 2A)... PS4_VeryStrong ≥ 16 points"; there are no reported case counts or phenotypic data for this variant; therefore, this criterion is not applied.
PM1 (Not Applied)
According to VCEP guidelines, the rule for PM1 is: "Supporting Strength: Supporting Residues affecting critical functional domains provided in Table 4 for each gene"; this nonsense variant does not map to a defined critical domain residue in PIK3CA; therefore, this criterion is not applied.
PM2 (Supporting)
According to VCEP guidelines, the rule for PM2 is: "Supporting Strength: Supporting Absent/rare from controls in an ethnically-matched cohort population sample (≥1)"; the evidence for this variant shows it is absent from gnomAD and other population databases; therefore, this criterion is applied at Supporting strength because the variant is absent from controls.
PM3 (Not Applied)
According to standard ACMG guidelines, the rule for PM3 is: "For recessive disorders detected in trans with a pathogenic variant"; PIK3CA-related disease is not recessive and no trans data exist; therefore, this criterion is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, the rule for PM4 is: "Protein length changes due to in-frame deletions/insertions OR stop-loss variants"; this variant is a nonsense change and does not produce an in-frame indel or stop-loss; therefore, this criterion is not applied.
PM5 (Not Applied)
According to standard ACMG guidelines, the rule for PM5 is: "Novel missense change at an amino acid residue where a different missense change is pathogenic"; this variant is nonsense, not missense, and no other truncating variant at this residue is reported; therefore, this criterion is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines, the rule for PM6 is: "Assumed de novo, but without confirmation of paternity and maternity"; there are no de novo data for this somatic variant; therefore, this criterion is not applied.
PP1 (Not Applied)
According to standard ACMG guidelines, the rule for PP1 is: "Co-segregation with disease in multiple affected family members"; no segregation data are available; therefore, this criterion is not applied.
PP2 (Not Applied)
According to VCEP guidelines, the rule for PP2 is: "Supporting Strength: Supporting Missense constraint computed in ExAC/gnomAD (z-score > 3.09)"; this variant is nonsense, not missense; therefore, this criterion is not applied.
PP3 (Not Applied)
According to standard ACMG guidelines, the rule for PP3 is: "Multiple lines of computational evidence support a deleterious effect on the gene or gene product"; in silico predictions are mixed and do not strongly support pathogenicity; therefore, this criterion is not applied.
PP4 (Not Applied)
According to standard ACMG guidelines, the rule for PP4 is: "Patient’s phenotype or family history is highly specific for a disease with a single genetic etiology"; no phenotype data are provided; therefore, this criterion is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, the rule for PP5 is: "Reputable source recently reports variant as pathogenic"; this variant is not found in ClinVar or other databases; therefore, this criterion is not applied.
BA1 (Not Applied)
According to VCEP guidelines, the rule for BA1 is: "Stand Alone Strength: Stand Alone Allele frequency (>0.0926%)"; this variant is absent from population databases; therefore, this criterion is not applied.
BS1 (Not Applied)
According to VCEP guidelines, the rule for BS1 is: "Strong Strength: Strong Allele frequency (>0.0185%)"; the variant is absent from controls; therefore, this criterion is not applied.
BS2 (Not Applied)
According to VCEP guidelines, the rule for BS2 is: "Strong Strength: Strong Award BS2 if ≥3 homozygotes present in gnomAD or ≥3 heterozygous in well phenotyped family members"; no such observations exist; therefore, this criterion is not applied.
BS3 (Not Applied)
According to VCEP guidelines, the rule for BS3 is: "Strong Strength: Strong Follow recommendations set forth by the SVI..."; no functional studies demonstrating no impact are available; therefore, this criterion is not applied.
BS4 (Not Applied)
According to standard ACMG guidelines, the rule for BS4 is: "Lack of segregation in affected members of a family"; no segregation data are available; therefore, this criterion is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, the rule for BP1 is: "Missense variant in a gene for which primarily truncating variants cause disease"; this variant is truncating but PIK3CA disease mechanism is gain-of-function missense; therefore, this criterion is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, the rule for BP2 is: "Observed in trans or cis with a pathogenic variant"; no such observations exist; therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, the rule for BP3 is: "In-frame deletions/insertions in a repetitive region without a known function"; this is a nonsense variant, not an in-frame indel; therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP guidelines, the rule for BP4 is: "Supporting Strength: Supporting Award BP4 for synonymous, intronic positions... if two of three splicing prediction tools predict no impact"; this is a nonsense variant, not applicable; therefore, this criterion is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, the rule for BP5 is: "Variant found in a gene for which an alternative molecular basis is known to cause disease"; not applicable here; therefore, this criterion is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, the rule for BP6 is: "Reputable source reports variant as benign"; no such reports exist; therefore, this criterion is not applied.
BP7 (Not Applied)
According to VCEP guidelines, the rule for BP7 is: "Supporting Strength: Supporting For synonymous, intronic positions... if the nucleotide is non-conserved"; this is a nonsense variant; therefore, this criterion is not applied.