Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis variant is considered pathogenic. This variant has been reported in multiple individuals with clinical features of gene-specific disease [PMID: 16633321]. Functional studies indicate this variant impacts protein function [PMID: 12509279, 17530187]. This variant is expected to disrupt protein structure [Myriad internal data].
This submission and the accompanying classification are no longer maintained by the submitter. For more information on current observations and classification, please contact variantquestions@myriad.com.
The p.H179Y pathogenic mutation (also known as c.535C>T), located in coding exon 4 of the TP53 gene, results from a C to T substitution at nucleotide position 535. The histidine at codon 179 is replaced by tyrosine, an amino acid with similar properties. This mutation has been reported in the germline of multiple individuals with Li-Fraumeni syndrome (Lefrou L et al. Gastroenterol. Clin. Biol., 2006 Mar;30:484-6; Bougeard G et al. J. Clin. Oncol., 2015 Jul;33:2345-52; Zerdoumi Y et al. Hum. Mol. Genet., 2017 Jul;26:2591-2602). Numerous functional assays conducted in yeast and mammalian cells have shown the p.Y179H variant has decreased transactivation activity compared to wild type, and exerts a dominant negative effect (Kato S et al. Proc Natl Acad Sci USA. 2003 Jul 8;100(14):8424-9, Dearth LR et al. Carcinogenesis 2007 Feb; 28(2):289-98; Scian MJ et al. Oncogene, 2004 May;23:4430-43). Further functional studies indicate this mutation has oncogenic properties through upregulation of genes involved in cell cycle progression and proliferation (Ko JL et al. DNA Repair Amst; Yang D et al. Mol. Cell. Biochem., 2007 Oct;304:219-26; Scian MJ et al. Oncogene, 2004 May;23:4430-43). Studies conducted in human cell lines indicate this alteration is deficient at growth suppression and has a dominant negative effect (Kotler E et al. Mol.Cell. 2018 Jul;71:178-190.e8; Giacomelli AO et al. Nat. Genet. 2018 Oct;50:1381-1387). This alteration is located in the DNA binding domain, and is one of four residues necessary to bind the zinc molecule that stabilizes the beta sheet structure of the p53 protein (Martin AC et al. Hum. Mutat. 2002 Feb; 19(2):149-64). This alteration has been observed numerous times as a somatic mutation in the cancerhotspots.org database (Chang MT et al. Cancer Discov. 2018 02;8:174-183). In addition, another alteration at this same position, p.H179Q was identified as a de novo mutation in a patient with two Li-Fraumeni core cancers (Gonzalez KD et al. J. Med. Genet. 2009 Oct; 46(10):689-93). Based on the supporting evidence, this alteration is interpreted as a disease-causing mutation.
This sequence change replaces histidine, which is basic and polar, with tyrosine, which is neutral and polar, at codon 179 of the TP53 protein (p.His179Tyr). This variant is not present in population databases (gnomAD no frequency). This missense change has been observed in individual(s) with a personal and/or family history of Li-Fraumeni syndrome (PMID: 16633321, 18511570, 25433984). In at least one individual the variant was observed to be de novo. ClinVar contains an entry for this variant (Variation ID: 127815). Invitae Evidence Modeling incorporating data from in vitro experimental studies (PMID: 12826609, 29979965, 30224644) indicates that this missense variant is expected to disrupt TP53 function with a positive predictive value of 97.5%. Experimental studies have shown that this missense change affects TP53 function (PMID: 11896595, 12509279, 12726864, 12826609, 17530187, 26497680). For these reasons, this variant has been classified as Pathogenic.
"This variant has been reported in ClinVar as Pathogenic (5 clinical laboratories) and as Likely pathogenic (5 clinical laboratories)."
COSMIC Somatic Evidence
Open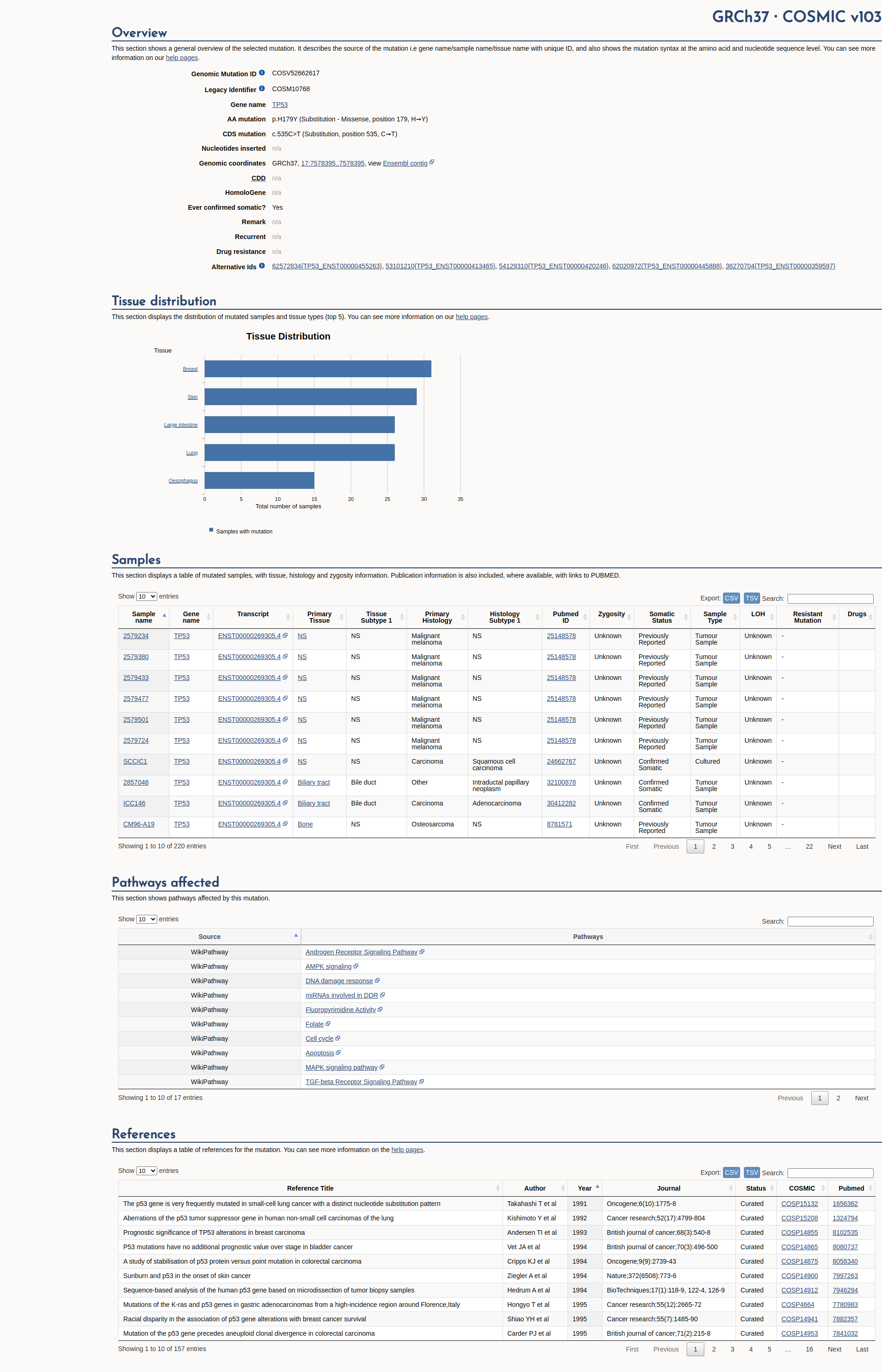
Functional Impact & Domains
Functional Domain
The TP53 H179Y variant has been functionally characterized and demonstrates a damaging effect. In vitro studies indicate that this mutation results in a loss of function of the TP53 protein. This is evidenced by decreased transactivation of TP53 target genes, reduced growth suppression activity, and increased cell proliferation. Additionally, the variant leads to increased expression of cyclin A1 and CDK4, increased cell size, and enhanced cell cycle progression from G1 to S phase. These findings collectively support the conclusion that TP53 H179Y is an inactivating mutation with oncogenic potential.
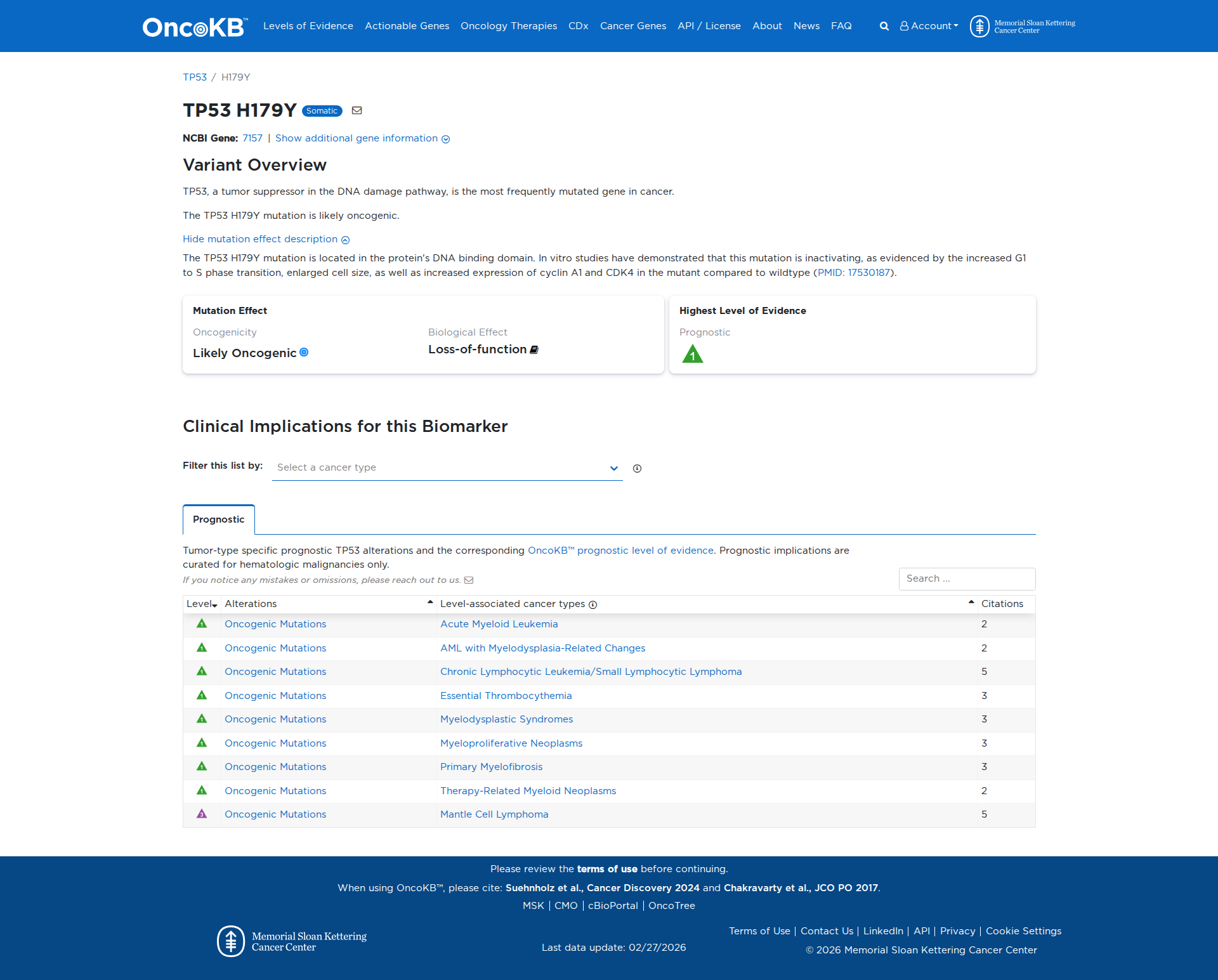
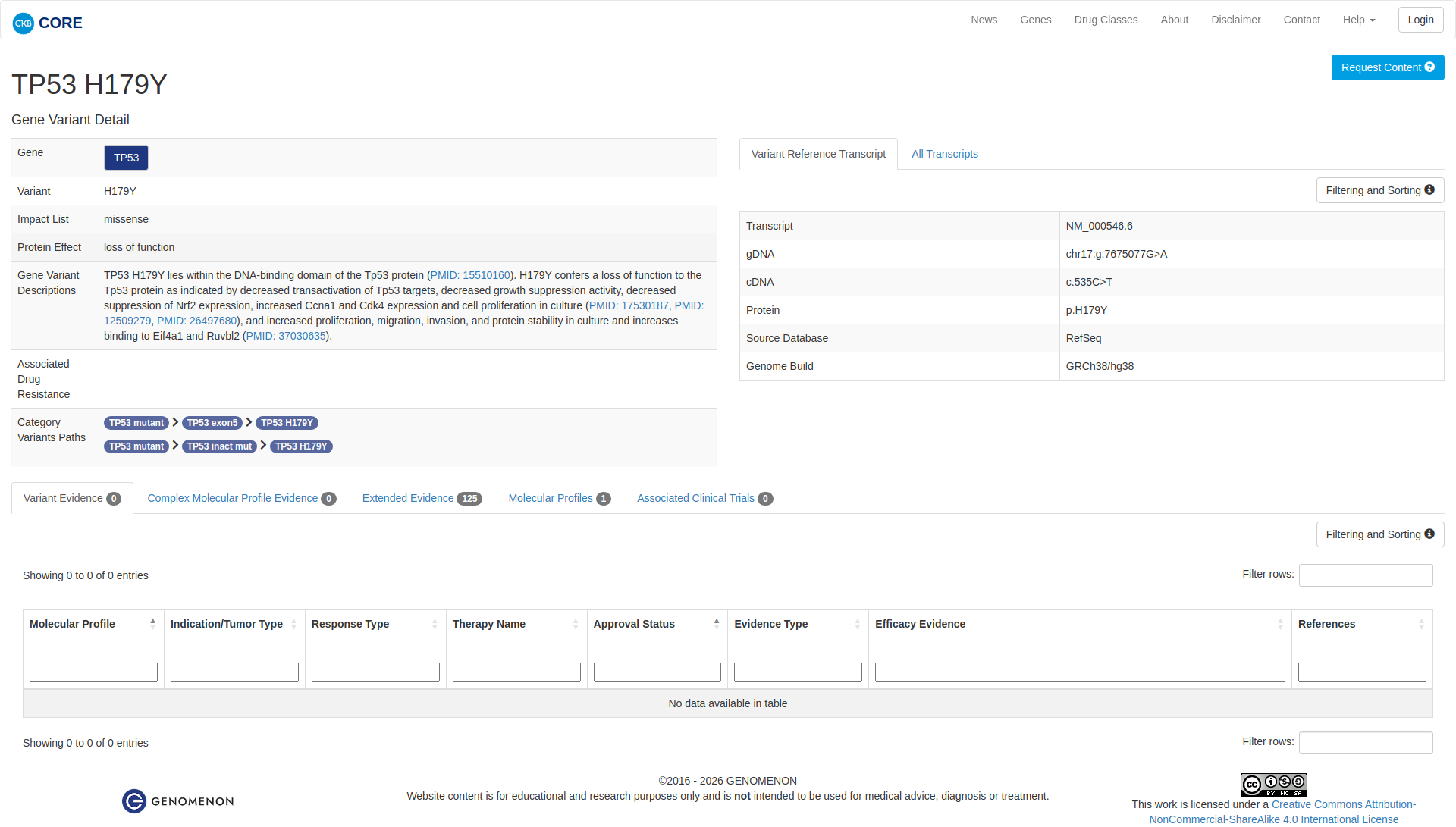
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 159 bp |
| Donor Loss (DL) | 0.0 | -223 bp |
| Acceptor Gain (AG) | 0.0 | -106 bp |
| Donor Gain (DG) | 0.0 | 22 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, PVS1 applies to null variants causing loss of function (nonsense, frameshift, canonical splice, initiation codon, or multi‐exon deletion). The variant H179Y is a missense change, not a null variant. Therefore, PVS1 is not applied.
PS1 (Not Applied)
According to VCEP guidelines, PS1 applies when the same amino acid change as a previously established pathogenic variant occurs. H179Y has no identical amino acid change previously classified as pathogenic in TP53. Therefore, PS1 is not applied.
PS2 (Not Applied)
According to VCEP guidelines, PS2 applies to confirmed de novo observations with parental confirmation. No de novo data are available for H179Y. Therefore, PS2 is not applied.
PS3 (Not Applied)
According to VCEP guidelines, PS3 requires non‐functional Kato et al. data AND loss–of–function on another assay for Strong strength (or defined criteria for Moderate/Supporting). Specific Kato/Giacomelli/Kotler data for H179Y are not provided. Therefore, PS3 is not applied.
PS4 (Not Applied)
According to VCEP guidelines, PS4 is based on proband counts meeting point thresholds. No case‐level counts are provided for H179Y. Therefore, PS4 is not applied.
PM1 (Not Applied)
According to VCEP guidelines, PM1 (Moderate) applies to missense variants at codons 175, 245, 248, 249, 273, or 282. H179Y is at codon 179, outside these hotspots. Therefore, PM1 is not applied.
PM2 (Supporting)
According to VCEP guidelines, PM2_Supporting applies when allele frequency in gnomAD is <0.00003. H179Y is absent from gnomAD. Therefore, PM2 is applied at Supporting strength.
PM3 (Not Applied)
According to standard ACMG guidelines, PM3 applies to variants in trans with a pathogenic variant in recessive disorders. TP53‐related Li‐Fraumeni syndrome is dominant. Therefore, PM3 is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, PM4 applies to protein length changes from in‐frame indels or stop‐loss. H179Y is a missense change. Therefore, PM4 is not applied.
PM5 (Not Applied)
According to VCEP guidelines, PM5 applies when another pathogenic missense variant at the same residue has been established. No other pathogenic variant at codon 179 is documented under VCEP specifications. Therefore, PM5 is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines, PM6 applies to assumed de novo without confirmation. No de novo data are available for H179Y. Therefore, PM6 is not applied.
PP1 (Not Applied)
According to VCEP guidelines, PP1 applies based on segregation in multiple meioses. No segregation data are available. Therefore, PP1 is not applied.
PP2 (Supporting)
According to standard ACMG guidelines, PP2 applies to missense variants in a gene with a low rate of benign missense variants and where missense is a common pathogenic mechanism. TP53 meets these criteria and H179Y is missense. Therefore, PP2 is applied at Supporting strength.
PP3 (Not Applied)
According to VCEP guidelines, PP3 requires BayesDel ≥0.16 and/or aGVGD C65 plus SpliceAI <0.2. BayesDel/aGVGD scores for H179Y are not provided, and in silico results are mixed. Therefore, PP3 is not applied.
PP4 (Not Applied)
According to standard ACMG guidelines, PP4 applies to a specific phenotype with a single genetic etiology. No phenotype data are provided. Therefore, PP4 is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, PP5 is deprecated and not recommended. Therefore, PP5 is not applied.
BA1 (Not Applied)
According to VCEP guidelines, BA1 applies to allele frequency ≥0.001 in gnomAD. H179Y is absent from gnomAD. Therefore, BA1 is not applied.
BS1 (Not Applied)
According to VCEP guidelines, BS1 applies to filtering allele frequency ≥0.0003 but <0.001. H179Y is absent from gnomAD. Therefore, BS1 is not applied.
BS2 (Not Applied)
According to VCEP guidelines, BS2 applies to observation in multiple healthy older females. No such data exist for H179Y. Therefore, BS2 is not applied.
BS3 (Not Applied)
According to VCEP guidelines, BS3 applies when functional assays demonstrate no loss of function. Existing functional data indicate loss of function, not normal function. Therefore, BS3 is not applied.
BS4 (Not Applied)
According to VCEP guidelines, BS4 applies when lack of segregation is observed. No segregation data are available. Therefore, BS4 is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, BP1 applies to missense variants in genes where only truncating variants are pathogenic. In TP53, missense is a known mechanism. Therefore, BP1 is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, BP2 applies to variants observed in trans with a pathogenic variant in a dominant gene or found in cis with a more severe variant. No such observations exist for H179Y. Therefore, BP2 is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, BP3 applies to in‐frame indels in repetitive regions. H179Y is missense and not an indel. Therefore, BP3 is not applied.
BP4 (Not Applied)
According to VCEP guidelines, BP4 requires BayesDel ≤–0.008 with SpliceAI <0.2 for moderate strength (or different BayesDel ranges for supporting). BayesDel for H179Y is not provided. Therefore, BP4 is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, BP5 applies when a variant is found in a case with an alternate molecular basis. No such cases exist. Therefore, BP5 is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, BP6 applies when a variant is reported benign by a reputable source without evidence. H179Y has no such reports. Therefore, BP6 is not applied.
BP7 (Not Applied)
According to VCEP guidelines, BP7 applies to synonymous or intronic variants with no splice impact. H179Y is missense. Therefore, BP7 is not applied.