Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_001754.3 | Alternative | 6190 nt | 400–1842 |
| NM_001754.4 | RefSeq Select | 5967 nt | 191–1633 |
| NM_001754.5 | MANE Select | 5971 nt | 195–1637 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Open""
COSMIC Somatic Evidence
Open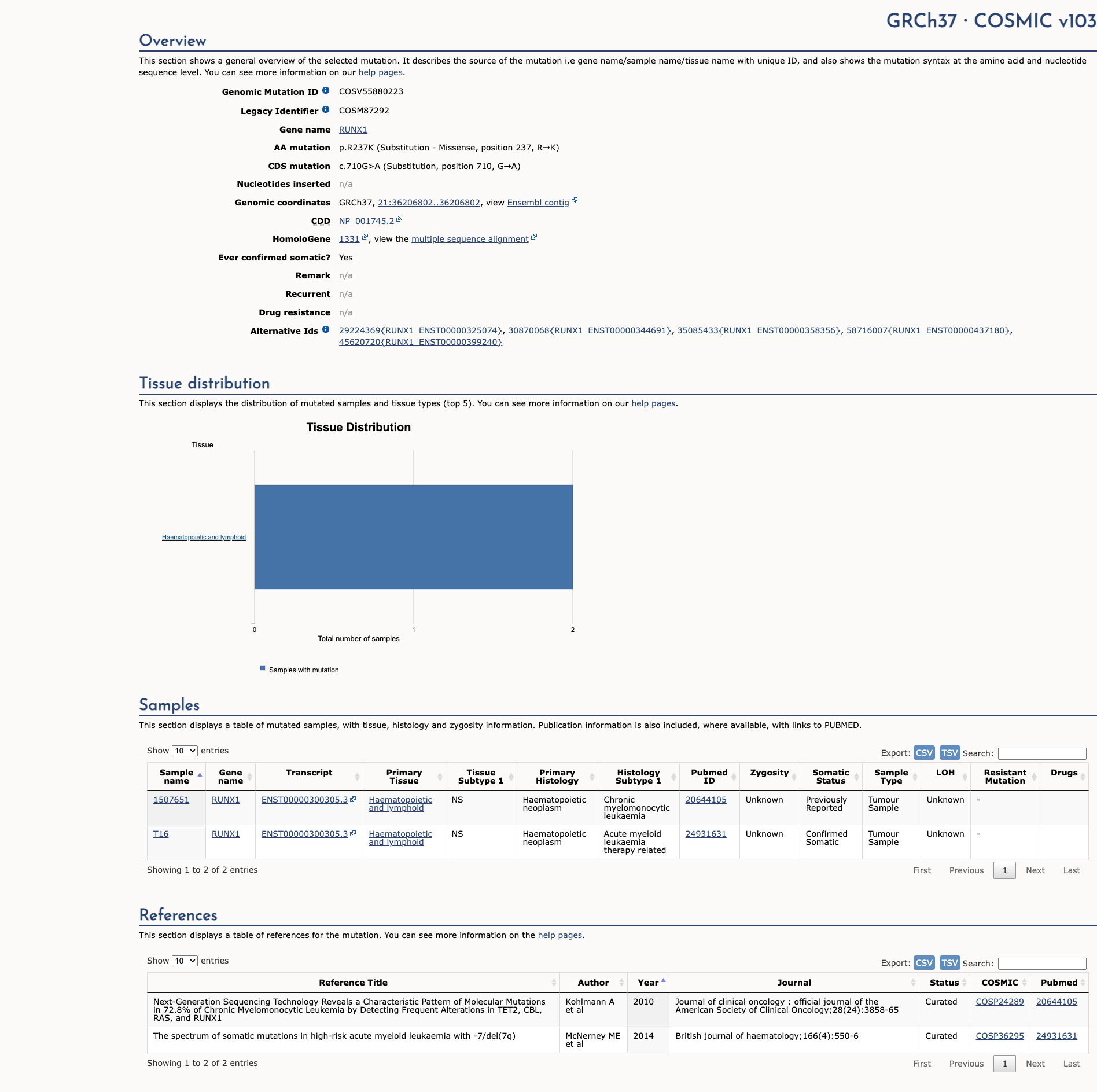
Functional Impact & Domains
Functional Domain
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.04 | -212 bp |
| Donor Loss (DL) | 0.27 | -95 bp |
| Acceptor Gain (AG) | 0.01 | 96 bp |
| Donor Gain (DG) | 0.7 | -1 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, the rule for PVS1 is: 'Very Strong Per modified RUNX1 PVS1 decision tree for SNVs and CNVs and table of splicing effects.' The evidence shows the variant is a missense change (R237K), not a null variant. Therefore, this criterion is not applied.
PS1 (Not Applied)
According to VCEP guidelines, the rule for PS1 is: 'Strong: Same amino acid change as a previously established pathogenic variant regardless of nucleotide change.' There is no reported pathogenic variant resulting in R237K. Therefore, this criterion is not applied.
PS2 (Not Applied)
According to VCEP guidelines, the rule for PS2 is: 'Moderate: Phenotypic specificity category ... For each proven de novo case give 0.5 points; assumed de novo give 0.25.' There is no de novo information. Therefore, this criterion is not applied.
PS3 (Not Applied)
According to VCEP guidelines, the rule for PS3 is: 'Strong: Transactivation assays demonstrating altered transactivation <20% of wt AND data from a secondary assay demonstrating altered function.' No functional assay data are available for R237K. Therefore, this criterion is not applied.
PS4 (Not Applied)
According to VCEP guidelines, the rule for PS4 is: 'Strong: ≥4 probands meeting RUNX1‐phenotypic criteria; Moderate: 2–3 probands; Supporting: 1 proband.' No case‐level data are available. Therefore, this criterion is not applied.
PM1 (Not Applied)
According to VCEP guidelines, the rule for PM1 (Moderate) is: 'Variant affecting one of the following residues within the RHD: R107, K110, A134, R162, R166, S167, R169, G170, K194, T196, D198, R201, R204.' R237 lies outside the RHD region. Therefore, this criterion is not applied.
PM2 (Supporting)
According to VCEP guidelines, the rule for PM2 (Supporting) is: 'Variant must be completely absent from all population databases.' The variant R237K is absent from gnomAD and other control databases. Therefore, this criterion is applied at Supporting strength.
PM3 (Not Applied)
According to standard ACMG guidelines, the rule for PM3 is: 'For recessive disorders, detected in trans with a pathogenic variant.' RUNX1‐related disease is dominant and no in trans data exist. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to VCEP guidelines, the rule for PM4 is: 'Moderate: In-frame indel affecting key RHD residues; Supporting: other RHD in-frame indels.' This is a missense substitution. Therefore, this criterion is not applied.
PM5 (Not Applied)
According to VCEP guidelines, the rule for PM5 (Moderate) is: 'Missense change at a residue with a different missense pathogenic change.' No other pathogenic missense at residue 237 has been reported. Therefore, this criterion is not applied.
PM6 (Not Applied)
According to VCEP guidelines, the rule for PM6 is: 'Moderate: points for assumed de novo cases.' No de novo data are available. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to VCEP guidelines, the rule for PP1 (Supporting) is: '3–4 meioses co-segregation with disease.' No family segregation data are available. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, the rule for PP2 is: 'Missense variant in a gene with low rate of benign missense variation.' Insufficient evidence on RUNX1 missense constraint. Therefore, this criterion is not applied.
PP3 (Supporting)
According to VCEP guidelines, the rule for PP3 (Supporting) is: 'For missense, SpliceAI ≥0.38 OR REVEL ≥0.88.' SpliceAI predicts a donor gain score of 0.70 (≥0.38). Therefore, this criterion is applied at Supporting strength.
PP4 (Not Applied)
According to standard ACMG guidelines, the rule for PP4 is: 'Patient’s phenotype highly specific for a gene.' No phenotype data provided. Therefore, this criterion is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, the rule for PP5 is: 'Reputable source reports variant as pathogenic.' The variant is not present in ClinVar or other sources. Therefore, this criterion is not applied.
BA1 (Not Applied)
According to VCEP guidelines, the rule for BA1 (Stand Alone) is: 'MAF ≥0.0015 in any continental population.' The variant is absent. Therefore, this criterion is not applied.
BS1 (Not Applied)
According to VCEP guidelines, the rule for BS1 (Strong) is: 'MAF between 0.00015 and 0.0015.' The variant is absent. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to standard ACMG guidelines, the rule for BS2 is: 'Observed in healthy adults for a dominant disorder.' No such observations. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to VCEP guidelines, the rule for BS3 (Strong) is: 'Normal transactivation (80–115% of wt) AND normal secondary assay.' No functional data available. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to standard ACMG guidelines, the rule for BS4 is: 'Lack of segregation in ≥2 informative meioses.' No segregation data. Therefore, this criterion is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, the rule for BP1 is: 'Missense in gene where only truncating variants cause disease.' RUNX1 has known pathogenic missense variants. Therefore, this criterion is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, the rule for BP2 is: 'Observed in cis or trans with a pathogenic variant.' No such data. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, the rule for BP3 is: 'In-frame indel in repetitive region without known function.' Not applicable for a missense change. Therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP guidelines, the rule for BP4 (Supporting) is: 'REVEL <0.50 AND SpliceAI ≤0.20.' REVEL is 0.72 and SpliceAI is 0.70. Therefore, this criterion is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, the rule for BP5 is: 'Variant found in a case with an alternate molecular basis for disease.' No such data. Therefore, this criterion is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, the rule for BP6 is: 'Reputable source reports variant as benign.' No such reports. Therefore, this criterion is not applied.
BP7 (Not Applied)
According to VCEP guidelines, the rule for BP7 (Supporting) is: 'Synonymous or intronic variant with SpliceAI ≤0.20 and low conservation.' This is a missense variant. Therefore, this criterion is not applied.