Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_024675.4 | MANE Select | 4008 nt | 154–3714 |
| NM_024675.3 | RefSeq Select | 4069 nt | 201–3761 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenCurators: Marc Tischkowitz, Arleen D. Auerbach. Submitter to LOVD: Yukihide Momozawa.
"This variant has been reported in ClinVar as Likely benign (7 clinical laboratories) and as Benign (1 clinical laboratories)."
COSMIC Somatic Evidence
Open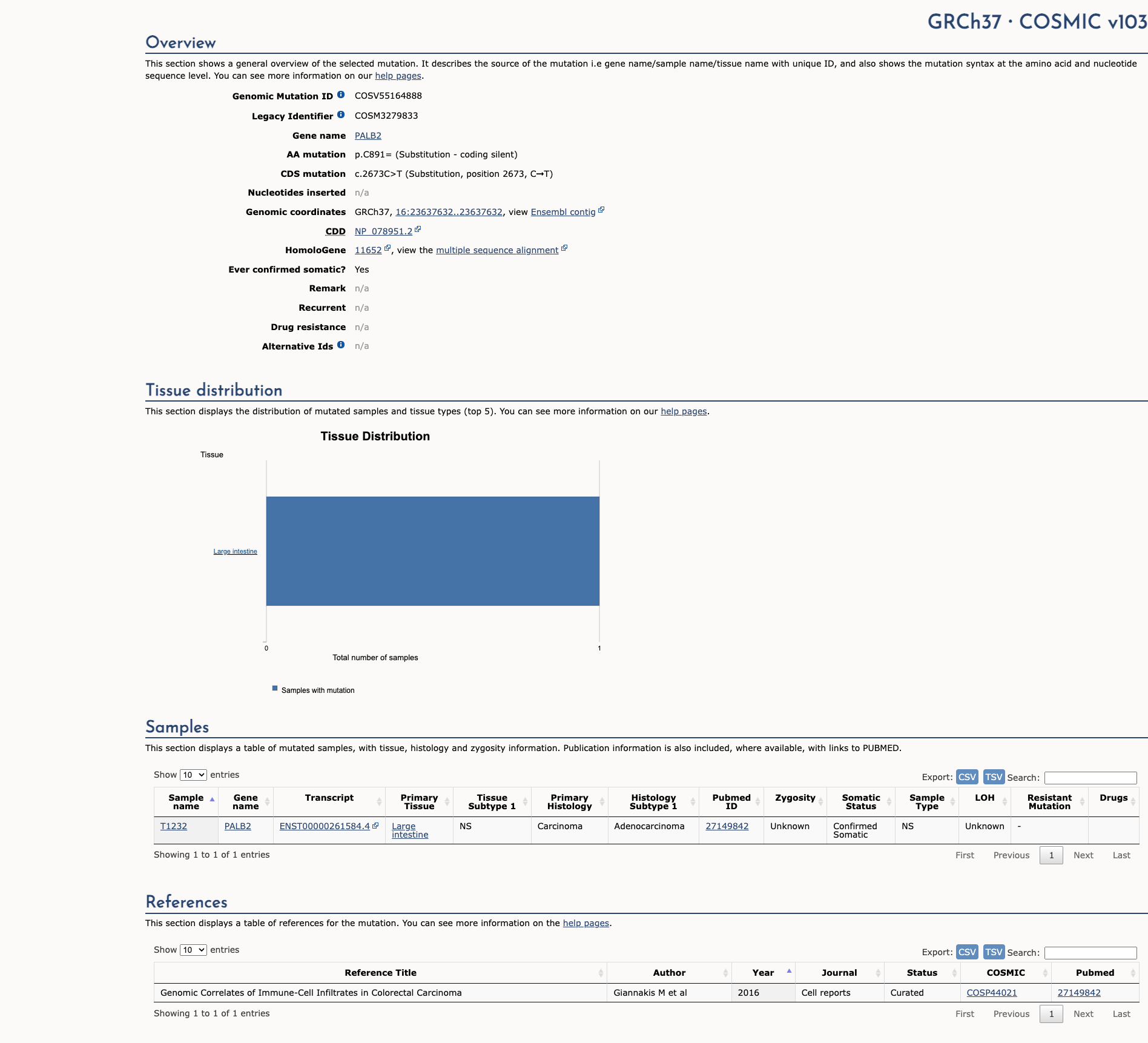
Functional Impact & Domains
Functional Domain
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.06 | 86 bp |
| Donor Loss (DL) | 0.04 | -75 bp |
| Acceptor Gain (AG) | 0.0 | -26 bp |
| Donor Gain (DG) | 0.0 | 60 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, the rule for PVS1 is: "Use PALB2 PVS1 Decision Tree Modification Type: Gene-specific; very strong strength for null variants." The evidence for this variant shows: it is a synonymous (p.C891=) change not predicted to impact splicing. Therefore, this criterion is not applied because the variant is not a null (loss-of-function) allele.
PS1 (Not Applied)
According to VCEP PS1 guidelines, the rule for PS1 is: "Use PALB2 PS1 Splicing table; strong for same amino acid change via different nucleotide when pathogenic." The evidence for this variant shows: no known pathogenic variant alters the same codon to the same amino acid, and this is a synonymous change. Therefore, PS1 is not applied.
PS2 (Not Applied)
According to standard ACMG guidelines, the rule for PS2 is: "De novo (both maternity and paternity confirmed) in a patient with the disease and no family history." The evidence for this variant shows: no de novo data are available. Therefore, PS2 is not applied.
PS3 (Not Applied)
According to standard ACMG guidelines, the rule for PS3 is: "Well-established in vitro or in vivo functional studies supportive of a damaging effect on the gene or gene product." The evidence for this variant shows: no functional studies have been reported. Therefore, PS3 is not applied.
PS4 (Not Applied)
According to VCEP PS4 guidelines, the rule for PS4 is: "Strong if case-control studies with significant odds ratio/hazard ratio ≥3 (lower 95% CI ≥1.5)." The evidence for this variant shows: no case-control or family segregation data. Therefore, PS4 is not applied.
PM1 (Not Applied)
According to standard ACMG guidelines, the rule for PM1 is: "Located in a mutational hot spot and/or critical and well-established functional domain without benign variation." The evidence for this variant shows: it is a synonymous change outside known functional domains. Therefore, PM1 is not applied.
PM2 (Not Applied)
According to VCEP guidelines, the rule for PM2 is: "Supporting if variant absent in gnomAD or present in ≤1/300,000 alleles." The evidence for this variant shows: MAF=0.00239% (6/251,486 alleles), which exceeds the ≤1/300,000 (0.00033%) threshold. Therefore, PM2 is not applied.
PM3 (Not Applied)
According to VCEP PM3 guidelines, the rule for PM3 is: "Use Fanconi Anemia PM3 tables for recessive in trans observations." The evidence for this variant shows: no compound heterozygous or homozygous observations in affected individuals consistent with recessive inheritance. Therefore, PM3 is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, the rule for PM4 is: "Protein length changes as a result of in-frame indels or stop-loss variants." The evidence for this variant shows: it is a synonymous substitution with no length change. Therefore, PM4 is not applied.
PM5 (Not Applied)
According to VCEP PM5 guidelines, the rule for PM5 is: "Supporting for frameshifting or truncating variants upstream of p.Tyr1183." The evidence for this variant shows: it is synonymous, not truncating. Therefore, PM5 is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines, the rule for PM6 is: "Assumed de novo (without confirmation of paternity and maternity)." The evidence for this variant shows: no de novo information. Therefore, PM6 is not applied.
PP1 (Not Applied)
According to VCEP PP1 guidelines, the rule for PP1 is: "Supporting/Moderate/Strong based on segregation LOD scores in familial cases." The evidence for this variant shows: no segregation data. Therefore, PP1 is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, the rule for PP2 is: "Missense variant in a gene with low rate of benign missense variation and in which missense are a common mechanism of disease." The evidence for this variant shows: it is synonymous, not missense. Therefore, PP2 is not applied.
PP3 (Not Applied)
According to VCEP PP3 guidelines, the rule for PP3 is: "Supporting RNA: at least one predictor (e.g. SpliceAI) shows impact on splicing." The evidence for this variant shows: SpliceAI score=0.06 indicating no splice impact. Therefore, PP3 is not applied.
PP4 (Not Applied)
According to standard ACMG guidelines, the rule for PP4 is: "Patient’s phenotype or family history is highly specific for a disease with a single genetic etiology." The evidence for this variant shows: no phenotype data provided. Therefore, PP4 is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, the rule for PP5 is: "Reputable source reports variant as pathogenic but evidence not available for independent evaluation." The evidence for this variant shows: ClinVar reports benign/likely benign. Therefore, PP5 is not applied.
BA1 (Not Applied)
According to VCEP BA1 guidelines, the rule for BA1 is: "Stand alone if allele frequency >0.1%." The evidence for this variant shows: MAF=0.00239%, which is below 0.1%. Therefore, BA1 is not applied.
BS1 (Not Applied)
According to VCEP BS1 guidelines, the rule for BS1 is: "Strong if filtering AF >0.01%." The evidence for this variant shows: MAF=0.00239% (0.0000239 fraction), which is below 0.01%. Therefore, BS1 is not applied.
BS2 (Not Applied)
According to VCEP BS2 guidelines, the rule for BS2 is: "Use Fanconi Anemia BS2 tables for observations in healthy adults." The evidence for this variant shows: no such observations. Therefore, BS2 is not applied.
BS3 (Not Applied)
According to standard ACMG guidelines, the rule for BS3 is: "Well-established functional studies show no damaging effect." The evidence for this variant shows: no functional data. Therefore, BS3 is not applied.
BS4 (Not Applied)
According to VCEP BS4 guidelines, the rule for BS4 is: "Strong/Moderate/Supporting segregation data showing lack of cosegregation." The evidence for this variant shows: no segregation data. Therefore, BS4 is not applied.
BP1 (Not Applied)
According to VCEP BP1 guidelines, the rule for BP1 is: "Supporting for missense variants in genes where only LoF causes disease." The evidence for this variant shows: it is synonymous. Therefore, BP1 is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, the rule for BP2 is: "Observed in trans with a pathogenic variant for dominant disorders or in cis with a pathogenic variant." The evidence for this variant shows: no such observations. Therefore, BP2 is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, the rule for BP3 is: "In-frame deletions/insertions in repetitive regions without known function." The evidence for this variant shows: it is a single-nucleotide change, not an indel. Therefore, BP3 is not applied.
BP4 (Supporting)
According to VCEP BP4 guidelines, the rule for BP4 is: "Supporting if RNA: at least one well-established predictor (e.g. SpliceAI) shows no impact on splicing." The evidence for this variant shows: SpliceAI maximum score=0.06, indicating no splicing impact. Therefore, BP4 is applied at Supporting strength.
BP5 (Not Applied)
According to standard ACMG guidelines, the rule for BP5 is: "Variant found in a case with an alternate molecular basis for disease." The evidence for this variant shows: no such case data. Therefore, BP5 is not applied.
BP6 (Supporting)
According to standard ACMG guidelines, the rule for BP6 is: "Reputable source reports variant as benign but evidence is not available for independent evaluation." The evidence for this variant shows: ClinVar entries from eight laboratories report the variant as benign/likely benign without primary evidence. Therefore, BP6 is applied at Supporting strength.
BP7 (Not Applied)
According to VCEP BP7 guidelines, the rule for BP7 is: "Supporting for synonymous variants with observed lack of aberrant RNA defect." The evidence for this variant shows: no RNA assay data, only in silico predictions. Therefore, BP7 is not applied.