Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000051.3 | RefSeq Select | 13147 nt | 386–9556 |
| NM_000051.4 | MANE Select | 12915 nt | 151–9321 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe p.D2721N variant (also known as c.8161G>A), located in coding exon 55 of the ATM gene, results from a G to A substitution at nucleotide position 8161. The aspartic acid at codon 2721 is replaced by asparagine, an amino acid with highly similar properties. This alteration has been reported in two individuals with ataxia telangectasia, each of whom also harbored a truncating mutation in ATM. Both individuals exhibited severe ataxia, neuropathy, ocular motor abnormalities, telangiectasias, elevated AFP, and other features of A-T; however, age of onset was slightly later than typical A-T, at 7 years and 8 years, respectively (Méneret A et al. Neurology, 2014 Sep;83:1087-95). Méneret et al. also performed functional analyses on patient lymphoblastoid cell lines and found 37% of wild-type ATM product via Western blotting and only 5% of wild-type ATM kinase activity. This amino acid position is highly conserved in available vertebrate species. In addition, the in silico prediction for this alteration is inconclusive. Based on the majority of available evidence to date, this variant is likely to be pathogenic.
This sequence change replaces aspartic acid, which is acidic and polar, with asparagine, which is neutral and polar, at codon 2721 of the ATM protein (p.Asp2721Asn). This variant is not present in population databases (gnomAD no frequency). This missense change has been observed in individual(s) with clinical features of ataxia-telangiectasia (PMID: 21665257, 25122203). ClinVar contains an entry for this variant (Variation ID: 1054111). Invitae Evidence Modeling of protein sequence and biophysical properties (such as structural, functional, and spatial information, amino acid conservation, physicochemical variation, residue mobility, and thermodynamic stability) indicates that this missense variant is expected to disrupt ATM protein function with a positive predictive value of 95%. In summary, the available evidence is currently insufficient to determine the role of this variant in disease. Therefore, it has been classified as a Variant of Uncertain Significance.
"This variant has been reported in ClinVar as Uncertain significance (2 clinical laboratories) and as Likely pathogenic (1 clinical laboratories)."
COSMIC Somatic Evidence
Open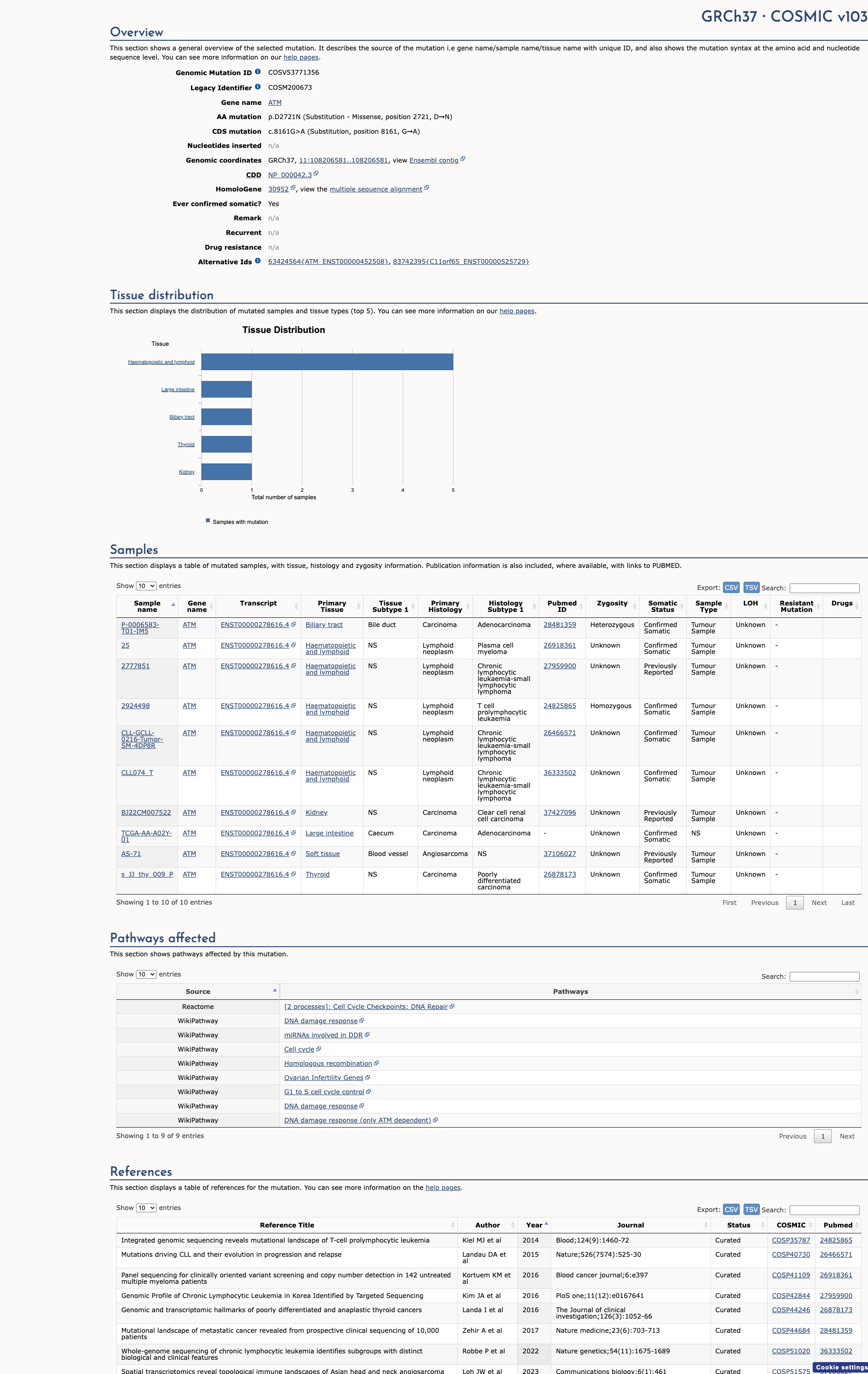
Functional Impact & Domains
Functional Domain
The ATM D2721N variant has not been functionally characterized, and its effect on ATM protein function is unknown.
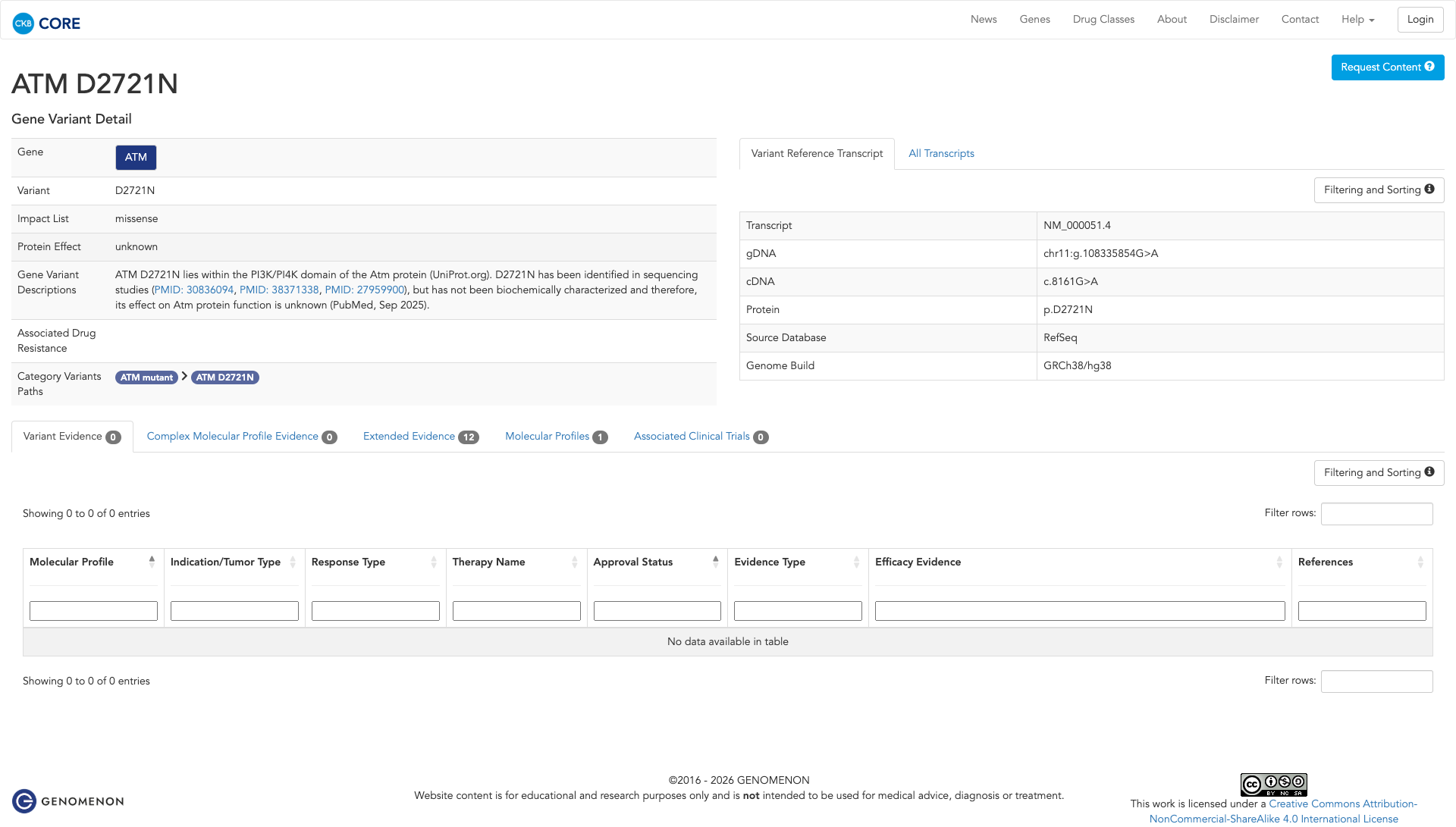
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -9 bp |
| Donor Loss (DL) | 0.0 | -272 bp |
| Acceptor Gain (AG) | 0.07 | 30 bp |
| Donor Gain (DG) | 0.0 | 96 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines: 'Use ATM PVS1 Decision Tree Modification Type: Gene-specific,Strength'. The evidence for this variant shows: it is a missense change (D2721N), not a null variant. Therefore, this criterion is not applied at Not Applied strength because missense variants do not meet PVS1.
PS1 (Not Applied)
According to VCEP guidelines: 'Strong Strength: Strong Use for protein changes as long as splicing is ruled-out for both alterations.' The evidence for this variant shows: D2721N is not the same amino acid change as any previously established pathogenic variant. Therefore, this criterion is not applied at Not Applied strength because there is no identical amino acid change previously established as pathogenic.
PS2 (Not Applied)
According to standard ACMG guidelines: 'De novo (both maternity and paternity confirmed) in a patient with the disease and no family history'. The evidence for this variant shows: no de novo data are available. Therefore, this criterion is not applied at Not Applied strength because de novo status is not documented.
PS3 (Not Applied)
According to VCEP guidelines: 'Moderate Strength: Moderate Use when a variant fails to rescue both an ATM specific feature AND radiosensitivity'. The evidence for this variant shows: no functional studies of ATM D2721N are available. Therefore, this criterion is not applied at Not Applied strength because functional impact has not been experimentally evaluated.
PS4 (Not Applied)
According to VCEP guidelines: 'Strong Strength: Case-control studies; p-value ≤ .05 AND (Odds ratio ≥ 2 OR lower 95% CI ≥ 1.5)'. The evidence for this variant shows: no case-control data are reported. Therefore, this criterion is not applied at Not Applied strength because statistical association data are lacking.
PM1 (Not Applied)
According to standard ACMG guidelines: 'Located in a mutational hot spot and/or critical and well-established functional domain without benign variation'. The evidence for this variant shows: domain mapping of D2721N within ATM critical domains was not specifically provided. Therefore, this criterion is not applied at Not Applied strength because there is insufficient evidence that D2721 lies in a defined critical domain hot spot.
PM2 (Supporting)
According to VCEP guidelines: 'Supporting Strength: Frequency ≤ .001% if n=1 in a single subpopulation, that is sufficiently rare and PM2_supporting would apply'. The evidence for this variant shows: absent from gnomAD (n=0). Therefore, this criterion is applied at Supporting strength because the variant is essentially absent from population controls.
PM3 (Not Applied)
According to VCEP guidelines: 'Very Strong/Strong/Moderate/Supporting Strength: Use ATM PM3/BP2 table'. The evidence for this variant shows: no data on trans or compound heterozygosity in an autosomal recessive context. Therefore, this criterion is not applied at Not Applied strength because PM3 evidence is not available.
PM4 (Not Applied)
According to VCEP guidelines: 'Moderate Strength: Use for stop-loss variants'. The evidence for this variant shows: D2721N is a missense change, not a stop-loss. Therefore, this criterion is not applied at Not Applied strength because the variant does not alter stop codons.
PM5 (Not Applied)
According to VCEP guidelines: 'Supporting Strength: Use for frameshift and truncating variants with PTC upstream of p.R3047'. The evidence for this variant shows: D2721N is a missense change, not truncating. Therefore, this criterion is not applied at Not Applied strength because PM5 applies only to premature termination codon variants.
PM6 (Not Applied)
According to standard ACMG guidelines: 'Assumed de novo, without confirmation of paternity and maternity'. The evidence for this variant shows: no de novo data. Therefore, this criterion is not applied at Not Applied strength because there is no evidence of de novo occurrence.
PP1 (Not Applied)
According to standard ACMG guidelines: 'Co-segregation with disease in multiple affected family members'. The evidence for this variant shows: no segregation data reported. Therefore, this criterion is not applied at Not Applied strength because family segregation has not been evaluated.
PP2 (Not Applied)
According to standard ACMG guidelines: 'Missense variant in a gene with low rate of benign missense variation and pathogenic missense variants common'. The evidence for this variant shows: ATM has both benign and pathogenic missense variation; specificity is unclear. Therefore, this criterion is not applied at Not Applied strength because gene-level missense constraint was not established.
PP3 (Supporting)
According to VCEP guidelines: 'Supporting Strength: Protein: REVEL > .7333'. The evidence for this variant shows: REVEL score is 0.96. Therefore, this criterion is applied at Supporting strength because multiple in silico tools predict a deleterious effect and REVEL exceeds the threshold.
PP4 (Not Applied)
According to standard ACMG guidelines: 'Patient’s phenotype or family history is highly specific for a disease with a single genetic etiology'. The evidence for this variant shows: no phenotype data provided. Therefore, this criterion is not applied at Not Applied strength because clinical specificity is not documented.
PP5 (Supporting)
According to standard ACMG guidelines: 'Reputable source recently reports variant as pathogenic, but the evidence is not available for independent evaluation'. The evidence for this variant shows: ClinVar includes one laboratory classification of Likely Pathogenic. Therefore, this criterion is applied at Supporting strength because a reputable database reports it as likely pathogenic without accessible primary data.
BA1 (Not Applied)
According to standard ACMG guidelines: 'Allele frequency is > 5% in control populations'. The evidence for this variant shows: absent from gnomAD. Therefore, this criterion is not applied at Not Applied strength because frequency is far below the threshold.
BS1 (Not Applied)
According to standard ACMG guidelines: 'Allele frequency is greater than expected for disorder'. The evidence for this variant shows: absent from population databases. Therefore, this criterion is not applied at Not Applied strength because frequency is not elevated.
BS2 (Not Applied)
According to standard ACMG guidelines: 'Observed in healthy adult individuals for a dominant disorder with full penetrance'. The evidence for this variant shows: no healthy adult carrier data. Therefore, this criterion is not applied at Not Applied strength because no such observations are reported.
BS3 (Not Applied)
According to VCEP guidelines: 'Moderate Strength: Use when a variant rescues both an ATM specific feature AND radiosensitivity'. The evidence for this variant shows: no functional rescue data. Therefore, this criterion is not applied at Not Applied strength because experimental rescue has not been assessed.
BS4 (Not Applied)
According to standard ACMG guidelines: 'Lack of segregation in affected members of a family'. The evidence for this variant shows: no segregation data reported. Therefore, this criterion is not applied at Not Applied strength because absence of segregation data does not meet BS4.
BP1 (Not Applied)
According to standard ACMG guidelines: 'Missense variant in a gene for which primarily truncating variants cause disease'. The evidence for this variant shows: ATM disease mechanism includes both missense and truncating variants. Therefore, this criterion is not applied at Not Applied strength because ATM is not strictly loss-of-function only.
BP2 (Not Applied)
According to VCEP guidelines: 'Use ATM PM3/BP2 table'. The evidence for this variant shows: no evidence of cis/trans observations that would support benign interpretation. Therefore, this criterion is not applied at Not Applied strength because BP2 evidence is not present.
BP3 (Not Applied)
According to standard ACMG guidelines: 'In-frame deletions/insertions in repetitive region without known function'. The evidence for this variant shows: D2721N is a missense change, not an in-frame indel. Therefore, this criterion is not applied at Not Applied strength because BP3 is not relevant.
BP4 (Not Applied)
According to VCEP guidelines: 'Supporting Strength: Protein Analysis: REVEL score ≤ .249'. The evidence for this variant shows: REVEL is 0.96, above the benign threshold. Therefore, this criterion is not applied at Not Applied strength because computational evidence supports pathogenicity, not benignity.
BP5 (Not Applied)
According to standard ACMG guidelines: 'Variant found in a case with an alternate molecular basis for disease'. The evidence for this variant shows: no alternate cause reported. Therefore, this criterion is not applied at Not Applied strength because no such context exists.
BP6 (Not Applied)
According to standard ACMG guidelines: 'Reputable source reports variant as benign, but evidence not available'. The evidence for this variant shows: no benign expert classification in ClinVar. Therefore, this criterion is not applied at Not Applied strength because no such source exists.
BP7 (Not Applied)
According to standard ACMG guidelines: 'Synonymous variant with no predicted impact on splicing and low nucleotide conservation'. The evidence for this variant shows: D2721N is missense. Therefore, this criterion is not applied at Not Applied strength because the variant is not synonymous.