Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000059.4 | MANE Select | 11954 nt | 200–10456 |
| NM_000059.2 | Alternative | 11386 nt | 228–10484 |
| NM_000059.3 | RefSeq Select | 11386 nt | 228–10484 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe c.4005dupA pathogenic mutation, located in coding exon 10 of the BRCA2 gene, results from a duplication of A at nucleotide position 4005, causing a translational frameshift with a predicted alternate stop codon (p.F1336Ifs*2). This alteration has been reported in multiple large studies of BRCA1/2 mutation positive families (Tea MK et al. Maturitas, 2014 Jan;77:68-72; Palmero EI et al. Sci Rep, 2018 06;8:9188; Rebbeck TR et al. Hum. Mutat., 2018 05;39:593-620). This alteration, designated 4232insA, has also been reported in a Hungarian male breast cancer patient (Ramus SJ et al. Am. J. Hum. Genet., 1997 May;60:1242-6). In addition to the clinical data presented in the literature, this alteration is expected to result in loss of function by premature protein truncation or nonsense-mediated mRNA decay. As such, this alteration is interpreted as a disease-causing mutation.
This sequence change creates a premature translational stop signal (p.Phe1336Ilefs*2) in the BRCA2 gene. It is expected to result in an absent or disrupted protein product. Loss-of-function variants in BRCA2 are known to be pathogenic (PMID: 20104584). This variant is not present in population databases (gnomAD no frequency). This premature translational stop signal has been observed in individual(s) with a personal and/or family history of breast and/or ovarian cancer (PMID: 9150174, 24156927, 29446198, 29907814). This variant is also known as 4232insA. ClinVar contains an entry for this variant (Variation ID: 51581). For these reasons, this variant has been classified as Pathogenic.
"This variant has been reported in ClinVar as Pathogenic (3 clinical laboratories) and as Pathogenic by Evidence-based Network for the Interpretation of Germline Mutant Alleles (ENIGMA) expert panel."
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
The BRCA2 F1336Ifs*2 variant is a truncating mutation that results in the loss of critical protein domains, including the C-terminal DNA binding domain, nuclear localization signal, and CDK2 phosphorylation site. Functional studies indicate that such truncating mutations impair the nuclear localization of BRCA2, disrupting its role in maintaining homologous recombination during DNA damage response. This functional impairment supports a damaging effect of the variant.
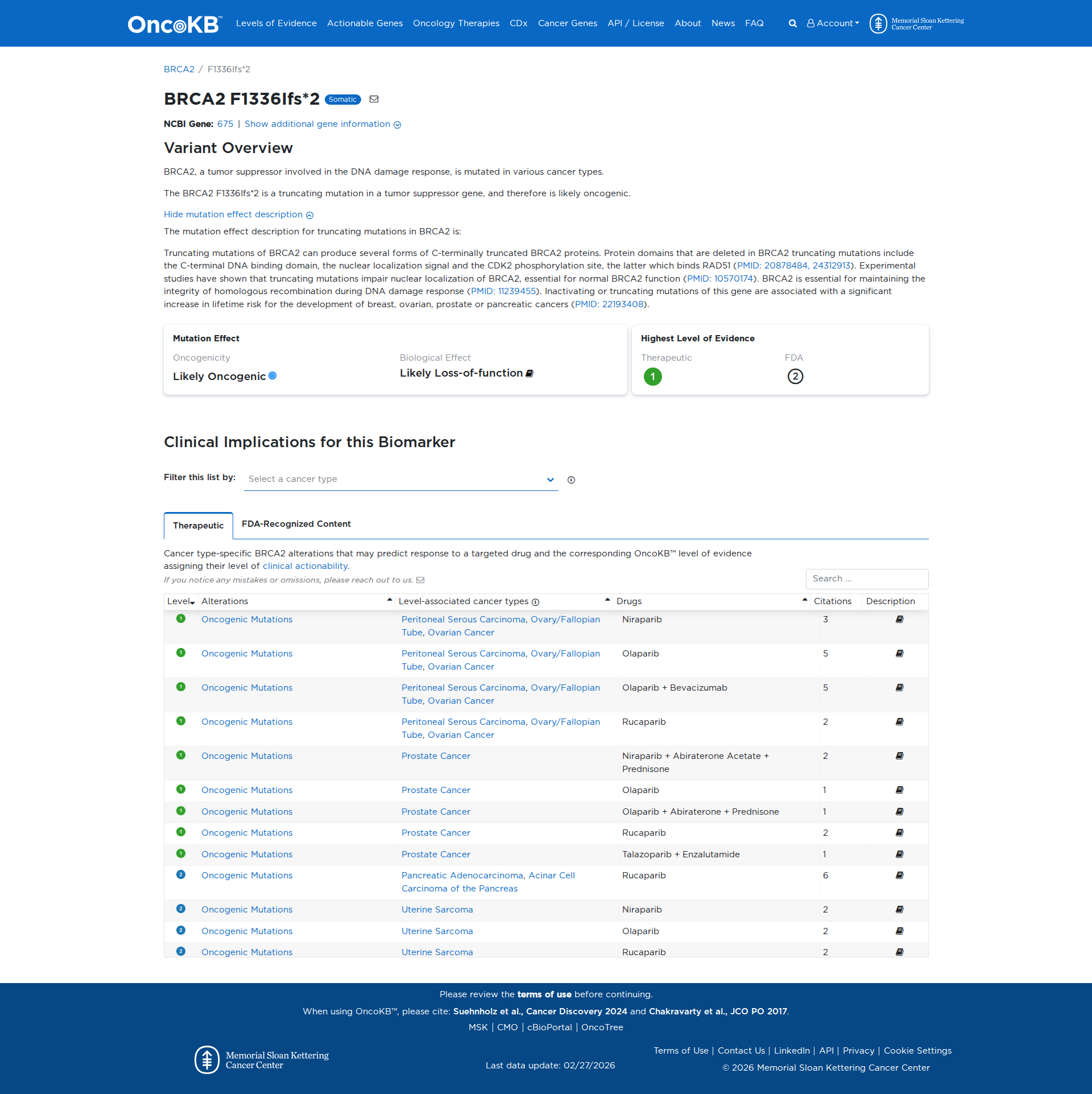
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 111 bp |
| Donor Loss (DL) | 0.0 | -453 bp |
| Acceptor Gain (AG) | 0.0 | 52 bp |
| Donor Gain (DG) | 0.01 | -157 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)