Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000314.8 | MANE Select | 8515 nt | 846–2057 |
| NM_000314.7 | RefSeq Select | 8514 nt | 845–2056 |
| NM_000314.5 | Alternative | 8719 nt | 1032–2243 |
| NM_000314.4 | Alternative | 5572 nt | 1032–2243 |
| NM_000314.3 | Alternative | 3416 nt | 1032–2243 |
| NM_000314.6 | Alternative | 8718 nt | 1032–2243 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis sequence change affects an acceptor splice site in intron 6 of the PTEN gene. It is expected to disrupt RNA splicing. Variants that disrupt the donor or acceptor splice site typically lead to a loss of protein function (PMID: 16199547), and loss-of-function variants in PTEN are known to be pathogenic (PMID: 9467011, 21194675). This variant is not present in population databases (gnomAD no frequency). Disruption of this splice site has been observed in individual(s) with PTEN-related conditions (PMID: 19265751, 21659347, 24379037). In at least one individual the variant was observed to be de novo. ClinVar contains an entry for this variant (Variation ID: 427598). Algorithms developed to predict the effect of sequence changes on RNA splicing suggest that this variant may disrupt the consensus splice site. For these reasons, this variant has been classified as Pathogenic.
The c.635-1G>A intronic variant results from a G to A substitution one nucleotide upstream from coding exon 7 of the PTEN gene. This variant was identified in 1 of 802 individuals with features of Cowden syndrome or Bannayan-Riley-Ruvalcaba syndrome undergoing PTEN analysis (Pilarski R et al. J Med Genet, 2011 Aug;48:505-12). This variant was reported in an individual who met clinical criteria for PTEN hamartoma tumor syndrome (Varga EA et al. Genet Med, 2009 Feb;11:111-7; Hansen-Kiss E et al. J Med Genet, 2017 Jul;54:471-478). This variant is considered to be rare based on population cohorts in the Genome Aggregation Database (gnomAD). This nucleotide position is highly conserved in available vertebrate species. In silico splice site analysis predicts that this alteration will weaken the native splice acceptor site and may result in the creation or strengthening of a novel splice acceptor site. In addition to the clinical data presented in the literature, alterations that disrupt the canonical splice site are expected to cause aberrant splicing, resulting in an abnormal protein or a transcript that is subject to nonsense-mediated mRNA decay. As such, this alteration is classified as likely pathogenic.
"This variant has been reported in ClinVar as Pathogenic (3 clinical laboratories) and as Likely pathogenic (2 clinical laboratories)."
COSMIC Somatic Evidence
Open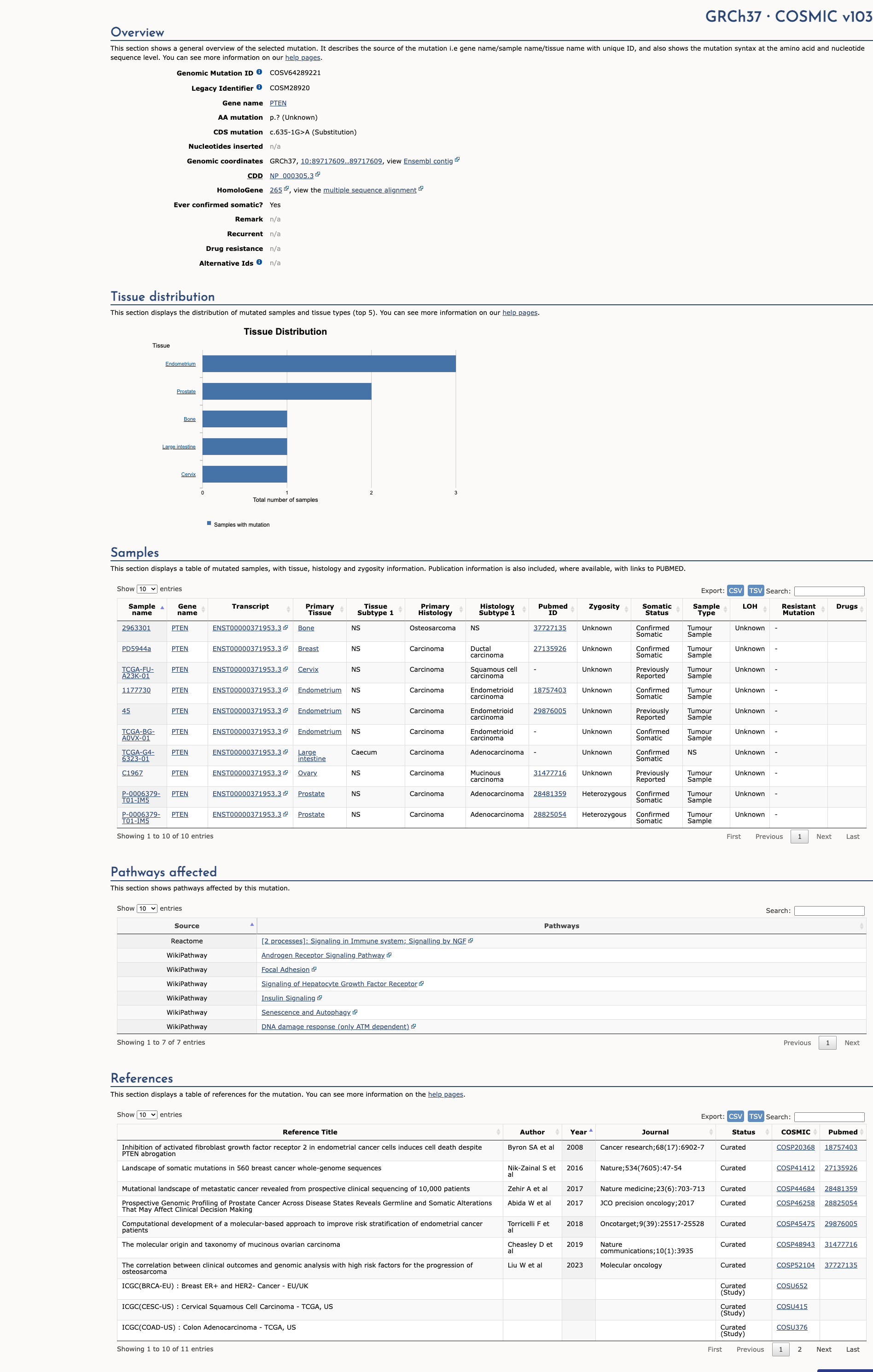
Functional Impact & Domains
Functional Domain
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.98 | 1 bp |
| Donor Loss (DL) | 0.01 | -97 bp |
| Acceptor Gain (AG) | 0.51 | 55 bp |
| Donor Gain (DG) | 0.07 | -20 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Very Strong)
According to VCEP guidelines for PTEN: "Very Strong Strength: Use PTEN PVS1 decision tree." The variant NM_000314.8:c.635-1G>A is a canonical splice acceptor change predicted to result in loss of function in PTEN, where loss of function is a known mechanism of disease. Therefore, this criterion is applied at Very Strong strength because it meets the PTEN-specific PVS1 decision tree for an essential splice site variant.
PS1 (Not Applied)
According to VCEP guidelines: "Different variant at same nucleotide position as a pathogenic splicing variant..." There is no previously established pathogenic splicing variant at this exact nucleotide position. Therefore, this criterion is not applied.
PS2 (Not Applied)
According to VCEP guidelines: de novo occurrence requires confirmed maternity and paternity. No de novo data are available for this variant. Therefore, this criterion is not applied.
PS3 (Not Applied)
According to PTEN pre-processing: variant not found in PTEN functional data file, and no in vitro or in vivo functional studies (e.g., splicing assays) are available. Therefore, this criterion is not applied.
PS4 (Not Applied)
According to VCEP guidelines: case-level or case-control data (specificity score) required. No case reports or case-control frequency data are available. Therefore, this criterion is not applied.
PM1 (Not Applied)
According to VCEP guidelines: location in critical functional domain (residues 90-94, 123-130, 166-168). This is a splice site variant, not a missense in those motifs. Therefore, this criterion is not applied.
PM2 (Supporting)
According to VCEP guidelines: "Supporting: Absent in population databases present at <0.00001 allele frequency in gnomAD." The variant is absent from gnomAD and other population databases. Therefore, this criterion is applied at Supporting strength because the allele frequency is below the threshold.
PM3 (Not Applied)
According to standard ACMG guidelines: PM3 requires observation in trans with a pathogenic variant in a recessive gene. No trans-phase data are available. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to VCEP guidelines: in-frame insertions/deletions or stop-loss variants required. This is a splice site variant. Therefore, this criterion is not applied.
PM5 (Not Applied)
According to VCEP guidelines: missense change at a residue with a different known pathogenic missense. This is a splice variant. Therefore, this criterion is not applied.
PM6 (Not Applied)
According to VCEP guidelines: presumed de novo without confirmation. No such data are available. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to VCEP guidelines: segregation data across multiple affected family members required. No segregation information is available. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines: missense variant in gene with low benign missense rate. This is a splice variant, not missense. Therefore, this criterion is not applied.
PP3 (Supporting)
According to VCEP guidelines: "Supporting: Multiple lines of computational evidence support a deleterious effect on the gene or gene product. Splicing variants: Concordance of SpliceAl and VarSeak." SpliceAI predicts an acceptor loss with a score of 0.98, indicating high impact on splicing. Therefore, this criterion is applied at Supporting strength because computational tools concordantly predict deleterious impact.
PP4 (Not Applied)
According to standard ACMG guidelines: specific phenotype with single genetic etiology required. No detailed phenotype or family history matching PTEN-related syndromes provided. Therefore, this criterion is not applied.
PP5 (Supporting)
According to standard ACMG guidelines: "Supporting: Reputable source recently reports variant as pathogenic, but evidence not available for independent evaluation." ClinVar reports this variant as Pathogenic by three laboratories and Likely Pathogenic by two. Therefore, this criterion is applied at Supporting strength.
BA1 (Not Applied)
According to VCEP guidelines: filtering allele frequency >0.00056 required. The variant is absent. Therefore, this criterion is not applied.
BS1 (Not Applied)
According to VCEP guidelines: allele frequency 0.000043–0.00056 required. The variant is absent. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to VCEP guidelines: observed homozygous in unaffected individual required. No such observations. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to VCEP guidelines: well-established functional studies show no damaging effect. No functional studies demonstrate no impact. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to VCEP guidelines: lack of segregation evidence in affected members of ≥2 families required. No segregation data available. Therefore, this criterion is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines: missense in gene where only truncating variants are pathogenic. This is a splice variant. Therefore, this criterion is not applied.
BP2 (Not Applied)
According to VCEP guidelines: observed in trans or in cis with other pathogenic variant required. No such data. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines: in-frame indel in repetitive region. This is a splice site. Therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP guidelines: computational evidence suggests no impact. Here, computational evidence predicts high impact on splicing. Therefore, this criterion is not applied.
BP5 (Not Applied)
According to VCEP guidelines: variant found in case with alternate molecular basis. No evidence of alternate etiology. Therefore, this criterion is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines: reputable source reports benign. No such reports. Therefore, this criterion is not applied.
BP7 (Not Applied)
According to VCEP guidelines: synonymous or intronic variant beyond +7/-21 with no splicing impact. This is a -1 splice site. Therefore, this criterion is not applied.