Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_033360.2 | Alternative | 5436 nt | 182–751 |
| NM_033360.4 | Alternative | 5430 nt | 191–760 |
| NM_033360.3 | Alternative | 5889 nt | 193–762 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Open"Present in ClinVar, however no clinical evidence available for this variant."
COSMIC Somatic Evidence
Open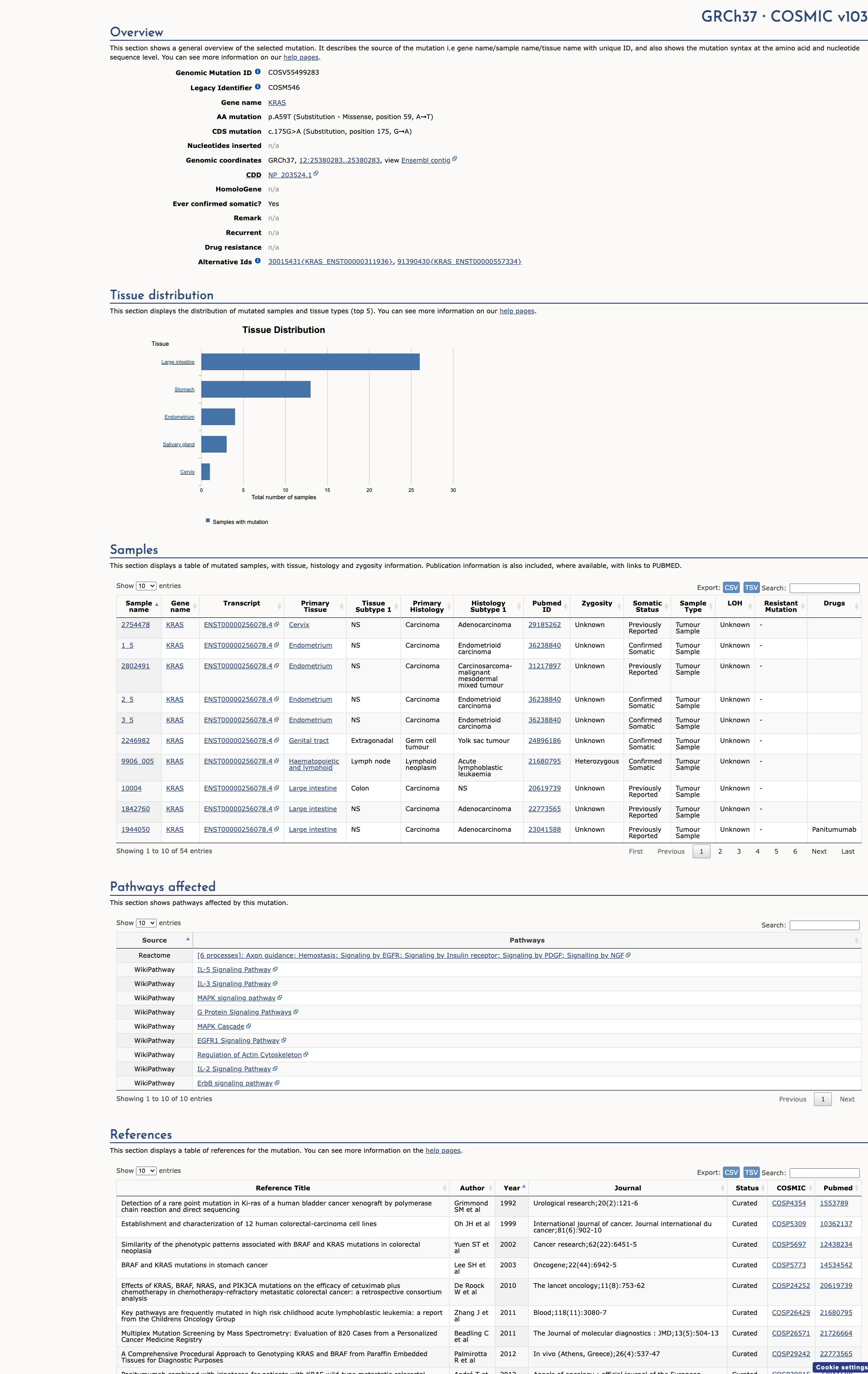
Functional Impact & Domains
Functional Domain
The KRAS A59T variant has been functionally characterized as a gain-of-function mutation. Experimental evidence demonstrates that this variant is oncogenic, leading to increased cell proliferation and migration in NIH3T3 cells, as well as enhanced MAPK signaling in Ba/F3 cells compared to the wildtype. These findings support a damaging effect of the KRAS A59T mutation.
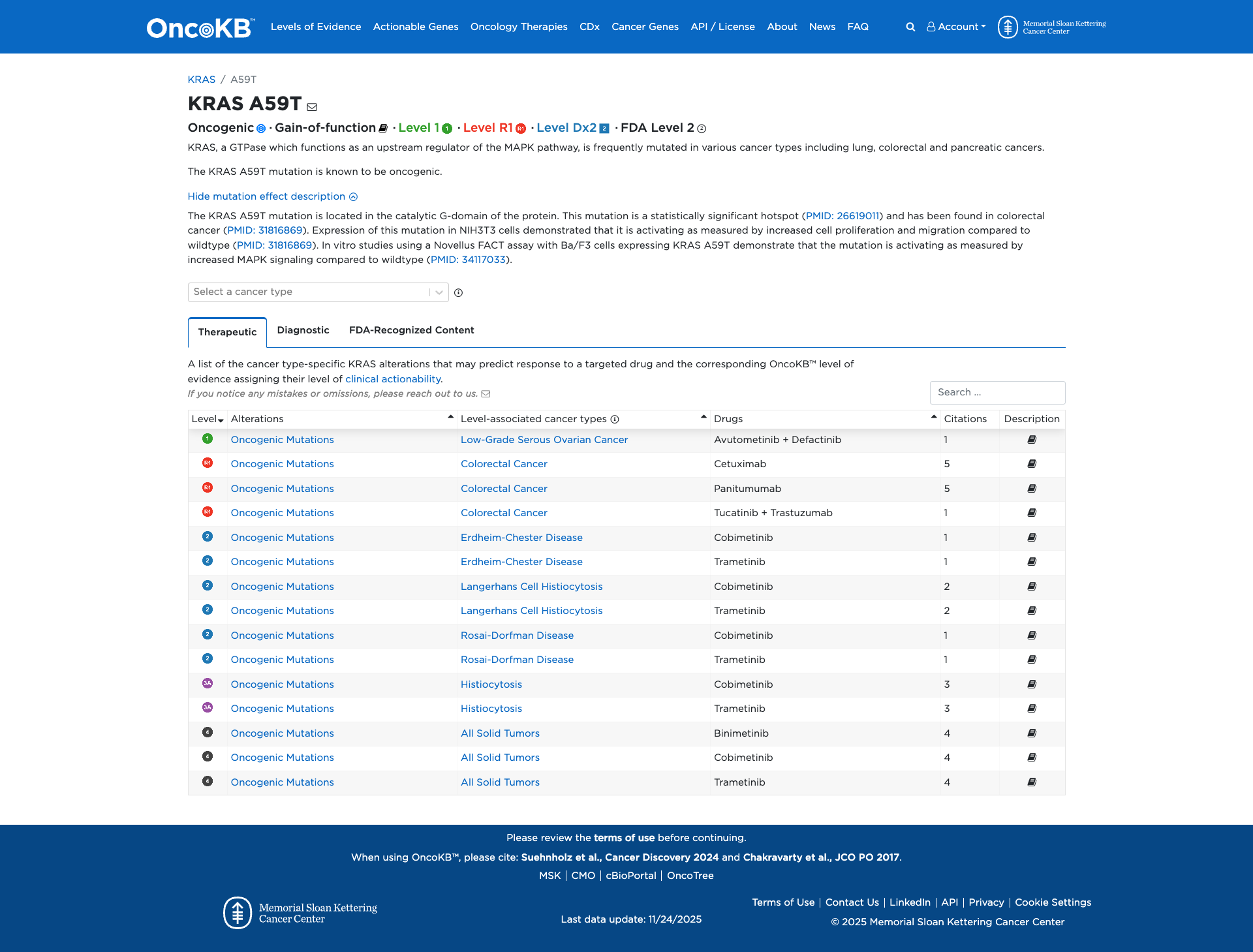
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -1 bp |
| Donor Loss (DL) | 0.0 | -65 bp |
| Acceptor Gain (AG) | 0.01 | -4 bp |
| Donor Gain (DG) | 0.07 | -3 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, PVS1 addresses null variants in genes where loss-of-function is a known mechanism. KRAS-associated disease mechanism is gain-of-function and this variant is a missense change. The evidence shows a missense alteration (A59T). Therefore, this criterion is not applied.
PS1 (Not Applied)
According to VCEP guidelines, PS1 (Strong) requires the same amino acid change as a previously established pathogenic variant. The evidence shows no identical A59T nucleotide change previously reported as pathogenic. Therefore, this criterion is not applied.
PS2 (Not Applied)
According to standard ACMG guidelines, PS2 (Strong) applies to de novo occurrences with confirmed parentage. No de novo data are available for this variant. Therefore, this criterion is not applied.
PS3 (Moderate)
According to VCEP guidelines, PS3 (Moderate) is met by two or more different approved functional assays. The evidence shows oncogenic gain-of-function demonstrated by increased cell proliferation in NIH3T3 cells and enhanced MAPK signaling in Ba/F3 cells. Therefore, this criterion is applied at Moderate strength.
PS4 (Not Applied)
According to VCEP guidelines, PS4 (Strong) requires ≥5 proband points. No case-level data or point counts are available. Therefore, this criterion is not applied.
PM1 (Moderate)
According to VCEP guidelines, PM1 (Moderate) applies to variants in the SW2 region (AA 57–64). A59T lies within SW2. Therefore, this criterion is applied at Moderate strength.
PM2 (Supporting)
According to VCEP guidelines, PM2 (Supporting) applies when the variant is absent from gnomAD. The evidence shows A59T is not present in gnomAD. Therefore, this criterion is applied at Supporting strength.
PM3 (Not Applied)
According to standard ACMG guidelines, PM3 (Moderate) applies to recessive disorders with detected trans variants. KRAS A59T is associated with a dominant oncogenic mechanism, and no trans data are available. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, PM4 (Moderate) applies to in-frame insertions/deletions or stop-loss variants. A59T is a missense change. Therefore, this criterion is not applied.
PM5 (Moderate)
According to VCEP guidelines, PM5 (Moderate) applies when one other pathogenic residue change at the same codon is reported. A59G has been established as pathogenic. Therefore, this criterion is applied at Moderate strength.
PM6 (Not Applied)
According to standard ACMG guidelines, PM6 (Supporting) applies to assumed de novo variants without confirmation. No de novo evidence is available. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to standard ACMG guidelines, PP1 (Supporting) requires segregation data. No family segregation data are available. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, PP2 (Supporting) applies when a gene has a low rate of benign missense variation. No specific constraint metrics or gene-level data were provided. Therefore, this criterion is not applied.
PP3 (Supporting)
According to VCEP guidelines, PP3 (Supporting) applies for missense variants with REVEL ≥0.7. The evidence shows a REVEL score of 0.87. Therefore, this criterion is applied at Supporting strength.
PP4 (Not Applied)
According to standard ACMG guidelines, PP4 (Supporting) requires a highly specific phenotype. No phenotype data are provided. Therefore, this criterion is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, PP5 (Supporting) applies when reputable sources report the variant as pathogenic without evidence. ClinVar has no assertion for this variant. Therefore, this criterion is not applied.
BA1 (Not Applied)
According to standard ACMG guidelines, BA1 (Stand Alone) requires population allele frequency ≥5%. The variant is absent from controls. Therefore, this criterion is not applied.
BS1 (Not Applied)
According to standard ACMG guidelines, BS1 (Strong) requires allele frequency ≥2.5%. The variant is absent from controls. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to standard ACMG guidelines, BS2 (Strong/Supporting) applies when observed in multiple healthy individuals. No such data are available. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to standard ACMG guidelines, BS3 (Strong) applies when well-established functional studies show no damaging effect. Functional studies for A59T demonstrate gain-of-function. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to standard ACMG guidelines, BS4 (Strong) requires non-segregation in affected members. No segregation data are available. Therefore, this criterion is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, BP1 (Supporting) applies to truncating variants in genes not associated with loss-of-function. A59T is missense. Therefore, this criterion is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, BP2 (Supporting) applies to observation in trans with a pathogenic variant in dominant disorders. No such data are available. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, BP3 (Supporting) applies to in-frame indels in repetitive regions. A59T is a missense change. Therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP guidelines, BP4 (Supporting) applies for missense variants with REVEL ≤0.3. The REVEL score is 0.87, well above threshold. Therefore, this criterion is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, BP5 (Supporting) applies when a variant is found in a case with an alternate molecular cause. No alternate cause data are available. Therefore, this criterion is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, BP6 (Supporting) applies when reputable sources report as benign without evidence. No such reports exist. Therefore, this criterion is not applied.
BP7 (Not Applied)
According to standard ACMG guidelines, BP7 (Supporting) applies to synonymous changes with no predicted splice impact. A59T is missense. Therefore, this criterion is not applied.