Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000249.3 | RefSeq Select | 2662 nt | 199–2469 |
| NM_000249.2 | Alternative | 2524 nt | 61–2331 |
| NM_000249.4 | MANE Select | 2494 nt | 31–2301 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe c.1513delA pathogenic mutation, located in coding exon 13 of the MLH1 gene, results from a deletion of one nucleotide at nucleotide position 1513, causing a translational frameshift with a predicted alternate stop codon (p.S505Vfs*3). This alteration is expected to result in loss of function by premature protein truncation or nonsense-mediated mRNA decay. As such, this alteration is interpreted as a disease-causing mutation.
"This variant has been reported in ClinVar as Pathogenic (1 clinical laboratories)."
COSMIC Somatic Evidence
Open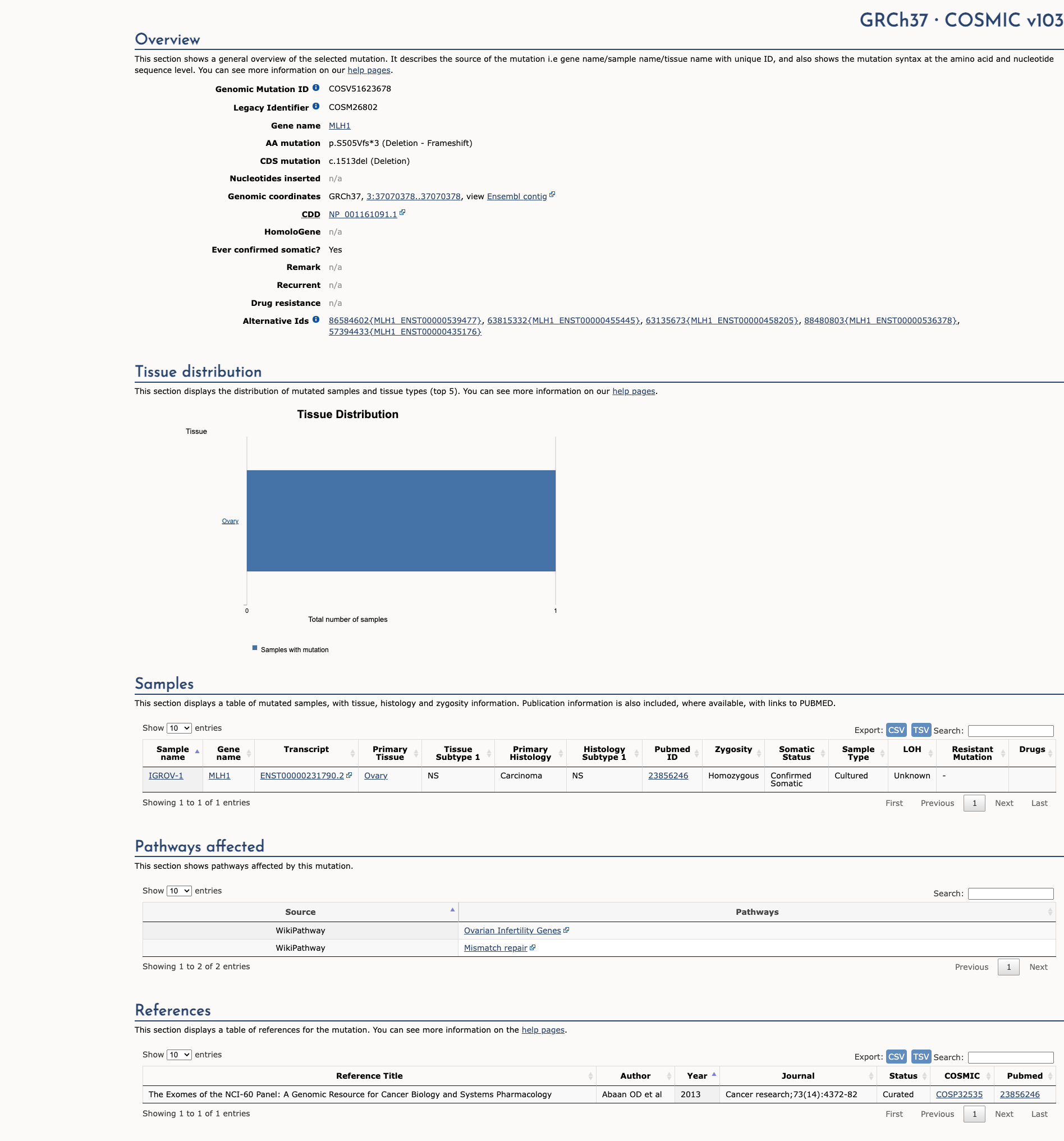
Functional Impact & Domains
Functional Domain
The MLH1 S505Vfs*3 variant is a truncating mutation that results in a premature stop codon, leading to the loss of the C-terminal domain of the MLH1 protein. Functional evidence indicates that this truncation impairs the protein's ability to bind PMS2, disrupting DNA mismatch repair and resulting in a loss of MLH1 function. This loss of function is associated with increased mutation burden and tumor development, as observed in MLH1-deficient mice. Therefore, the functional evidence supports a damaging effect of the MLH1 S505Vfs*3 variant.
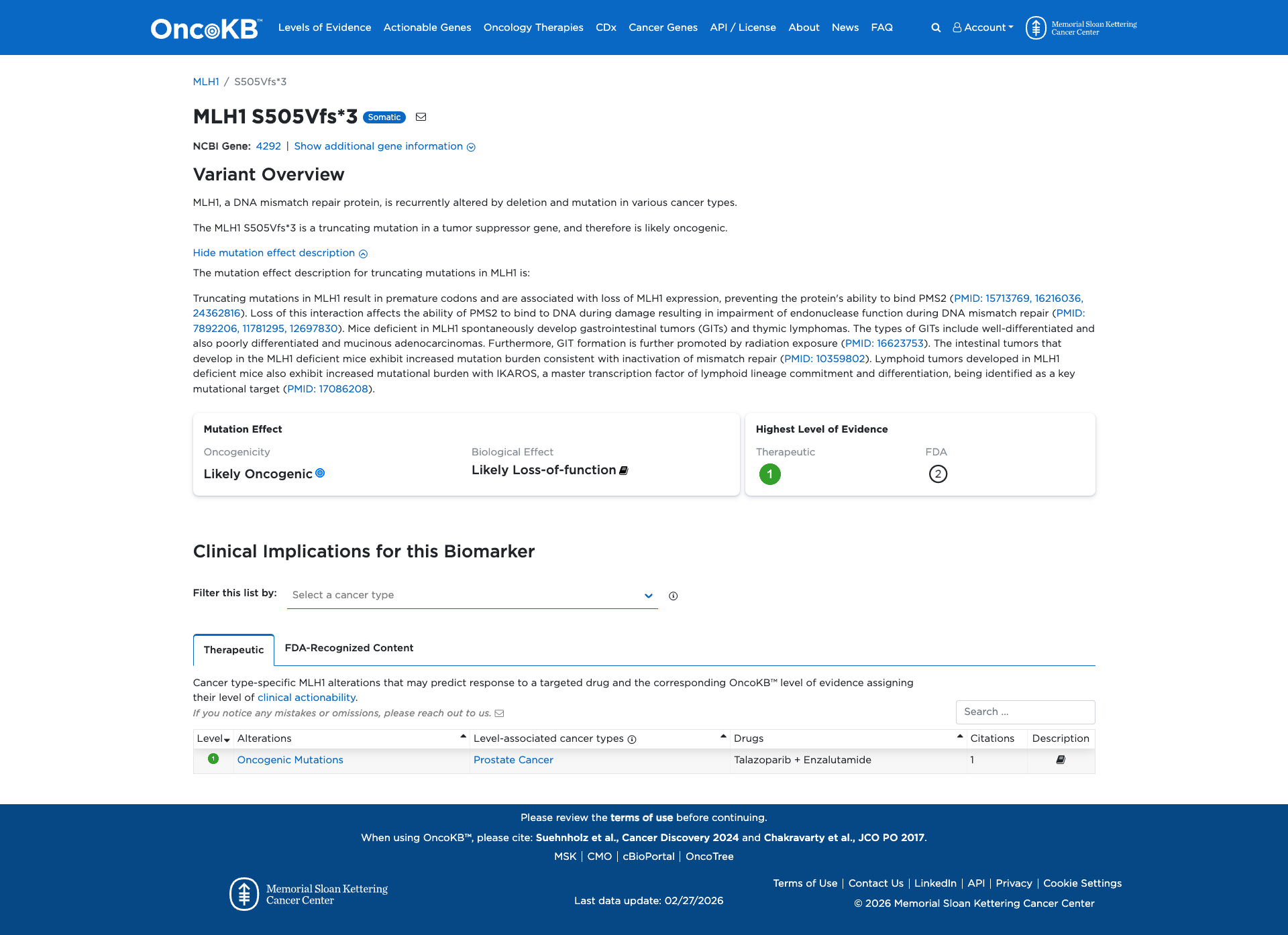
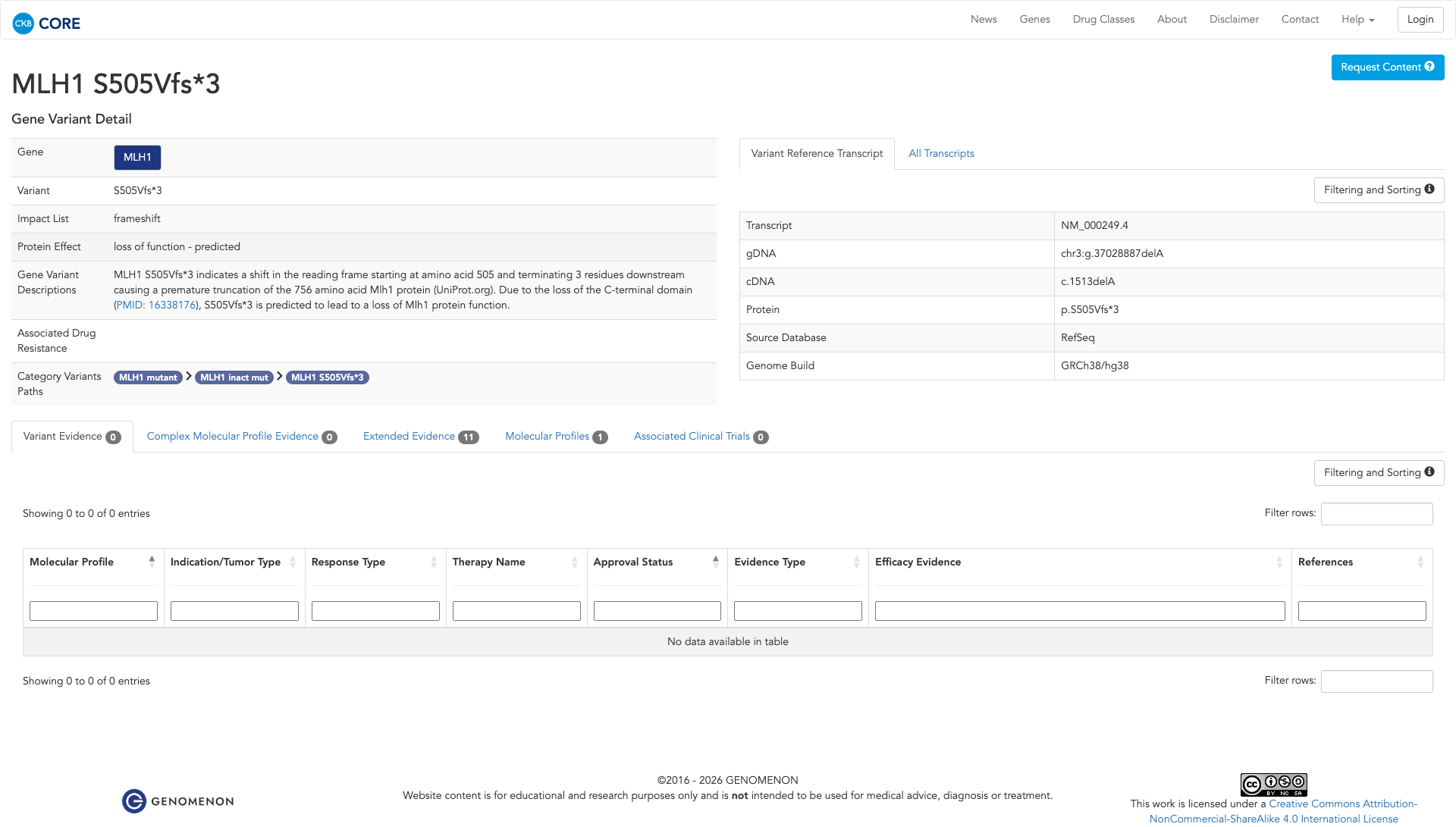
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -102 bp |
| Donor Loss (DL) | 0.0 | 1 bp |
| Acceptor Gain (AG) | 0.0 | 19 bp |
| Donor Gain (DG) | 0.0 | -32 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Very Strong)
According to VCEP guidelines, the rule for PVS1 is: "Very Strong Nonsense/frameshift variant introducing Premature Termination Codon (PTC) at or before codon 753 in MLH1". The evidence for this variant shows: NM_000249.4:c.1513del causes a frameshift at codon 505 and introduces a stop codon at codon 507, which is before codon 753. Therefore, this criterion is applied at Very Strong strength because it is a null frameshift variant predicted to undergo NMD in a gene where loss of function is a known mechanism of disease.
PS1 (Not Applied)
According to VCEP guidelines, the rule for PS1 is: "Strong A predicted missense substitution that encodes the same amino acid change with a different underlying nucleotide change previously established by this VCEP as Pathogenic (not a predicted or confirmed splice defect)". The evidence for this variant shows: it is a frameshift variant (S505Vfs*3), not a missense substitution encoding the same amino acid change. Therefore, this criterion is not applied.
PS2 (Not Applied)
According to VCEP guidelines, the rule for PS2 is: "Very Strong ≥ 4 de novo points; Strong 2–3 de novo points; Moderate 1 de novo point; Supporting 0.5 de novo points". The evidence for this variant shows: no information on de novo occurrence or parental testing. Therefore, this criterion is not applied.
PS3 (Moderate)
According to VCEP guidelines, the rule for PS3 is: "Moderate Calibrated functional assays with functional odds for pathogenicity >4.3 and ≤18.7 OR MMR function defect following functional assay flowchart* OR Variants with monoallelic expression: complete loss of expression (<10% of wild-type in cDNA without puromycin) of the variant allele". The evidence for this variant shows: in vitro and in vivo studies demonstrate that the truncation disrupts MLH1 binding to PMS2 and abolishes mismatch repair function. Therefore, this criterion is applied at Moderate strength because the variant causes an MMR function defect.
PS4 (Not Applied)
According to standard ACMG guidelines, PS4 is: "The prevalence of the variant in affected individuals is significantly increased compared with the prevalence in controls". The evidence for this variant shows: no case-control or statistical data are available. Therefore, this criterion is not applied.
PM1 (Not Applied)
According to standard ACMG guidelines, PM1 is: "Located in a mutational hot spot and/or critical and well-established functional domain without benign variation". The evidence for this variant shows: no data indicating location in a mutational hot spot or critical domain beyond LOF. Therefore, this criterion is not applied.
PM2 (Supporting)
According to VCEP guidelines, the rule for PM2 is: "Supporting Absent/extremely rare (<1 in 50,000 alleles) in gnomAD v4 dataset". The evidence for this variant shows: absent from gnomAD. Therefore, this criterion is applied at Supporting strength because the variant is absent from population databases.
PM3 (Not Applied)
According to VCEP guidelines, the rule for PM3 is: "Very Strong ≥4 points; Strong ≥2 and <4 points; Moderate ≥1 and <2 points; Supporting =0.5 points for evidence of in trans occurrence in recessive disease". The evidence for this variant shows: no evidence of occurrence in trans with another pathogenic MLH1 variant. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, PM4 is: "Protein length changes due to in-frame deletions/insertions in a non-repeat region or stop-loss variants". The evidence for this variant shows: this is a frameshift leading to truncation, not an in-frame change or stop-loss. Therefore, this criterion is not applied.
PM5 (Not Applied)
According to VCEP guidelines, the rule for PM5 is: "Moderate Missense change at an amino acid residue where a different missense change was classified by this VCEP as Pathogenic". The evidence for this variant shows: it is a frameshift variant, not a missense change. Therefore, this criterion is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines, PM6 is: "Assumed de novo, without confirmation of paternity and maternity". The evidence for this variant shows: no de novo data or parental testing. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to VCEP guidelines, the rule for PP1 is: "Strong, Moderate, or Supporting co-segregation based on Bayes likelihood ratio". The evidence for this variant shows: no family segregation data. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, PP2 is: "Missense variant in a gene with a low rate of benign missense variation and where missense variants are a common mechanism of disease". The evidence for this variant shows: this is a frameshift variant, not a missense. Therefore, this criterion is not applied.
PP3 (Not Applied)
According to VCEP guidelines, the rule for PP3 is: "Supporting Missense variant with HCI-prior probability >0.68 & ≤0.81 OR predicted splice defect for non-canonical nucleotides with SpliceAI delta ≥0.2". The evidence for this variant shows: SpliceAI score 0 and no missense; computational evidence does not support a deleterious effect. Therefore, this criterion is not applied.
PP4 (Not Applied)
According to VCEP guidelines, the rule for PP4 is: "Supporting phenotype data such as MSI-H tumors consistent with MLH1 variant". The evidence for this variant shows: no specific tumor phenotype or MSI/MMR data. Therefore, this criterion is not applied.
PP5 (Supporting)
According to standard ACMG guidelines, PP5 is: "Supporting Reputable source recently reports variant as pathogenic, but the evidence is not available to the laboratory to perform an independent evaluation". The evidence for this variant shows: ClinVar lists it as Pathogenic by one clinical laboratory. Therefore, this criterion is applied at Supporting strength because of the credible external assertion.
BA1 (Not Applied)
According to VCEP guidelines, the rule for BA1 is: "Stand Alone GnomAD v4 Grpmax filtering allele frequency ≥0.001 (0.1%) and variant is excluded as founder pathogenic variant". The evidence for this variant shows: allele frequency is 0%. Therefore, this criterion is not applied.
BS1 (Not Applied)
According to VCEP guidelines, the rule for BS1 is: "Strong GnomAD v4 Grpmax filtering allele frequency ≥0.0001 and <0.001". The evidence for this variant shows: allele frequency is 0%. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to VCEP guidelines, the rule for BS2 is: "Strong co-occurrence in trans with a known pathogenic variant in the same gene in a patient without CMMRD". The evidence for this variant shows: no such co-occurrence data. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to VCEP guidelines, the rule for BS3 is: "Strong Calibrated functional assays with functional odds for pathogenicity ≤0.05 OR synonymous/intronic with no mRNA aberration". The evidence for this variant shows: functional assays demonstrate loss of function, not normal function. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to VCEP guidelines, the rule for BS4 is: "Strong Lack of co-segregation with disease in pedigree(s) with Bayes LR <0.05". The evidence for this variant shows: no segregation data. Therefore, this criterion is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, BP1 is: "Missense variant in a gene for which primarily truncating variants are known to cause disease". The evidence for this variant shows: it is a truncating variant, not missense. Therefore, this criterion is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, BP2 is: "Observed in trans with a pathogenic variant in a dominant gene or in cis with a pathogenic variant". The evidence for this variant shows: no data on cis/trans observations. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, BP3 is: "In-frame deletions/insertions in a repetitive region without a known function". The evidence for this variant shows: this is a frameshift variant, not an in-frame change in a repeat region. Therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP guidelines, the rule for BP4 is: "Supporting Missense variant with HCI-prior probability <0.11 OR for intronic/synonymous SpliceAI delta ≤0.1". The evidence for this variant shows: it is a frameshift variant; computational data do not support a benign prediction. Therefore, this criterion is not applied.
BP5 (Not Applied)
According to VCEP guidelines, the rule for BP5 is: "Supporting presence of alternative molecular basis for disease in multiple tumors". The evidence for this variant shows: no tumor co-occurrence data. Therefore, this criterion is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, BP6 is: "Reputable source reports variant as benign, but evidence is not available". The evidence for this variant shows: no benign assertion. Therefore, this criterion is not applied.
BP7 (Not Applied)
According to VCEP guidelines, the rule for BP7 is: "Supporting A synonymous (silent) or intronic variant at or beyond -21/+7 (5′/3′ exonic)". The evidence for this variant shows: it is a frameshift in an exon, not a silent or intronic change. Therefore, this criterion is not applied.