Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_007294.2 | Alternative | 7191 nt | 201–5792 |
| NM_007294.3 | RefSeq Select | 7224 nt | 233–5824 |
| NM_007294.4 | MANE Select | 7088 nt | 114–5705 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis missense variant replaces glutamic acid with lysine at codon 1836 of the BRCA1 protein. Computational prediction tool is inconclusive regarding the impact of this variant on protein structure and function (internally defined REVEL score threshold 0.5 < inconclusive < 0.7, PMID: 27666373). Functional studies have reported discrepant findings on variant protein impacts, including modest (PMID: 21473589) to severe disruption (PMID: 20516115) in phosphopeptide binding activity, greater than 50% of wild-type transcriptional activation (PMID: 20516115), and normal function in complementing BRCA1 deficiency and rescue cell proliferation in haploid cells (PMID: 30209399). To our knowledge, this variant has not been reported in individuals affected with hereditary cancer in the literature. This variant has been identified in 2/251336 chromosomes in the general population by the Genome Aggregation Database (gnomAD). Available evidence is insufficient to determine the role of this variant in disease conclusively. Therefore, this variant is classified as a Variant of Uncertain Significance.
This sequence change replaces glutamic acid, which is acidic and polar, with lysine, which is basic and polar, at codon 1836 of the BRCA1 protein (p.Glu1836Lys). This variant is present in population databases (rs80356942, gnomAD 0.003%). This variant has not been reported in the literature in individuals affected with BRCA1-related conditions. ClinVar contains an entry for this variant (Variation ID: 55605). Advanced modeling performed at Invitae incorporating data from internal and/or published experimental studies (PMID: 30209399) indicates that this missense variant is not expected to disrupt BRCA1 function with a negative predictive value of 95%. Experimental studies are conflicting or provide insufficient evidence to determine the effect of this variant on BRCA1 function (PMID: 20516115, 21473589, 24845084, 30209399, 30765603). In summary, the available evidence is currently insufficient to determine the role of this variant in disease. Therefore, it has been classified as a Variant of Uncertain Significance.
This missense variant replaces glutamic acid with lysine at codon 1836 of the BRCA1 protein. Computational prediction tool is inconclusive regarding the impact of this variant on protein structure and function (internally defined REVEL score threshold 0.5 < inconclusive < 0.7, PMID: 27666373). Functional studies have reported discrepant findings on variant protein impacts, including modest (PMID: 21473589) to severe disruption (PMID: 20516115) in phosphopeptide binding activity, greater than 50% of wild-type transcriptional activation (PMID: 20516115), and normal function in complementing BRCA1 deficiency and rescue cell proliferation in haploid cells (PMID: 30209399). To our knowledge, this variant has not been reported in individuals affected with hereditary cancer in the literature. This variant has been identified in 2/251336 chromosomes in the general population by the Genome Aggregation Database (gnomAD). Available evidence is insufficient to determine the role of this variant in disease conclusively. Therefore, this variant is classified as a Variant of Uncertain Significance.
Variant summary: BRCA1 c.5506G>A (p.Glu1836Lys) results in a conservative amino acid change located in the BRCT domain profile (IPR001357) of the encoded protein sequence. Three of five in-silico tools predict a damaging effect of the variant on protein function. The variant allele was found at a frequency of 8e-06 in 251336 control chromosomes. The available data on variant occurrences in the general population are insufficient to allow any conclusion about variant significance. To our knowledge, no occurrence of c.5506G>A in individuals affected with Hereditary Breast And Ovarian Cancer Syndrome has been reported. Experimental studies provide conflicting reports regarding the functional outcome of this variant (e.g. Findlay_2018, Lee_2010, Carvalho_2014). The following publications have been ascertained in the context of this evaluation (PMID: 30209399, 20516115, 24845084). ClinVar contains an entry for this variant (Variation ID: 55605). Based on the evidence outlined above, the variant was classified as uncertain significance.
The p.E1836K variant (also known as c.5506G>A and 5625G>A), located in coding exon 22 of the BRCA1 gene, results from a G to A substitution at nucleotide position 5506. The glutamic acid at codon 1836 is replaced by lysine, an amino acid with similar properties. One study, which assessed the effect of BRCT missense alterations by structure and sequence based computational analysis, found this alteration to have a probability of affecting structure and function of 0.48 and 0.42, using two sets of analytic features. Additionally, refined sequence based analysis predicted this alteration to affect function (Williams RS et al. J. Biol. Chem. 2003 Dec; 278(52):53007-16). Using several different structural and functional assays, another study found this alteration to be structurally stable; however, they did predict compromised binding activity and specificity, with less than 10% of wild type for both parameters, and found transcriptional activation to be at 55% of wild type, resulting in a classification of uncertain significance for this alteration (Lee MS et al. Cancer Res. 2010 Jun; 70(12):4880-90). Additionally, while one study suggested that this alteration could perturb phosphopeptide recognition and binding (Campbell SJ et al. Structure 2010 Feb; 18(2):167-76), other studies showed that this alteration does not dramatically affect function (Coquelle N et al. Biochemistry 2011 May; 50(21):4579-89; Findlay GM et al. Nature, 2018 Oct;562:217-222). This amino acid position is well conserved in available vertebrate species. In addition, this alteration is predicted to be deleterious by in silico analysis. Based on the available evidence, the clinical significance of this variant remains unclear.
"This variant has been reported in ClinVar as Uncertain significance (9 clinical laboratories) and as Uncertain Significance (1 clinical laboratories) and as Likely benign (1 clinical laboratories)."
COSMIC Somatic Evidence
Open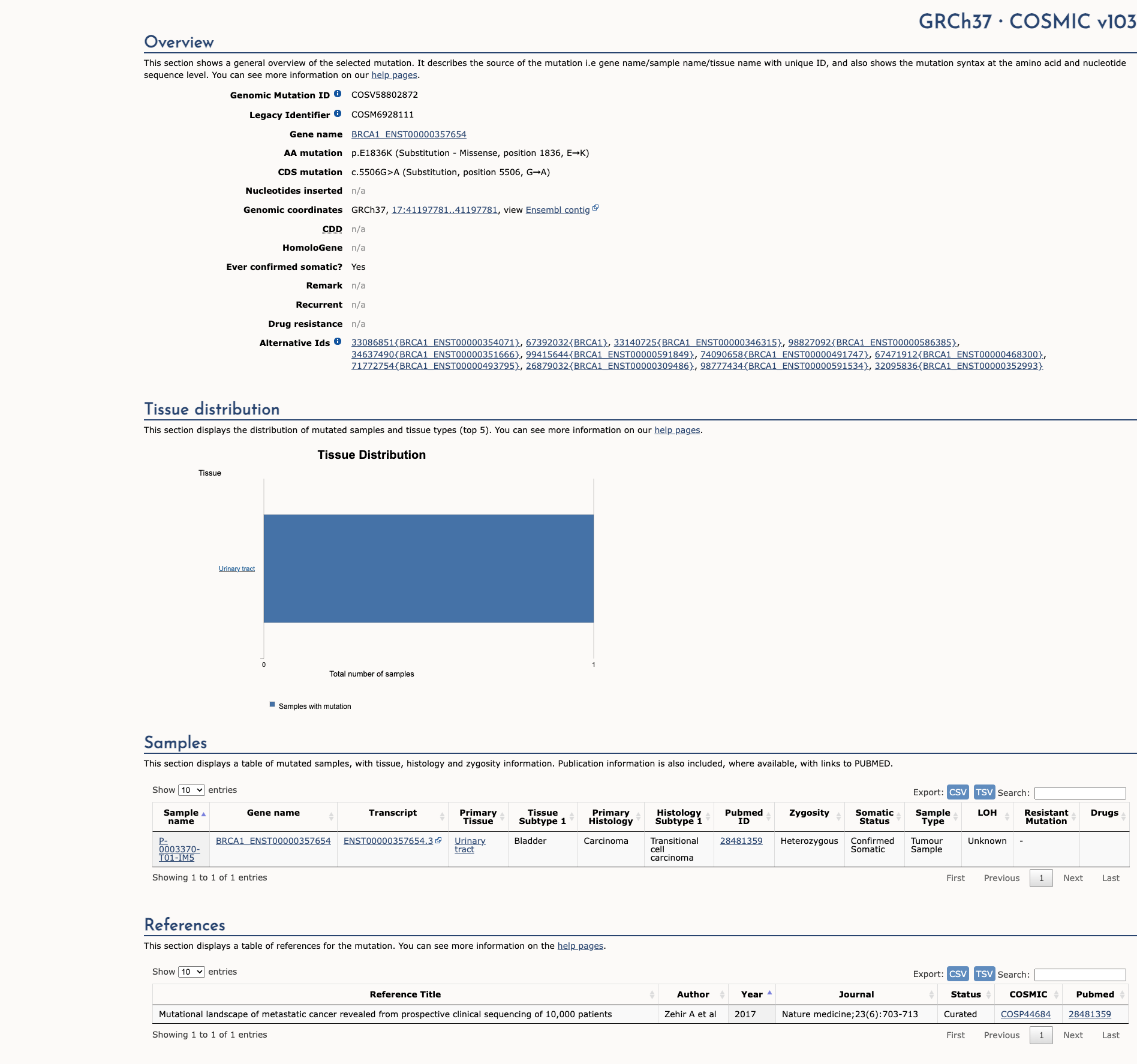
Functional Impact & Domains
Functional Domain
The BRCA1 E1836K variant has been functionally characterized with mixed results. It shows no folding defect and maintains wildtype homologous recombination efficiency and normal localization after mitomycin treatment. However, it exhibits compromised binding activity and specificity, with a minor defect in phosphopeptide binding. Overall, the functional evidence presents conflicting data regarding its impact.
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -222 bp |
| Donor Loss (DL) | 0.0 | -102 bp |
| Acceptor Gain (AG) | 0.0 | 27 bp |
| Donor Gain (DG) | 0.0 | -311 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, PVS1 applies to a null variant (nonsense, frameshift, canonical ±1/2 splice site) in a gene where loss of function is a known mechanism (“Very Strong Null variant…”). The variant NM_007294.4:c.5506G>A is a missense change (p.E1836K), not a null variant. Therefore, PVS1 is not applied.
PS1 (Not Applied)
According to VCEP guidelines, PS1 applies when a variant results in the same amino acid change as a previously established pathogenic variant. There is no known pathogenic variant producing p.E1836K at this codon. Therefore, PS1 is not applied.
PS2 (Not Applied)
According to standard ACMG, PS2 applies for de novo occurrences with confirmed maternity and paternity. No de novo data are available for this variant. Therefore, PS2 is not applied.
PS3 (Not Applied)
According to VCEP guidelines, PS3 applies for well‐established functional studies showing a damaging effect. Functional assays for E1836K show normal homologous recombination and localization but a minor phosphopeptide binding defect, yielding conflicting results. Therefore, PS3 is not applied.
PS4 (Not Applied)
According to VCEP guidelines, PS4 applies when variant prevalence in affected individuals is significantly increased (OR≥4, p≤0.05). No case‐control or case prevalence data are available. Therefore, PS4 is not applied.
PM1 (Not Applied)
According to VCEP guidelines, PM1 applies to variants in a mutational hot‐spot or well‐established functional domain with enrichment of pathogenic variants. Although E1836K lies within the BRCT domain, there is no evidence of enrichment of pathogenic missense changes at this residue or region. Therefore, PM1 is not applied.
PM2 (Supporting)
According to VCEP guidelines, PM2_Supporting applies when a variant is absent or extremely rare in population databases. NM_007294.4:c.5506G>A is exceedingly rare in gnomAD (MAF=0.000796%). Therefore, PM2 is applied at Supporting strength.
PM3 (Not Applied)
According to VCEP guidelines, PM3 applies for variants observed in trans with a pathogenic variant in Fanconi Anemia phenotypes. No such data exist. Therefore, PM3 is not applied.
PM4 (Not Applied)
According to standard ACMG, PM4 applies to protein length changes (in‐frame indels) not relevant to a missense variant. Therefore, PM4 is not applied.
PM5 (Not Applied)
According to VCEP guidelines, PM5 applies to a novel missense change at an amino acid residue where a different pathogenic missense change has been seen. No other pathogenic variants at residue 1836 are reported. Therefore, PM5 is not applied.
PM6 (Not Applied)
According to standard ACMG, PM6 applies to assumed de novo occurrences without confirmation. No such data exist. Therefore, PM6 is not applied.
PP1 (Not Applied)
According to VCEP guidelines, PP1 applies for co-segregation with disease in multiple affected family members. No segregation data are available. Therefore, PP1 is not applied.
PP2 (Not Applied)
According to standard ACMG, PP2 applies for missense variants in genes with low rates of benign missense variation. BRCA1 exhibits many missense variants. Therefore, PP2 is not applied.
PP3 (Not Applied)
According to VCEP guidelines, PP3 applies for missense variants in a functional domain with supportive computational predictions (BayesDel no-AF≥0.28 or SpliceAI≥0.2). BayesDel data are unavailable and SpliceAI predicts no splicing impact. Therefore, PP3 is not applied.
PP4 (Not Applied)
According to VCEP guidelines, PP4 applies to a phenotype highly specific to a single gene. No detailed clinical phenotype data are provided. Therefore, PP4 is not applied.
PP5 (Not Applied)
According to standard ACMG, PP5 applies to unverified assertions of pathogenicity by reputable sources. Existing ClinVar entries are uncertain or likely benign; no definitive pathogenic assertions. Therefore, PP5 is not applied.
BA1 (Not Applied)
According to VCEP guidelines, BA1 applies when FAF>0.1% (0.001). The highest allele frequency (South Asian MAF=0.00327%) equates to 0.0000327, below the BA1 threshold. Therefore, BA1 is not applied.
BS1 (Supporting)
According to VCEP guidelines, BS1_Supporting applies when FAF>0.002% (0.00002) and ≤0.01% (0.0001). The South Asian subpopulation MAF=0.00327% (0.0000327) meets this range. Therefore, BS1 is applied at Supporting strength.
BS2 (Not Applied)
According to VCEP guidelines, BS2 applies to variants observed in healthy adults inconsistent with a recessive phenotype. No such data are available. Therefore, BS2 is not applied.
BS3 (Not Applied)
According to VCEP guidelines, BS3 applies for well‐established functional studies showing no damaging effect. Functional data are conflicting. Therefore, BS3 is not applied.
BS4 (Not Applied)
According to VCEP guidelines, BS4 applies for lack of segregation in affected family members. No segregation data exist. Therefore, BS4 is not applied.
BP1 (Not Applied)
According to VCEP guidelines, BP1 applies to missense variants outside a functional domain with no splicing impact. E1836K is inside the BRCT domain. Therefore, BP1 is not applied.
BP2 (Not Applied)
According to standard ACMG, BP2 applies when co-observed in trans with a pathogenic variant. No such observations are reported. Therefore, BP2 is not applied.
BP3 (Not Applied)
According to standard ACMG, BP3 applies to in-frame deletions/insertions in repetitive regions. This is a missense variant. Therefore, BP3 is not applied.
BP4 (Supporting)
According to VCEP guidelines, BP4_Supporting applies for missense variants inside a clinically important domain with no predicted impact via protein change or splicing (BayesDel no-AF≤0.15 AND SpliceAI≤0.1). SpliceAI=0 and in silico protein predictions do not support a deleterious effect. Therefore, BP4 is applied at Supporting strength.
BP5 (Not Applied)
According to VCEP guidelines, BP5 applies to variants observed in trans with known pathogenic variants without specific phenotype. No such data. Therefore, BP5 is not applied.
BP6 (Not Applied)
According to standard ACMG, BP6 applies to unverified benign assertions. No unambiguous benign assertions exist. Therefore, BP6 is not applied.
BP7 (Not Applied)
According to VCEP guidelines, BP7 applies to silent or intronic variants with no splicing impact. This is a missense variant. Therefore, BP7 is not applied.