Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000314.8 | MANE Select | 8515 nt | 846–2057 |
| NM_000314.7 | RefSeq Select | 8514 nt | 845–2056 |
| NM_000314.5 | Alternative | 8719 nt | 1032–2243 |
| NM_000314.4 | Alternative | 5572 nt | 1032–2243 |
| NM_000314.3 | Alternative | 3416 nt | 1032–2243 |
| NM_000314.6 | Alternative | 8718 nt | 1032–2243 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Open"This variant has been reported in ClinVar as Likely pathogenic (1 clinical laboratories)."
COSMIC Somatic Evidence
Open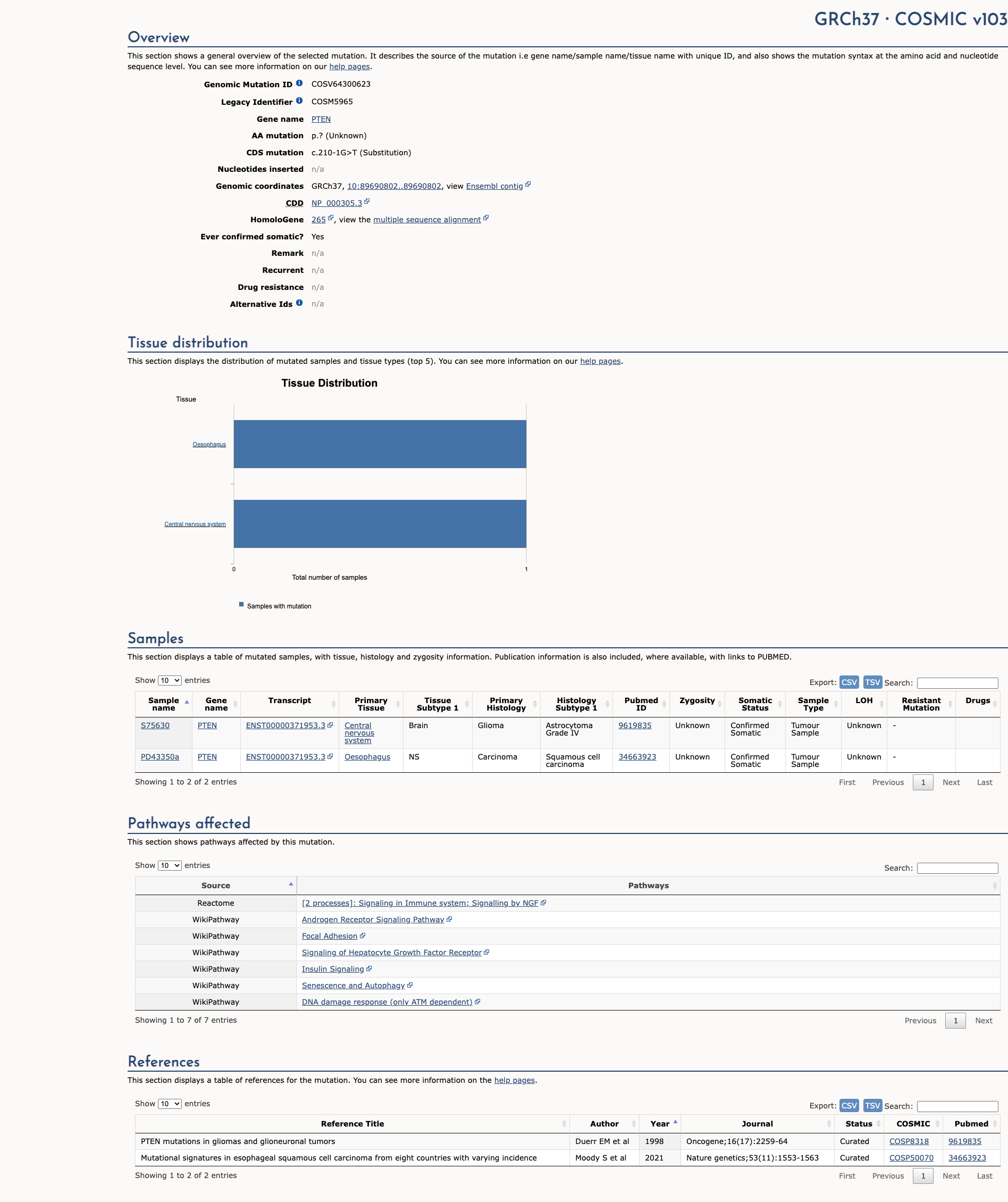
Functional Impact & Domains
Functional Domain
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.99 | 1 bp |
| Donor Loss (DL) | 0.66 | 44 bp |
| Acceptor Gain (AG) | 0.47 | 13 bp |
| Donor Gain (DG) | 0.0 | 404 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Very Strong)
According to VCEP guidelines, the rule for PVS1 is: "Null Variant (nonsense, fs, start codon, splicing +1/2 ...) in a gene where LOF is a known mechanism of disease". The evidence for this variant shows: c.210-1G>T affects the canonical splice acceptor site predicted to abolish normal splicing, resulting in loss of function of PTEN. Therefore, this criterion is applied at Very Strong strength because it is a canonical splice site variant in a LOF‐sensitive gene.
PS1 (Not Applied)
According to VCEP guidelines, the rule for PS1 is: "Same amino acid change as a previously established pathogenic variant regardless of nucleotide change OR different variant at same nucleotide position as a pathogenic splicing variant...". The evidence for this variant shows: no known pathogenic splicing variant reported at this exact nucleotide. Therefore, this criterion is not applied.
PS2 (Not Applied)
According to VCEP guidelines, the rule for PS2 is: "De novo (both maternity and paternity confirmed) observation in a patient with the disease and no family history". The evidence for this variant shows: no de novo inheritance data available. Therefore, this criterion is not applied.
PS3 (Not Applied)
According to PTEN Pre-processing, the finding for PS3 is: "The variant has not been functionally characterized." The evidence for this variant shows: no RNA, mini-gene, or in vitro functional studies available. Therefore, this criterion is not applied.
PS4 (Not Applied)
According to VCEP guidelines, the rule for PS4 is: "Increased prevalence of variant in affected individuals compared with controls or probands specificity score ≥4". The evidence for this variant shows: no case series, no specificity scores reported. Therefore, this criterion is not applied.
PM1 (Not Applied)
According to VCEP guidelines, the rule for PM1 is: "Located in a mutational hot spot and/or critical and well‐established functional domain (residues 90–94, 123–130, 166–168)". The evidence for this variant shows: position c.210-1 is outside of the defined catalytic motifs. Therefore, this criterion is not applied.
PM2 (Supporting)
According to VCEP guidelines, the rule for PM2 is: "Absent in population Databases present at <0.00001 allele frequency in gnomAD". The evidence for this variant shows: MAF = 0% in gnomAD. Therefore, this criterion is applied at Supporting strength because the variant is absent from large population databases.
PM3 (Not Applied)
According to standard ACMG guidelines, the rule for PM3 is: "For recessive disorders, detected in trans with a pathogenic variant". The evidence for this variant shows: PTEN is autosomal dominant and no trans data. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to VCEP guidelines, the rule for PM4 is: "Protein length changes due to in‐frame indels or stop‐loss variants". The evidence for this variant shows: it is a splice site variant, not an in‐frame indel or stop‐loss. Therefore, this criterion is not applied.
PM5 (Not Applied)
According to VCEP guidelines, the rule for PM5 is: "Missense change at a residue where a different missense change is known pathogenic". The evidence for this variant shows: it is a splice variant, not missense. Therefore, this criterion is not applied.
PM6 (Not Applied)
According to VCEP guidelines, the rule for PM6 is: "Assumed de novo without confirmation of paternity/maternity". The evidence for this variant shows: no de novo inheritance information. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to VCEP guidelines, the rule for PP1 is: "Co‐segregation with disease in multiple affected family members". The evidence for this variant shows: no segregation data reported. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, the rule for PP2 is: "Missense variant in a gene with low rate of benign missense variation and where missense is a common mechanism". The evidence for this variant shows: it is a splice site variant. Therefore, this criterion is not applied.
PP3 (Not Applied)
According to VCEP guidelines, the rule for PP3 is: "Multiple lines of computational evidence support a deleterious effect – for splicing variants: concordance of SpliceAI and VarSeak". The evidence for this variant shows: only SpliceAI predicts impact; VarSeak data unavailable. Therefore, this criterion is not applied due to lack of concordant in silico tools.
PP4 (Not Applied)
According to standard ACMG guidelines, the rule for PP4 is: "Patient’s phenotype is highly specific for a disease with single genetic etiology". The evidence for this variant shows: no phenotype data provided. Therefore, this criterion is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, the rule for PP5 is: "Reputable source recently reports variant as pathogenic but evidence not available". The evidence for this variant shows: ClinVar entry exists but evidence not accessible; however, ClinGen discourages PP5. Therefore, this criterion is not applied.
BA1 (Not Applied)
According to VCEP guidelines, the rule for BA1 is: "gnomAD filtering allele frequency >0.00056". The evidence for this variant shows: allele frequency = 0%. Therefore, this criterion is not applied.
BS1 (Not Applied)
According to VCEP guidelines, the rule for BS1 is: "Allele frequency 0.000043–0.00056 in gnomAD". The evidence for this variant shows: allele frequency = 0%. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to VCEP guidelines, the rule for BS2 is: "Observed homozygous in healthy individual". The evidence for this variant shows: no homozygous observations. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to VCEP guidelines, the rule for BS3 is: "Well‐established functional studies show no damaging effect". The evidence for this variant shows: no functional studies. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to VCEP guidelines, the rule for BS4 is: "Lack of segregation in affected members". The evidence for this variant shows: no segregation data. Therefore, this criterion is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, the rule for BP1 is: "Missense variant in a gene where only truncating variants cause disease". The evidence for this variant shows: it is a splice variant. Therefore, this criterion is not applied.
BP2 (Not Applied)
According to VCEP guidelines, the rule for BP2 is: "Observed in trans with a pathogenic variant". The evidence for this variant shows: no evidence of cis/trans observations. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, the rule for BP3 is: "In-frame indels in repetitive region". The evidence for this variant shows: splice site variant, not in repeat. Therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP guidelines, the rule for BP4 is: "Multiple lines of computational evidence suggest no impact". The evidence for this variant shows: conflicting predictions. Therefore, this criterion is not applied.
BP5 (Not Applied)
According to VCEP guidelines, the rule for BP5 is: "Variant found in case with alternative molecular basis". The evidence for this variant shows: no alternate diagnosis. Therefore, this criterion is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, the rule for BP6 is: "Reputable source reports variant as benign but evidence not available". The evidence for this variant shows: no benign classifications. Therefore, this criterion is not applied.
BP7 (Not Applied)
According to VCEP guidelines, the rule for BP7 is: "Synonymous or intronic variant at or beyond +7/-21 with no splicing impact". The evidence for this variant shows: position at -1, within the splice consensus. Therefore, this criterion is not applied.