Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis sequence change replaces serine, which is neutral and polar, with phenylalanine, which is neutral and non-polar, at codon 127 of the TP53 protein (p.Ser127Phe). This variant is not present in population databases (gnomAD no frequency). This missense change has been observed in individuals with clinical features of Li-Fraumeni syndrome and/or Li-Fraumeni syndrome (PMID: 32817165, 34240179; Invitae; external communication). It has also been observed to segregate with disease in related individuals. ClinVar contains an entry for this variant (Variation ID: 182928). Advanced modeling performed at Invitae incorporating data from internal and/or published experimental studies (PMID: 12826609, 29979965, 30224644) indicates that this missense variant is expected to disrupt TP53 function. Experimental studies have shown that this missense change affects TP53 function (PMID: 12826609, 16000567, 16322298, 29979965, 30224644). This variant disrupts the p.Ser127 amino acid residue in TP53. Other variant(s) that disrupt this residue have been determined to be pathogenic (PMID: 29470806; Invitae). This suggests that this residue is clinically significant, and that variants that disrupt this residue are likely to be disease-causing. For these reasons, this variant has been classified as Pathogenic.
This variant is considered likely pathogenic. Functional studies indicate this variant impacts protein function [PMID: 29979965]. This variant is expected to disrupt protein structure [Myriad internal data].
The p.S127F variant (also known as c.380C>T), located in coding exon 4 of the TP53 gene, results from a C to T substitution at nucleotide position 380. The serine at codon 127 is replaced by phenylalanine, an amino acid with highly dissimilar properties. This variant was detected in a cohort of 295 women considered at high risk for breast cancer (Guindalini RSC et al. Clin Cancer Res, 2019 Mar;25:1786-1794). This variant is in the DNA binding domain of the TP53 protein and is reported to have non-functional transactivation in yeast based assays (Kato S et al. Proc Natl Acad Sci U S A, 2003 Jul;100:8424-9). Studies conducted in human cell lines indicate this alteration is deficient at growth suppression and has a dominant negative effect (Kotler E et al. Mol.Cell. 2018 Jul;71:178-190.e8; Giacomelli AO et al. Nat. Genet. 2018 Oct;50:1381-1387). Another alteration at the same codon, p.S127T (c.379T>A), has been shown to also have non-functional transactivation in yeast based assays, be deficient at growth suppression, and have a dominant negative effect (Ambry internal data; Kato S et al. Proc. Natl. Acad. Sci. USA. 2003 Jul;100:8424-9; Kotler E et al. Mol.Cell. 2018 Jul;71:178-190.e8; Giacomelli AO et al. Nat. Genet. 2018 Oct;50:1381-1387). This variant is considered to be rare based on population cohorts in the Genome Aggregation Database (gnomAD). This amino acid position is highly conserved in available vertebrate species. In addition, this alteration is predicted to be deleterious by in silico analysis. Based on the majority of available evidence to date, this variant is likely to be pathogenic.
"This variant has been reported in ClinVar as Uncertain significance (1 clinical laboratories) and as Pathogenic (3 clinical laboratories) and as Likely pathogenic (5 clinical laboratories)."
COSMIC Somatic Evidence
Open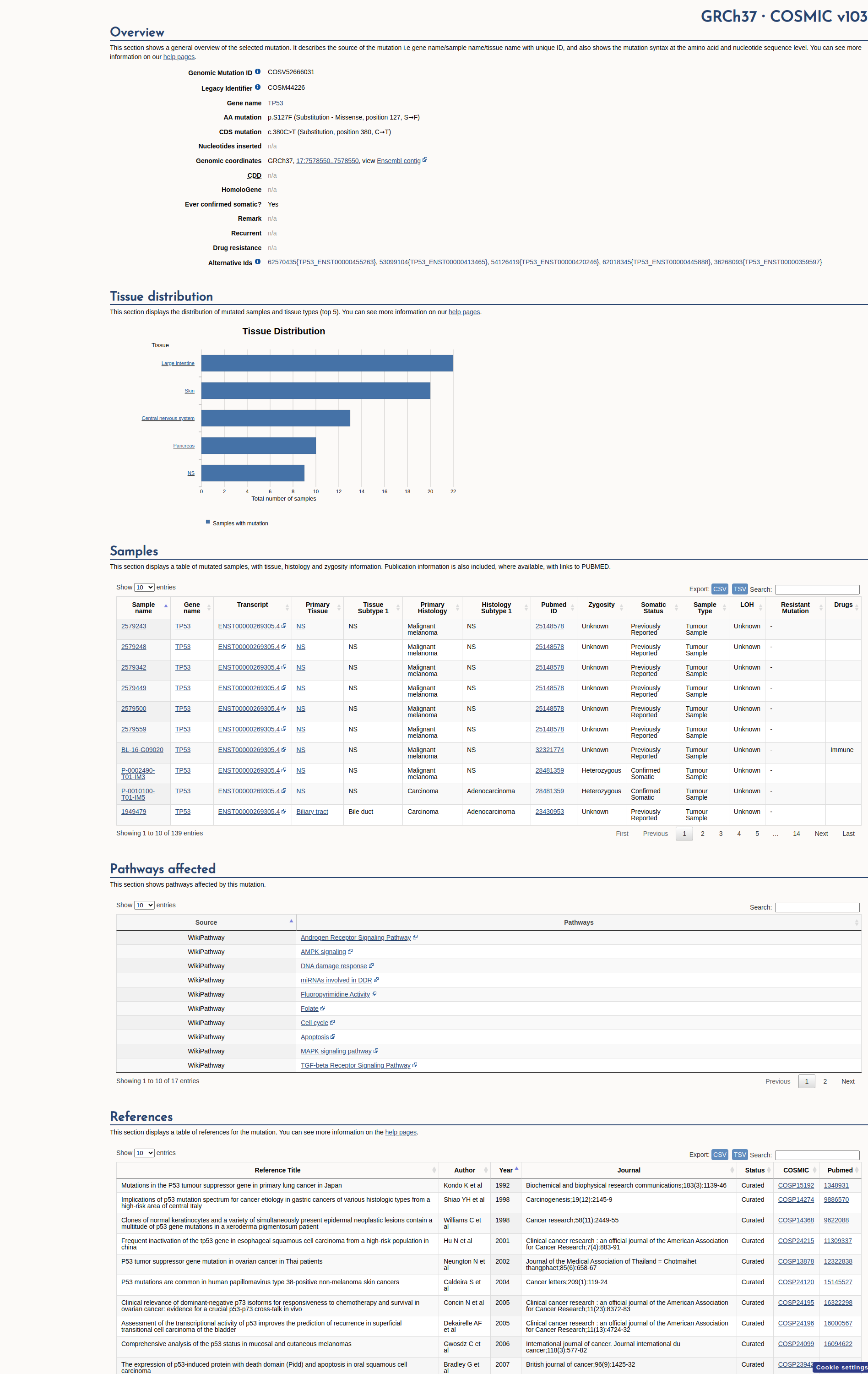
Functional Impact & Domains
Functional Domain
The TP53 S127F variant has been functionally characterized as inactivating. In vivo studies in yeast have shown a loss of transactivational activity, and in vitro studies in human cancer cell lines have demonstrated reduced growth suppression activity compared to the wildtype. These findings support a damaging effect on TP53 protein function.
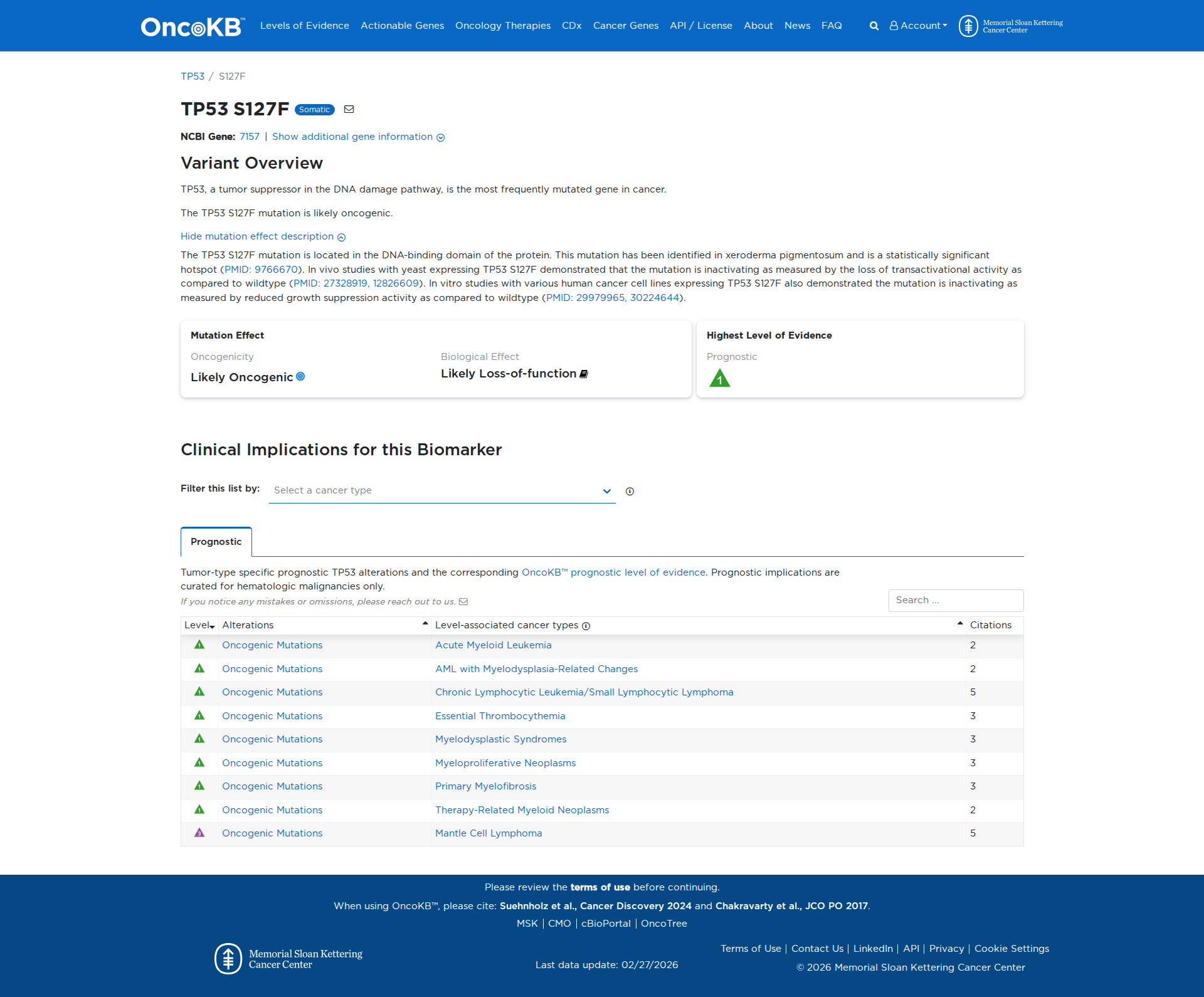
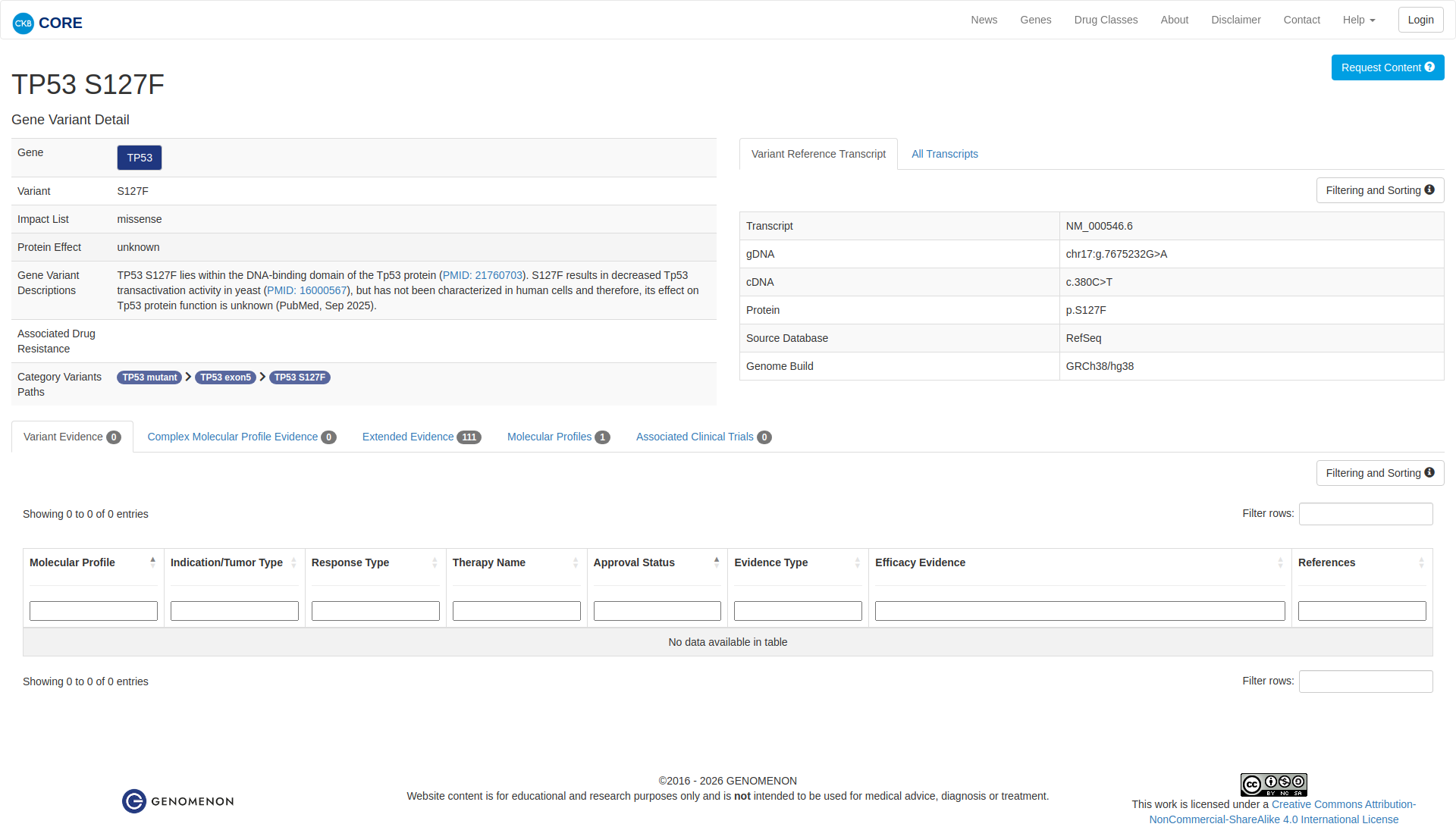
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -278 bp |
| Donor Loss (DL) | 0.0 | -378 bp |
| Acceptor Gain (AG) | 0.0 | -17 bp |
| Donor Gain (DG) | 0.0 | 104 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, PVS1 applies to null variants (nonsense, frameshift, canonical splice, initiation codon) following the TP53 decision tree. The c.380C>T (p.Ser127Phe) variant is a missense change, not predicted to cause a null allele. Therefore, PVS1 is not applied.
PS1 (Not Applied)
According to VCEP guidelines, PS1 applies when a variant results in the same amino acid change as a previously established pathogenic variant. No other variant yielding p.Ser127Phe has been classified as pathogenic. Therefore, PS1 is not applied.
PS2 (Not Applied)
According to VCEP guidelines, PS2 applies for confirmed de novo occurrences with appropriate clinical confirmation. No de novo evidence is available for this variant. Therefore, PS2 is not applied.
PS3 (Strong)
According to VCEP guidelines for TP53, PS3 at Strong strength applies when a variant is non-functional on Kato et al. data AND shows loss of function on another assay. The evidence shows loss of transactivation activity in yeast (Kato assay) and reduced growth suppression in human cancer cell lines (another functional assay). Therefore, PS3 is applied at Strong strength because the variant meets both non-functional and LOF requirements.
PS4 (Not Applied)
According to VCEP guidelines, PS4 applies based on proband points tally for Li-Fraumeni spectrum cancers. No such proband data or point total is reported for this variant. Therefore, PS4 is not applied.
PM1 (Not Applied)
According to VCEP guidelines, PM1 applies to missense variants at TP53 hotspot codons 175, 245, 248, 249, 273, or 282. p.Ser127 is outside these codons. Therefore, PM1 is not applied.
PM2 (Supporting)
According to VCEP guidelines for TP53, PM2 at Supporting strength applies when allele frequency is <0.00003 in gnomAD. The variant is absent from gnomAD. Therefore, PM2 is applied at Supporting strength.
PM3 (Not Applied)
According to standard ACMG guidelines, PM3 applies to recessive disorders with detected trans variants. TP53 is autosomal dominant and no phase data exist. Therefore, PM3 is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, PM4 applies to protein length changes due to in-frame indels or stop-loss variants. This is a missense variant. Therefore, PM4 is not applied.
PM5 (Not Applied)
According to VCEP guidelines, PM5 applies for missense at a residue with ≥1 TP53 pathogenic missense variants. No other pathogenic variants at codon 127 are documented by the VCEP. Therefore, PM5 is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines, PM6 applies to assumed de novo cases without confirmation. No de novo assumption or partial evidence exists. Therefore, PM6 is not applied.
PP1 (Not Applied)
According to VCEP guidelines, PP1 applies based on cosegregation in families. No familial segregation data are provided. Therefore, PP1 is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, PP2 applies when a gene has a low rate of benign missense variation and missense are common mechanism. TP53 has many pathogenic missense variants and benign variation; PP2 is not recommended by VCEP. Therefore, PP2 is not applied.
PP3 (Not Applied)
According to VCEP guidelines, PP3 at Moderate or Supporting requires BayesDel and aGVGD thresholds or SpliceAI ≥0.2. BayesDel score is unavailable and computational predictions are mixed. Therefore, PP3 is not applied.
PP4 (Not Applied)
According to VCEP guidelines, PP4 applies for somatic observation patterns (clonal hematopoiesis) with VAF metrics. No such VAF data are provided. Therefore, PP4 is not applied.
BA1 (Not Applied)
According to VCEP guidelines, BA1 applies for allele frequency ≥0.001 in gnomAD. The variant is absent. Therefore, BA1 is not applied.
BS1 (Not Applied)
According to VCEP guidelines, BS1 applies for filtering AF ≥0.0003 but <0.001. The variant is absent. Therefore, BS1 is not applied.
BS2 (Not Applied)
According to VCEP guidelines, BS2 applies for unrelated elderly female controls without cancer. No such data exist. Therefore, BS2 is not applied.
BS3 (Not Applied)
According to VCEP guidelines, BS3 applies when functional assays show no LOF on Kato et al. and other assays. The variant shows LOF. Therefore, BS3 is not applied.
BS4 (Not Applied)
According to VCEP guidelines, BS4 applies for lack of segregation in affected family members. No segregation data exist. Therefore, BS4 is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, BP1 applies to Missense in gene where only truncating variants cause disease. TP53 disease is missense dominant. Therefore, BP1 is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, BP2 applies when observed in trans with a pathogenic variant for autosomal dominant. No such evidence. Therefore, BP2 is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, BP3 applies to in-frame deletions in repetitive regions. This is a missense variant. Therefore, BP3 is not applied.
BP4 (Not Applied)
According to VCEP guidelines, BP4 at Moderate or Supporting requires BayesDel thresholds and SpliceAI <0.2. BayesDel is unavailable and in silico predictions are mixed. Therefore, BP4 is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, BP5 applies when variant found in a case with an alternate molecular cause. No alternate cause reported. Therefore, BP5 is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, BP6 applies when a reputable source reports benign without data. No such benign report. Therefore, BP6 is not applied.
BP7 (Not Applied)
According to VCEP guidelines, BP7 applies to synonymous variants with no splicing impact. This variant is missense. Therefore, BP7 is not applied.