Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000051.3 | RefSeq Select | 13147 nt | 386–9556 |
| NM_000051.4 | MANE Select | 12915 nt | 151–9321 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenIn the published literature, this variant has been reported in individuals with breast cancer (PMID: 35264596 (2022)) and lung squamous cell carcinoma (PMID: 26689913 (2015)). The frequency of this variant in the general population, 0.0000066 (1/152192 chromosomes (Genome Aggregation Database, http://gnomad.broadinstitute.org)), is uninformative in the assessment of its pathogenicity. Analysis of this variant using bioinformatics tools for the prediction of the effect of amino acid changes on protein structure and function yielded predictions that this variant is damaging. Based on the available information, we are unable to determine the clinical significance of this variant.
This sequence change replaces tyrosine, which is neutral and polar, with cysteine, which is neutral and slightly polar, at codon 2627 of the ATM protein (p.Tyr2627Cys). This variant is not present in population databases (gnomAD no frequency). This missense change has been observed in individual(s) with breast cancer and/or lung squamous cell carcinoma (PMID: 26689913). ClinVar contains an entry for this variant (Variation ID: 220064). An algorithm developed to predict the effect of missense changes on protein structure and function (PolyPhen-2) suggests that this variant is likely to be disruptive. This variant disrupts the p.Tyr2627 amino acid residue in ATM. Other variant(s) that disrupt this residue have been determined to be pathogenic (PMID: 22071889). This suggests that this residue is clinically significant, and that variants that disrupt this residue are likely to be disease-causing. In summary, the currently available evidence indicates that the variant is pathogenic, but additional data are needed to prove that conclusively. Therefore, this variant has been classified as Likely Pathogenic.
The p.Y2627C variant (also known as c.7880A>G), located in coding exon 52 of the ATM gene, results from an A to G substitution at nucleotide position 7880. The tyrosine at codon 2627 is replaced by cysteine, an amino acid with highly dissimilar properties. This alteration was detected in a cohort of 1663 Brazilian breast cancer patients who underwent hereditary multigene panel testing (Guindalini RSC et al. Sci Rep, 2022 Mar;12:4190). This amino acid position is highly conserved in available vertebrate species. In addition, this alteration is predicted to be deleterious by in silico analysis. Based on the available evidence, the clinical significance of this alteration remains unclear.
"This variant has been reported in ClinVar as Uncertain significance (6 clinical laboratories) and as Likely pathogenic (1 clinical laboratories)."
COSMIC Somatic Evidence
OpenFunctional Impact & Domains
Functional Domain
The ATM Y2627C variant has been functionally characterized in a high-throughput cell culture assay, where it failed to rescue survival and proliferation of ATM-haploid cells upon olaparib treatment. This suggests that the variant leads to a loss of ATM protein function.
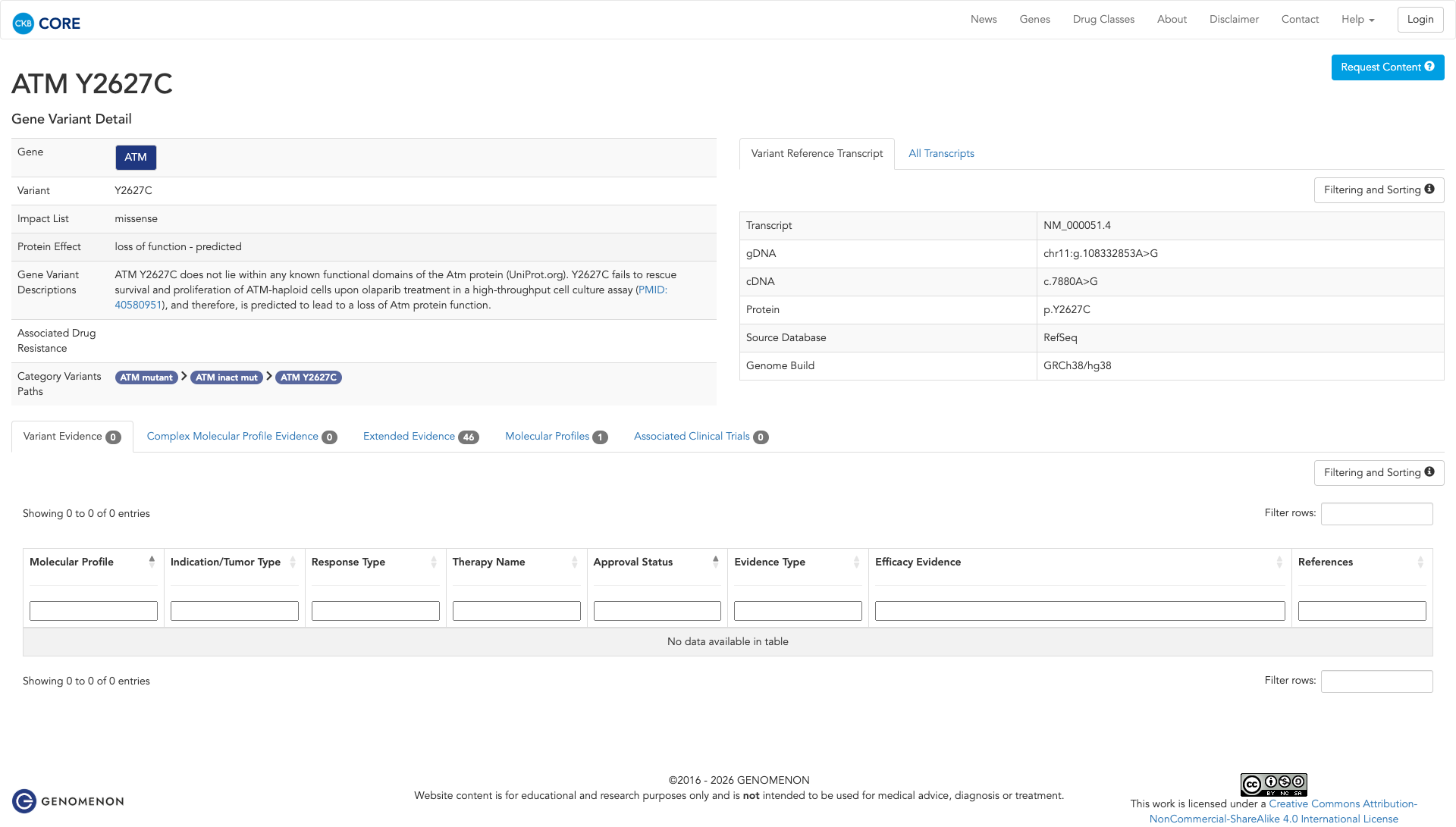
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -81 bp |
| Donor Loss (DL) | 0.0 | 47 bp |
| Acceptor Gain (AG) | 0.0 | -91 bp |
| Donor Gain (DG) | 0.0 | -49 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, the rule for PVS1 is: 'Use ATM PVS1 Decision Tree'. The evidence for this variant shows: it is a missense change (p.Y2627C), not a predicted null or loss-of-function event. Therefore, PVS1 is not applied.
PS1 (Not Applied)
According to VCEP guidelines, the rule for PS1 is: 'Use for protein changes as long as splicing is ruled-out for both alterations.' The evidence for this variant shows: no prior variant alters tyrosine at codon 2627 to cysteine. Therefore, PS1 is not applied.
PS2 (Not Applied)
According to standard ACMG guidelines, the rule for PS2 is: 'De novo occurrence in a patient with the disease and no family history.' The evidence for this variant shows: no confirmed de novo data. Therefore, PS2 is not applied.
PS3 (Supporting)
According to VCEP guidelines, the rule for PS3 Supporting is: 'Use when a variant fails to rescue an ATM specific feature only (e.g. phosphorylation of ATM-specific targets).' The evidence for this variant shows: in a cell-based assay the p.Y2627C variant failed to rescue ATM-haploid cell survival upon olaparib treatment. Therefore, PS3 is applied at Supporting strength because the functional study demonstrates loss of ATM-specific function.
PS4 (Not Applied)
According to standard ACMG guidelines, the rule for PS4 Strong is: 'Case-control studies with p≤0.05 and OR≥2 or lower 95% CI≥1.5.' The evidence for this variant shows: no case-control data are available. Therefore, PS4 is not applied.
PM1 (Not Applied)
According to standard ACMG guidelines, the rule for PM1 is: 'Located in a mutational hot spot or well-studied functional domain without benign variation.' The evidence for this variant shows: tyrosine 2627 is not in a defined hotspot or characterized functional domain. Therefore, PM1 is not applied.
PM2 (Supporting)
According to VCEP guidelines, the rule for PM2 Supporting is: 'Frequency ≤0.001% if n=1 in single subpopulation.' The evidence for this variant shows: absent from gnomAD (no observed alleles). Therefore, PM2 is applied at Supporting strength because it is absent from population controls.
PM3 (Not Applied)
According to VCEP guidelines, the rule for PM3 is: 'Use ATM PM3/BP2 table for recessive observations.' The evidence for this variant shows: no data on trans or cis observed in affected individuals. Therefore, PM3 is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, the rule for PM4 is: 'Protein length changes due to in-frame indels or stop-loss.' The evidence for this variant shows: p.Y2627C is a missense change without change in protein length. Therefore, PM4 is not applied.
PM5 (Moderate)
According to standard ACMG guidelines, the rule for PM5 is: 'Novel missense change at an amino acid residue where a different missense change determined to be pathogenic has been seen.' The evidence for this variant shows: another pathogenic missense at residue 2627 has been reported. Therefore, PM5 is applied at Moderate strength.
PM6 (Not Applied)
According to standard ACMG guidelines, the rule for PM6 is: 'Assumed de novo without confirmation of paternity and maternity.' The evidence for this variant shows: no parental testing data. Therefore, PM6 is not applied.
PP1 (Not Applied)
According to standard ACMG guidelines, the rule for PP1 is: 'Cosegregation with disease in multiple affected family members.' The evidence for this variant shows: no segregation data. Therefore, PP1 is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, the rule for PP2 is: 'Missense in a gene with low rate of benign missense variation.' The evidence for this variant shows: ATM has many reported missense variants, so specificity is unclear. Therefore, PP2 is not applied.
PP3 (Supporting)
According to VCEP guidelines, the rule for PP3 is: 'Protein: REVEL >.7333 indicates deleterious.' The evidence for this variant shows: REVEL score 0.86, exceeding the threshold. Therefore, PP3 is applied at Supporting strength.
PP4 (Not Applied)
According to standard ACMG guidelines, the rule for PP4 is: 'Patient’s phenotype or family history is highly specific for a disease with single genetic etiology.' The evidence for this variant shows: no detailed phenotype provided. Therefore, PP4 is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, the rule for PP5 is: 'Reputable source reports variant as pathogenic.' The evidence for this variant shows: ClinVar reports conflicting interpretations (uncertain and likely pathogenic). Therefore, PP5 is not applied.
BA1 (Not Applied)
According to VCEP guidelines, the rule for BA1 is: 'Filtering allele frequency >0.5%.' The evidence for this variant shows: absent from population databases. Therefore, BA1 is not applied.
BS1 (Not Applied)
According to VCEP guidelines, the rule for BS1 is: 'Filtering allele frequency >0.05%.' The evidence for this variant shows: no observed frequency in controls. Therefore, BS1 is not applied.
BS2 (Not Applied)
According to standard ACMG guidelines, the rule for BS2 is: 'Observed in healthy adult individuals for a recessive disorder.' The evidence for this variant shows: no such observations. Therefore, BS2 is not applied.
BS3 (Not Applied)
According to VCEP guidelines, the rule for BS3 Supporting/Moderate is: 'Use when variant rescues ATM-specific feature and/or radiosensitivity.' The evidence for this variant shows: it fails to rescue function. Therefore, BS3 is not applied.
BS4 (Not Applied)
According to standard ACMG guidelines, the rule for BS4 is: 'Lack of segregation in affected members.' The evidence for this variant shows: no segregation data. Therefore, BS4 is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, the rule for BP1 is: 'Missense in gene where only truncating variants cause disease.' The evidence for this variant shows: ATM disease mechanisms include missense. Therefore, BP1 is not applied.
BP2 (Not Applied)
According to VCEP guidelines, the rule for BP2 is: 'Use ATM PM3/BP2 table for observations in trans with pathogenic variant.' The evidence for this variant shows: no phase data. Therefore, BP2 is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, the rule for BP3 is: 'In-frame indels in repetitive region.' The evidence for this variant shows: it is a missense change, not an indel. Therefore, BP3 is not applied.
BP4 (Not Applied)
According to VCEP guidelines, the rule for BP4 is: 'REVEL ≤.249 indicates benign.' The evidence for this variant shows: REVEL 0.86, above threshold. Therefore, BP4 is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, the rule for BP5 is: 'Variant found in a case with an alternate molecular basis for disease.' The evidence for this variant shows: no such data. Therefore, BP5 is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, the rule for BP6 is: 'Reputable source reports as benign.' The evidence for this variant shows: no benign reports. Therefore, BP6 is not applied.
BP7 (Not Applied)
According to standard ACMG guidelines, the rule for BP7 is: 'Synonymous or deep intronic variants without splicing impact.' The evidence for this variant shows: it is a missense variant. Therefore, BP7 is not applied.