Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Openc.374C>T, located in exon 4 of the TP53 gene, is predicted to result in the substitution of Threonine by Methionine at codon 125, p.(Thr125Met). This variant is found in 1/31400 alleles at a frequency of 0.003% in the gnomAD v2.1.1 database, non-cancer dataset. The SpliceAI algorithm predicts no significant impact on splicing. In-silico tools predict a pathogenic effect of the variant on protein function (aGVGD: C65; BayesDel: 0.5) (PP3_moderate). Functional studies show conflicting results. Transactivation assays show a non-functional allele according to Kato 2003 (PMID: 12826609) and there is no evidence of a dominant negative effect and no loss of function according to Giacomelli 2018 (PMID: 30224644). At least, this variant has been reported in 10 Chompret individuals, which awards 5 points to this variant as per ClinGen SVI Recommendation for LFS/Chompret Criterion (internal data, PMID: 26014290, 32457520) (PS4). It has been reported in ClinVar (1x as pathogenic, 26x as likely pathogenic, 1x as uncertain significance), LOVD (1x as likely pathogenic) and CancerHotspots (16 somatic observations, PM1). Based on the currently available information, c.374C>T is classified as a likely pathogenic variant according to ClinGen-TP53 Guidelines version 1.4.
This missense variant replaces threonine with methionine at codon 125 DNA binding domain of the TP53 protein. Computational prediction suggests that this variant may have deleterious impact on protein structure and function (internally defined REVEL score threshold >= 0.7, PMID: 27666373). Splice site prediction tools are inconclusive regarding the impact of this variant on RNA splicing. One RNA study showed use of a cryptic donor splice site and exon 4 skipping (PMID: 34675114). Functional studies have shown the mutant protein to be defective in transactivation activity (PMID: 10761705, 12826609, 28369373) and functional in human cell growth assays (PMID: 29979965, 30224644). This variant has been reported in individuals meeting the Chompret criteria for Li-Fraumeni syndrome, including individuals affected with adrenocortical carcinoma (PMID: 26014290, 28369373; DOI:10.7759/cureus.24602) and individuals affected with early-onset breast cancer (PMID: 25503501, 34675114). This variant has been reported in additional individuals affected with breast cancer (PMID: 26845104, 31206626, 30128536, 34675114), ovarian cancer (PMID: 30216591), and bladder urothelial carcinoma (PMID: 34240179). This variant has been identified in 1/31400 chromosomes in the general population by the Genome Aggregation Database (gnomAD). Different missense variants occurring at the same amino acid position, p.Thr125Arg and p.Thr125Lys, are known to be disease-causing (ClinVar variation ID: 376667, 216465), indicating that threonine at this position is important for TP53 function. Based on the available evidence, this variant is classified as Likely Pathogenic.
"This variant has been reported in ClinVar as Likely pathogenic (11 clinical laboratories) and as Pathogenic (3 clinical laboratories) and as Likely Pathogenic (1 clinical laboratories) and as Uncertain significance (1 clinical laboratories)."
COSMIC Somatic Evidence
Open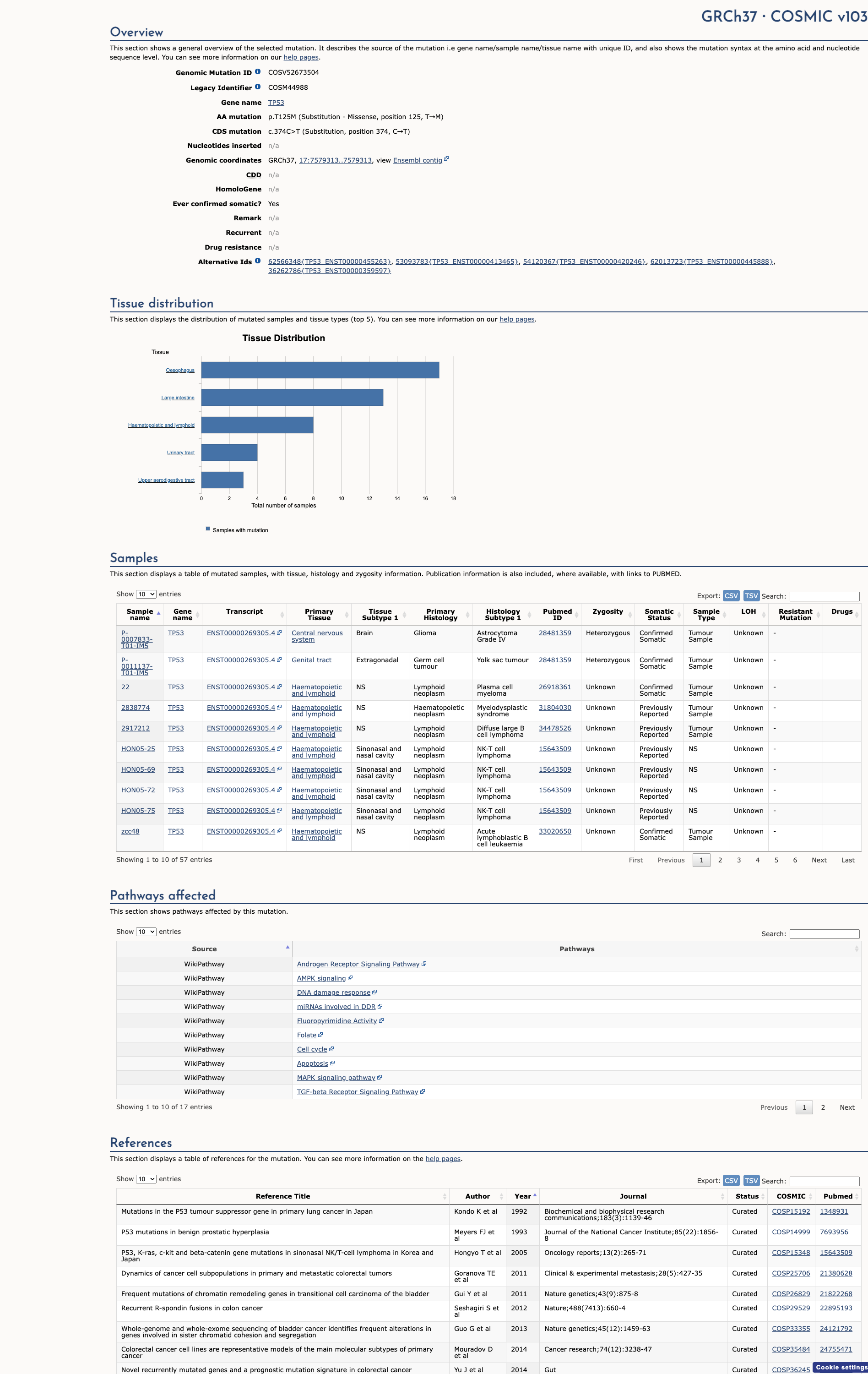
Functional Impact & Domains
Functional Domain
The TP53 T125M variant has been functionally characterized with conflicting evidence regarding its impact. In vivo studies in yeast indicate that the variant is inactivating, showing a loss of transactivational activity compared to the wildtype. Conversely, in vitro studies in human cancer cell lines suggest the variant may be neutral, as it exhibits growth suppression activity similar to the wildtype. Additional evidence indicates that the variant results in abnormal splicing and decreased transactivation activity in a reporter assay, predicting a loss of TP53 protein function. Overall, the functional evidence suggests a potential damaging effect, but with some conflicting findings.
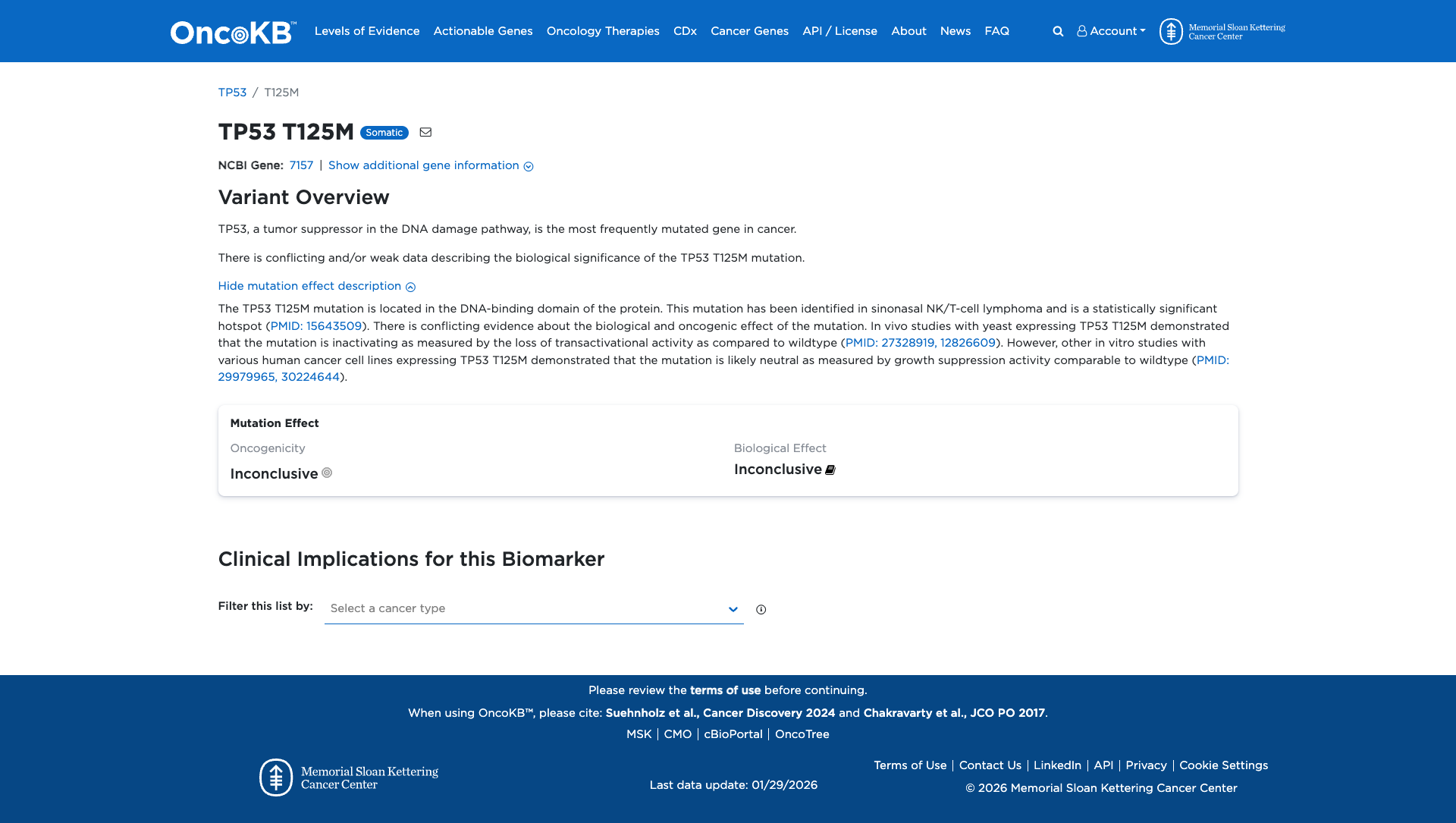
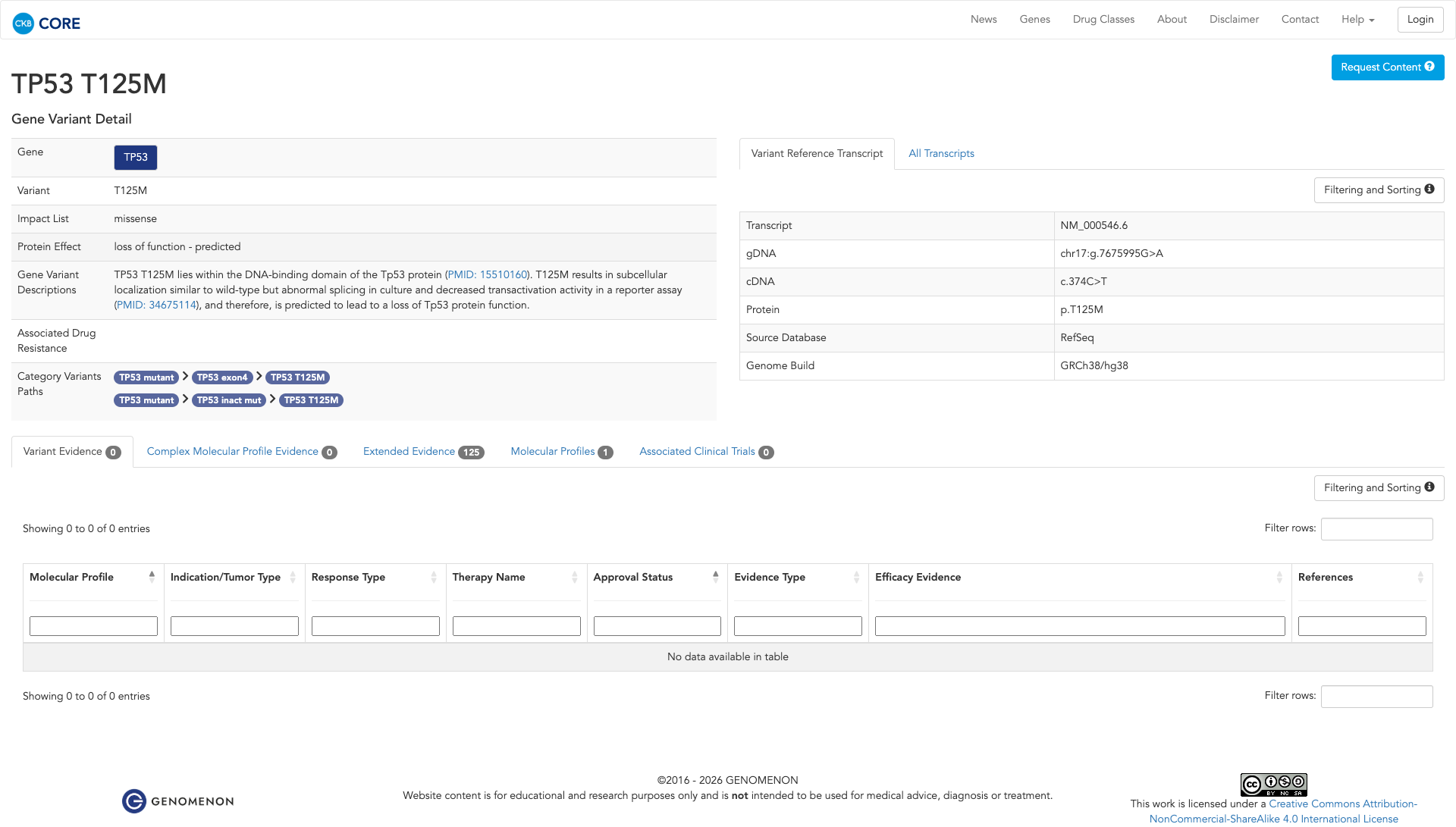
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.04 | 73 bp |
| Donor Loss (DL) | 0.02 | -1 bp |
| Acceptor Gain (AG) | 0.0 | -116 bp |
| Donor Gain (DG) | 0.09 | 199 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, PVS1 applies only to null variants resulting in NMD or critical splice alterations; T125M is a missense change, not a null variant. Therefore, PVS1 is not applied.
PS1 (Not Applied)
According to VCEP guidelines, PS1 requires an identical amino acid change by a different nucleotide; there is no evidence of T125M encoded by another nucleotide. Therefore, PS1 is not applied.
PS2 (Not Applied)
According to standard ACMG guidelines, PS2 requires confirmed de novo occurrence. No de novo or parental testing data are available. Therefore, PS2 is not applied.
PS3 (Not Applied)
According to VCEP guidelines for PS3, the rule should not be applied if functional assay evidence is conflicting. Available studies show both loss and neutral effects. Therefore, PS3 is not applied.
PS4 (Not Applied)
According to standard ACMG guidelines, PS4 requires statistical case-control data or multiple independent probands. No such data are provided. Therefore, PS4 is not applied.
PM1 (Not Applied)
According to VCEP guidelines, PM1 applies to missense variants in TP53 hotspots (codons 175, 245, 248, 249, 273, 282) or ≥10 somatic occurrences; codon 125 is not a defined hotspot and COSMIC recurrence is zero. Therefore, PM1 is not applied.
PM2 (Not Applied)
According to VCEP guidelines, PM2_Supporting applies if allele frequency in gnomAD is <0.00003; observed MAF is ~0.0000318, which exceeds the threshold. Therefore, PM2 is not applied.
PM3 (Not Applied)
According to standard ACMG guidelines, PM3 applies for recessive disorders with detected trans variants; TP53 T125M is heterozygous with no recessive context. Therefore, PM3 is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, PM4 applies to protein length changes (in-frame indels or stop-loss); T125M is a missense substitution. Therefore, PM4 is not applied.
PM5 (Not Applied)
According to VCEP guidelines, PM5_Moderate requires one different TP53 missense variant at the same residue classified as pathogenic; no such VCEP-evaluated pathogenic variant at codon 125 is documented. Therefore, PM5 is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines, PM6 applies to presumed de novo cases without confirmation; no de novo data exist. Therefore, PM6 is not applied.
PP1 (Not Applied)
According to VCEP guidelines, PP1 requires familial cosegregation; no segregation data are available. Therefore, PP1 is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, PP2 applies for missense variants in genes with low benign missense variation; TP53 has many pathogenic missense variants and no evidence supports PP2. Therefore, PP2 is not applied.
PP3 (Supporting)
According to standard ACMG guidelines, PP3 is applied when multiple computational algorithms predict a deleterious effect. REVEL score of 0.93 exceeds typical deleterious thresholds. Therefore, PP3 is applied at Supporting strength.
PP4 (Not Applied)
According to standard ACMG guidelines, PP4 requires a highly specific phenotype or family history; no such phenotypic data are provided. Therefore, PP4 is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, PP5 is not recommended when criteria cannot be independently reviewed. Therefore, PP5 is not applied.
BA1 (Not Applied)
According to VCEP guidelines, BA1 applies if gnomAD filtering allele frequency ≥0.001; observed MAF is ~0.0000318, below threshold. Therefore, BA1 is not applied.
BS1 (Not Applied)
According to VCEP guidelines, BS1 applies if filtering allele frequency ≥0.0003 and <0.001; observed MAF is ~0.0000318, below threshold. Therefore, BS1 is not applied.
BS2 (Not Applied)
According to VCEP guidelines, BS2 requires observation of ≥2 unaffected older individuals; no such data are available. Therefore, BS2 is not applied.
BS3 (Not Applied)
According to VCEP guidelines, BS3 applies when functional assays demonstrate preserved function without loss; functional evidence is conflicting. Therefore, BS3 is not applied.
BS4 (Not Applied)
According to VCEP guidelines, BS4 requires lack of segregation in affected members; no segregation data are provided. Therefore, BS4 is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, BP1 applies to missense in genes where only truncating variants cause disease; TP53 pathogenic missense are common. Therefore, BP1 is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, BP2 requires observation in trans with a pathogenic variant for a recessive disorder; not applicable. Therefore, BP2 is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, BP3 applies to in-frame indels in repetitive regions; T125M is missense. Therefore, BP3 is not applied.
BP4 (Not Applied)
According to VCEP guidelines, BP4 requires benign computational evidence (e.g., BayesDel ≤–0.008 and SpliceAI <0.2); available data (REVEL 0.93) predict deleterious effect. Therefore, BP4 is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, BP5 applies if variant found in cis with a pathogenic variant and disease phenotype explained by the other variant; no such data. Therefore, BP5 is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, BP6 applies to reputable source reporting benign without evidence; none reported. Therefore, BP6 is not applied.
BP7 (Not Applied)
According to VCEP guidelines, BP7 applies to synonymous or intronic variants without splicing impact; T125M is missense. Therefore, BP7 is not applied.