Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis variant is considered likely pathogenic. Functional studies indicate this variant impacts protein function [PMID: 7651740, 9635828, 15037740, 16861262]. This variant is expected to disrupt protein structure [Myriad internal data].
The p.V272M pathogenic mutation (also known as c.814G>A), located in coding exon 7 of the TP53 gene, results from a G to A substitution at nucleotide position 814. The valine at codon 272 is replaced by methionine, an amino acid with highly similar properties. This alteration has been detected in a French family with features suggestive of Li-Fraumeni syndrome, a patient diagnosed with rhabdomyosarcoma at age 1 and osteosarcoma at age 2, as well as a patient diagnosed with medulloblastoma at age 3 (Bougeard G et al. J. Med. Genet. 2008 Aug;45(8):535-8; Renaux-Petel M et al. J. Med. Genet. 2017 Oct 25). This alteration was also identified in a patient with late onset adrenal cortical carcinoma (Raymond VM et al. J. Clin. Endocrinol. Metab. 2013 Jan; 98(1):E119-25). The p.V272M alteration has been reported as a somatic alteration in a variety of tumors (Marinelli M et al. Haematologica 2013 Mar; 98(3):371-5; Chiaretti S et al. Genes Chromosomes Cancer 2011 Apr; 50(4):263-74; Chang MT et al. Cancer Discov. 2018 02;8:174-183). This alteration is a well-known temperature sensitive alteration that causes abnormal protein folding and prevents DNA binding at temperatures above 37 degrees Celsius (da Costa NM et al. Cell Cycle 2012 Dec;11(24):4570-8). Functional studies have demonstrated that this alteration interferes with p53 up-regulation in response to DNA damaging agents and shows loss of transactivation capacity in yeast based assays (Barnas Cet al. Int. J. Cancer 1997 Mar;71(1):79-87). Studies conducted in human cell lines indicate this alteration has a dominant negative effect and is deficient at growth suppression (Kotler E et al. Mol. Cell 2018 Jul;71:178-190.e8; Giacomelli AO et al. Nat. Genet. 2018 Oct;50:1381-1387). This amino acid position is not well conserved in available vertebrate species. In addition, this alteration is predicted to be deleterious by in silico analysis. Based on the supporting evidence, this alteration is interpreted as a disease-causing mutation.
PS3; PM1; PS4_MOD; PM6_Strong
Variant summary: TP53 c.814G>A (p.Val272Met) results in a conservative amino acid change located in the DNA-binding domain (IPR011615) of the encoded protein sequence. Four of five in-silico tools predict a damaging effect of the variant on protein function. The variant allele was found at a frequency of 4e-06 in 250892 control chromosomes. c.814G>A has been reported in the literature in individuals affected with Li-Fraumeni Syndrome as de novo occurances (Renaux-Petel_2018, Ohlsen_2022). These data indicate that the variant may be associated with disease. A different variant affecting the same codon has been classified as pathogenic by our lab (c.814G>T, p.Val272Leu), supporting the critical relevance of codon 272 to TP53 protein function. At least one publication reports experimental evidence evaluating an impact on protein function. The most pronounced variant effect results in absent DNA binding and absent activation in response to DNA-damaging treatment (North_2002). The following publications have been ascertained in the context of this evaluation (PMID: 11870884, 35484874, 29070607). ClinVar contains an entry for this variant (Variation ID: 185814). Based on the evidence outlined above, the variant was classified as pathogenic.
"This variant has been reported in ClinVar as Likely Pathogenic (1 clinical laboratories) and as Likely pathogenic (1 clinical laboratories) and as Pathogenic (6 clinical laboratories) and as Pathogenic by ClinGen TP53 Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open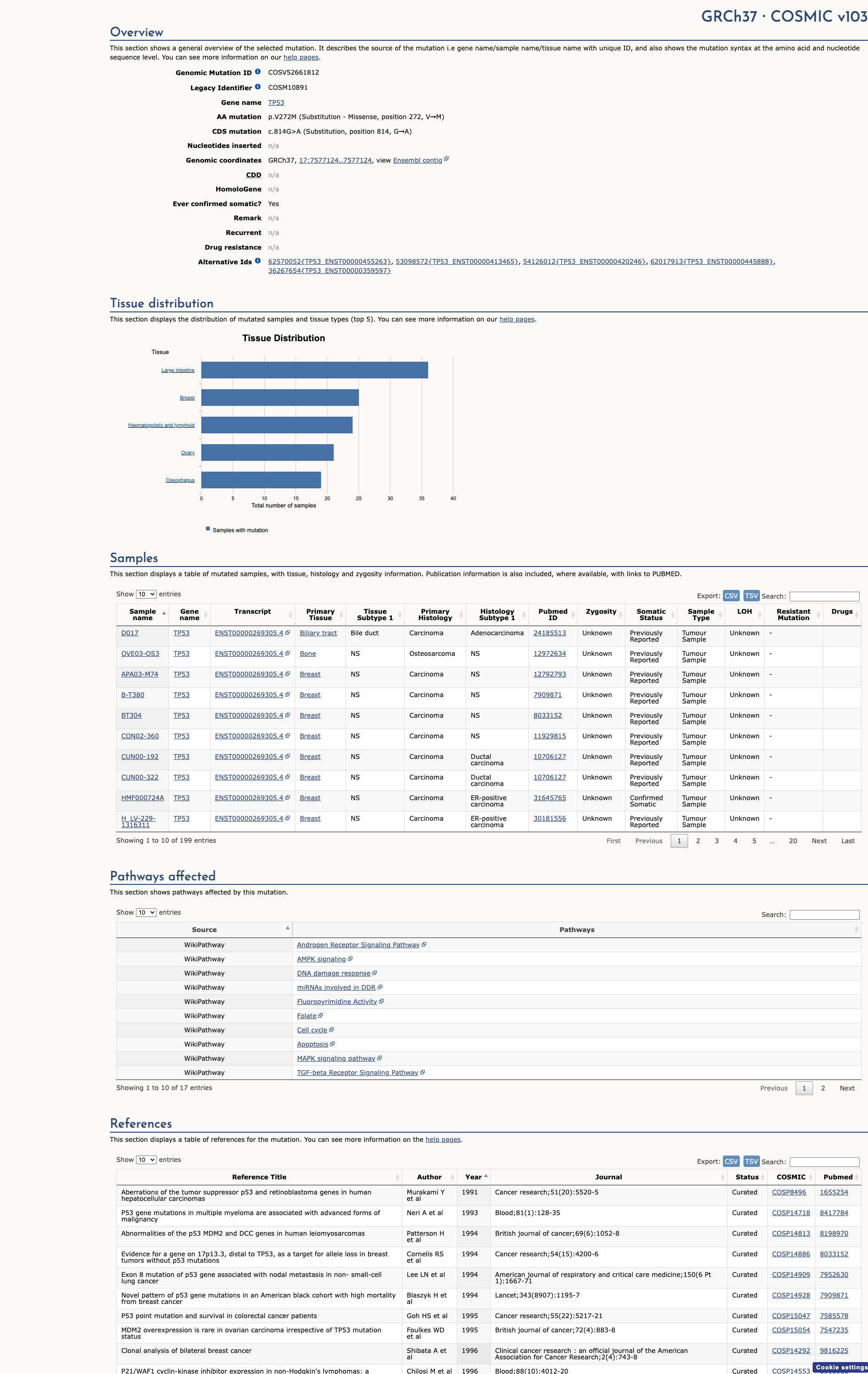
Functional Impact & Domains
Functional Domain
The TP53 V272M variant has been functionally characterized and is likely damaging. It is located in the DNA-binding domain of the TP53 protein and has been shown to be inactivating. In vitro studies demonstrate that this variant results in reduced reactivity with a wildtype conformation-specific antibody, reduced accumulation in cells exposed to DNA-damaging agents, and an inability to rescue TP53-mediated transcriptional activity in TP53-null cells. Additionally, it leads to increased proliferation, migration, invasion, altered subcellular localization, loss of DNA-binding, and transactivation ability in cell culture.
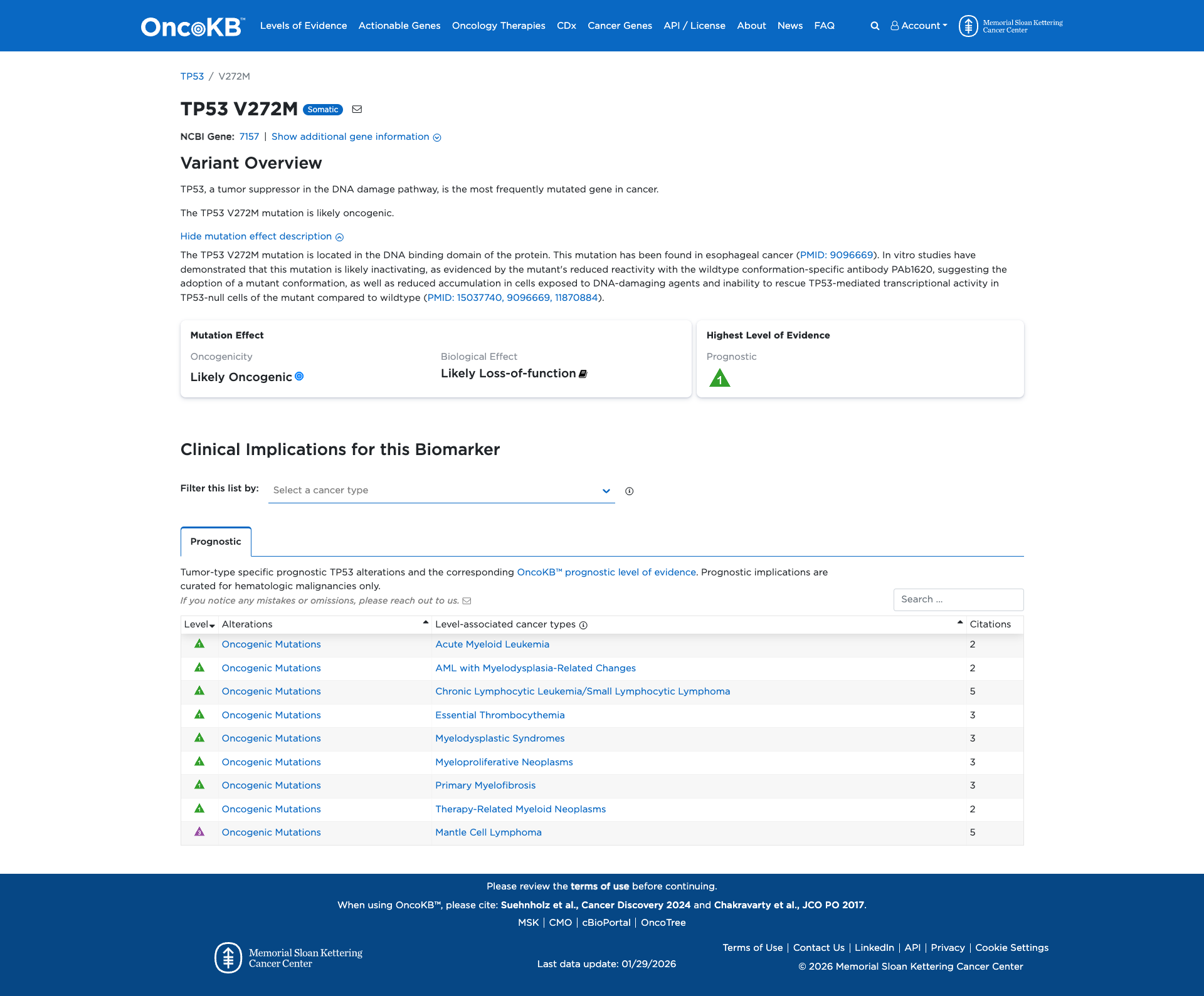
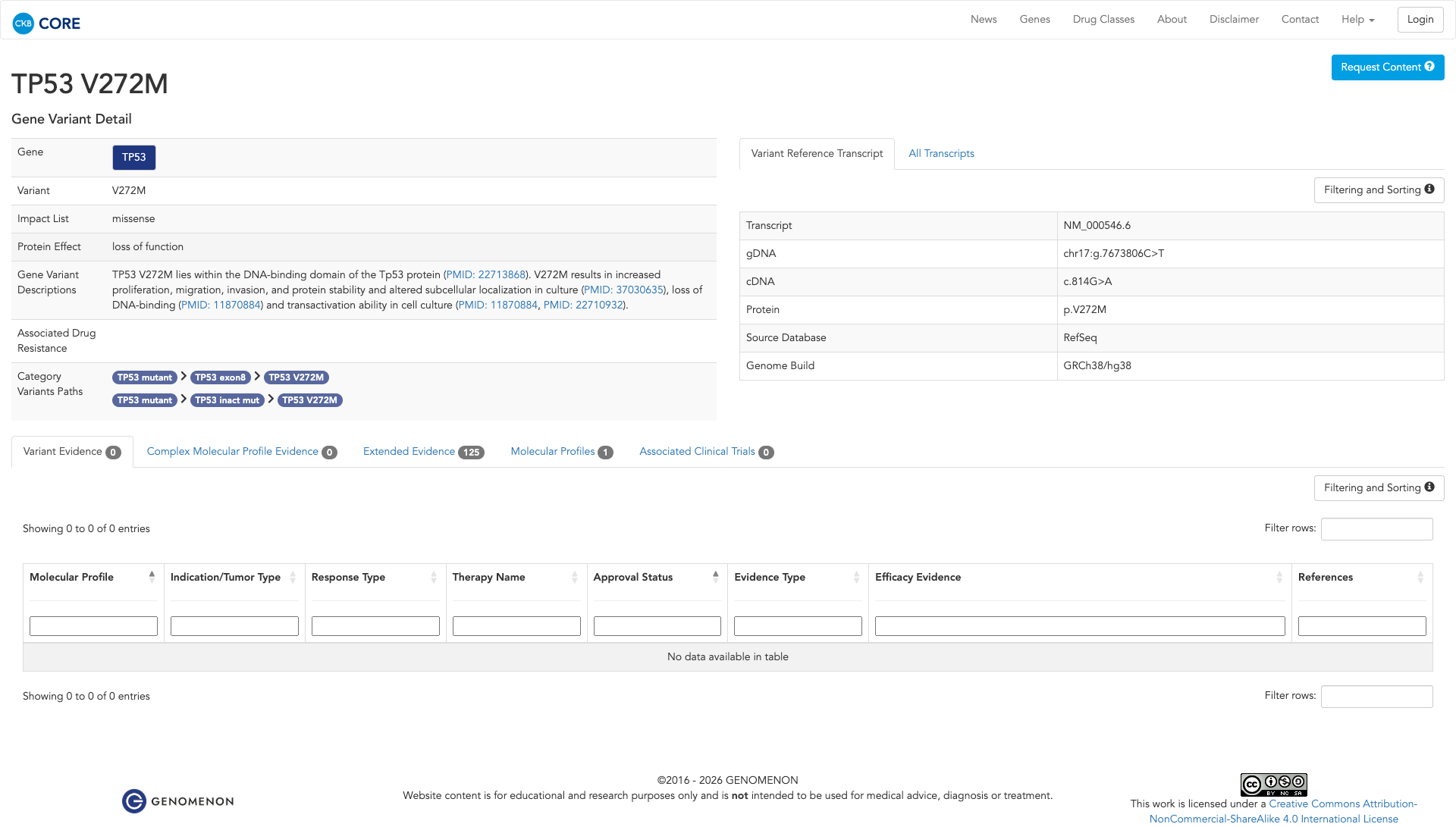
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.01 | -467 bp |
| Donor Loss (DL) | 0.0 | 1 bp |
| Acceptor Gain (AG) | 0.01 | 34 bp |
| Donor Gain (DG) | 0.0 | -105 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, PVS1 applies only to null variants predicted to undergo NMD or critical splice site variants. This is a missense variant (V272M) and does not meet PVS1 specifications. Therefore, PVS1 is not applied.
PS1 (Not Applied)
According to VCEP guidelines, PS1 applies when a different nucleotide change results in the same amino acid change as a previously established pathogenic variant. There is no record of a different nucleotide change producing V272M. Therefore, PS1 is not applied.
PS2 (Not Applied)
According to VCEP guidelines, PS2 applies based on de novo occurrence with confirmed parental testing. No de novo or parental data are available for this case. Therefore, PS2 is not applied.
PS3 (Not Applied)
According to VCEP guidelines, PS3 requires specific functional assay data (e.g., non‐functional on Kato et al. data AND LOF on another assay) to be applied. The available functional evidence is not from the designated VCEP assays. Therefore, PS3 is not applied.
PS4 (Not Applied)
According to VCEP guidelines, PS4 applies based on proband case‐level observations meeting defined point thresholds. No case‐level or proband data are provided. Therefore, PS4 is not applied.
PM1 (Not Applied)
According to VCEP guidelines, PM1 (Moderate) applies to missense variants at TP53 codons 175, 245, 248, 249, 273, or 282. V272M is at codon 272, which is not one of the specified hotspot codons. Therefore, PM1 is not applied.
PM2 (Supporting)
According to VCEP guidelines, PM2 applies at Supporting strength for variants with allele frequency <0.00003 in gnomAD. The variant’s MAF is 0.00000399 (<0.00003). Therefore, PM2 is applied at Supporting strength because the variant is extremely rare in population databases per VCEP threshold.
PM3 (Not Applied)
According to standard ACMG guidelines, PM3 applies to variants observed in trans with a pathogenic variant in recessive disorders. TP53‐related disease is dominant and no trans observations are relevant. Therefore, PM3 is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, PM4 applies to protein length changes such as in‐frame indels. This is a missense substitution with no change in protein length. Therefore, PM4 is not applied.
PM5 (Not Applied)
According to VCEP guidelines, PM5 applies when ≥1 different pathogenic missense variant has been observed at the same residue. There are no other pathogenic variants reported at codon 272. Therefore, PM5 is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines, PM6 applies to assumed de novo variants without confirmation of paternity/maternity. No de novo or parental data are available. Therefore, PM6 is not applied.
PP1 (Not Applied)
According to VCEP guidelines, PP1 (Supporting) requires segregation in ≥3 meioses; higher strengths require more segregation. No family segregation data are available. Therefore, PP1 is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, PP2 is for missense variants in genes with low benign missense variation and a common disease mechanism. TP53 has both benign and pathogenic missense variation and no specific VCEP recommendation for PP2. Therefore, PP2 is not applied.
PP3 (Not Applied)
According to VCEP guidelines, PP3 requires specific computational scores (e.g., aGVGD and BayesDel thresholds). Those values are not available. Therefore, PP3 is not applied.
PP4 (Not Applied)
According to standard ACMG guidelines, PP4 applies when a patient’s phenotype highly specific for a disease is observed. No phenotype data are provided. Therefore, PP4 is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, PP5 is for assertions from reputable sources without available evidence for independent evaluation and is discouraged. Therefore, PP5 is not applied.
BA1 (Not Applied)
According to VCEP guidelines, BA1 (Stand Alone) requires filtering allele frequency ≥0.001. The variant’s frequency is ≪0.001. Therefore, BA1 is not applied.
BS1 (Not Applied)
According to VCEP guidelines, BS1 (Strong) requires filtering allele frequency ≥0.0003. The variant’s frequency is ≪0.0003. Therefore, BS1 is not applied.
BS2 (Not Applied)
According to VCEP guidelines, BS2 requires observation in ≥2 unrelated unaffected older females. No such data are available. Therefore, BS2 is not applied.
BS3 (Not Applied)
According to VCEP guidelines, BS3 requires well‐established functional assays showing preserved function. The available functional data show loss of function but are not from specified VCEP assays; in any case, there is no evidence of normal function. Therefore, BS3 is not applied.
BS4 (Not Applied)
According to VCEP guidelines, BS4 requires lack of segregation in affected family members. No segregation data are available. Therefore, BS4 is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, BP1 applies to missense variants in genes where only truncating variants cause disease. TP53 has a well‐established missense mechanism. Therefore, BP1 is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, BP2 applies to observations in cis with a pathogenic variant in dominant disorders. No such data are available. Therefore, BP2 is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, BP3 applies to in‐frame deletions/insertions in repetitive regions. This is a missense substitution. Therefore, BP3 is not applied.
BP4 (Not Applied)
According to VCEP guidelines, BP4 requires benign computational scores (e.g., BayesDel ≤ –0.008 and SpliceAI <0.2). Those scores are not provided. Therefore, BP4 is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, BP5 applies when a variant is found in a case with an alternate molecular cause. No such context is provided. Therefore, BP5 is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, BP6 applies to assertions of benign by reputable sources without available evidence. No such assertion exists. Therefore, BP6 is not applied.
BP7 (Not Applied)
According to VCEP guidelines, BP7 applies to synonymous or intronic variants with no splicing impact. This is a missense variant. Therefore, BP7 is not applied.