Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000051.3 | RefSeq Select | 13147 nt | 386–9556 |
| NM_000051.4 | MANE Select | 12915 nt | 151–9321 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenAvailable data are insufficient to determine the clinical significance of the variant at this time. The frequency of this variant in the general population is uninformative in assessment of its pathogenicity (http://gnomad.broadinstitute.org). Computational tools disagree on the variant's effect on normal protein function.
Variant summary: ATM c.610G>A (p.Gly204Arg) results in a non-conservative amino acid change in the encoded protein sequence. Four of five in-silico tools predict a damaging effect of the variant on protein function. The variant allele was found at a frequency of 8e-05 in 1613552 control chromosomes. This frequency is not significantly higher than estimated for a pathogenic variant in ATM causing Breast Cancer (8e-05 vs 0.001), allowing no conclusion about variant significance. c.610G>A has been reported in the literature in individuals affected with breast and/or ovarian cancer, prostate cancer, chronic lymphocytic leukemia and rhabdosarcoma in settings of multigene panel testing and in unaffected control cohorts (example, Tung 2015, Decker 2017, Singh 2018, Nadeu 2016, Tiao 2017, Zhang 2015, Bonache_2018, Girard_2019, Karlsson_2021). These reports do not provide unequivocal conclusions about association of the variant with Ataxia Telangiectasia or Breast Cancer. At-least one co-occurrence with another pathogenic variant has been reported in an individual undergoing multigene panel testing for breast cancer (BRCA1 c.5172dupA, p.Glu1725Argfs*7, Tung_2015), providing supporting evidence for a benign role. A recent study that included this variant reports that carriers of loss of function variants in the ATM gene have a significantly higher risk of developing breast cancer than carriers of an ATM missense variant (OR for LOF =17.4; OR for missense = 1.6) (Girard_2019). The following publications have been ascertained in the context of this evaluation (PMID: 20305132, 30306255, 28779002, 33471991, 34204722, 30303537, 29522266, 33436325, 27720647, 26837699, 29470806, 28652578, 25186627, 26580448, 37762649, 36685941). ClinVar contains an entry for this variant (Variation ID: 127419). Based on the evidence outlined above, the variant was classified as likely benign.
BP4
This alteration is classified as likely benign based on a combination of the following: seen in unaffected individuals, population frequency, intact protein function, lack of segregation with disease, co-occurrence, RNA analysis, in silico models, amino acid conservation, lack of disease association in case-control studies, and/or the mechanism of disease or impacted region is inconsistent with a known cause of pathogenicity.
"This variant has been reported in ClinVar as Uncertain significance (20 clinical laboratories) and as Likely benign (4 clinical laboratories)."
COSMIC Somatic Evidence
Open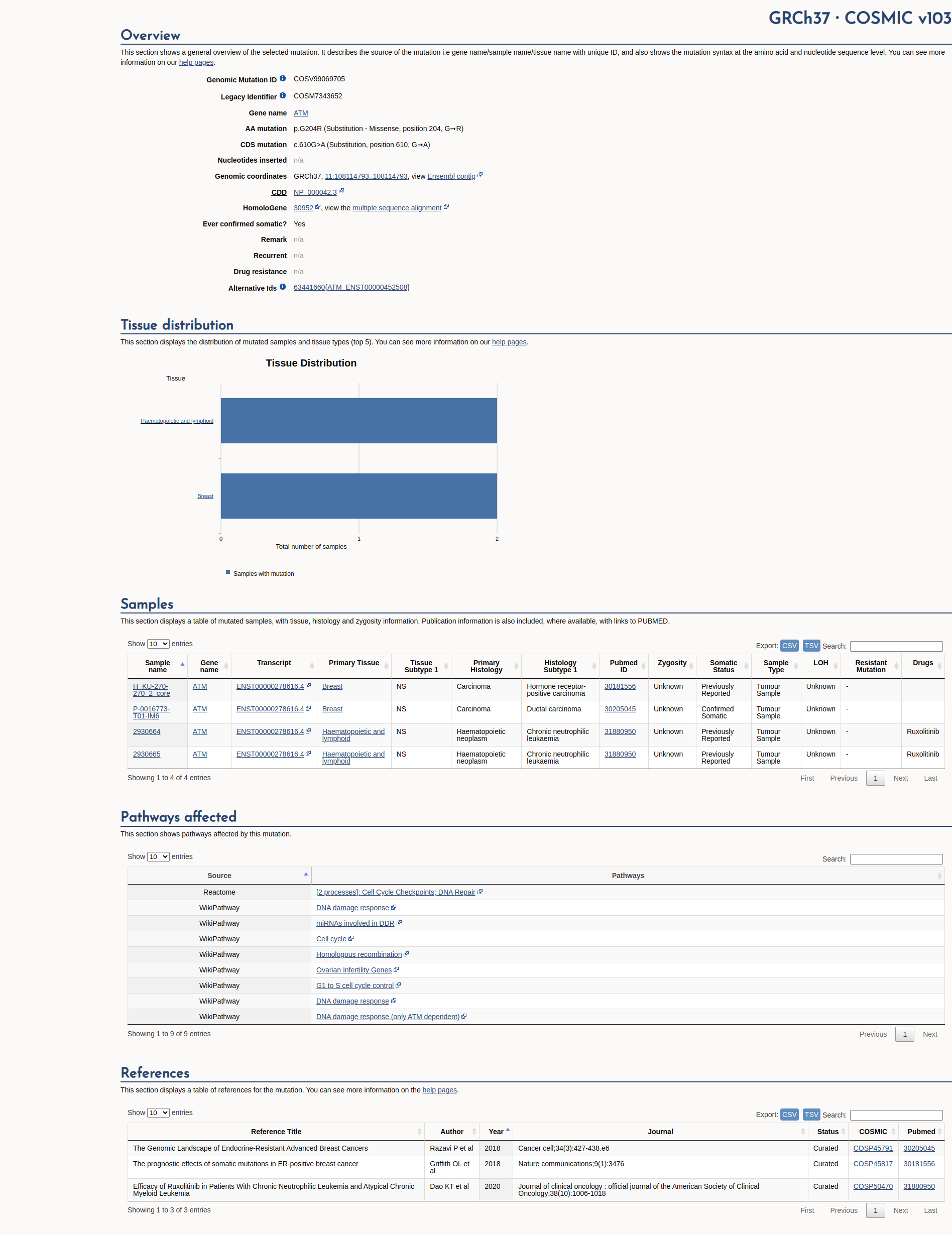
Functional Impact & Domains
Functional Domain
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.24 | -91 bp |
| Donor Loss (DL) | 0.13 | 52 bp |
| Acceptor Gain (AG) | 0.0 | 108 bp |
| Donor Gain (DG) | 0.0 | -416 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, the rule for PVS1 is: "Very Strong Strength: Very Strong Use ATM PVS1 Decision Tree". The evidence for this variant shows it is a missense change (G204R) not predicted to cause loss of function. Therefore, this criterion is not applied at Not Applied strength because PVS1 applies only to null variants in ATM per the VCEP decision tree.
PS1 (Not Applied)
According to VCEP guidelines, the rule for PS1 is: "Strong Strength: Strong Use for protein changes as long as splicing is ruled-out for both alterations". The evidence for this variant shows no other nucleotide change resulting in the same p.G204R amino acid has been reported as pathogenic. Therefore, this criterion is not applied at Not Applied strength because there is no established pathogenic variant with the same amino acid change.
PS2 (Not Applied)
According to standard ACMG guidelines, the rule for PS2 is: "De novo (both maternity and paternity confirmed) in a patient with the disease and no family history". The evidence for this variant shows no de novo or parental origin data. Therefore, this criterion is not applied at Not Applied strength because parental confirmation and de novo status cannot be evaluated.
PS3 (Not Applied)
According to VCEP guidelines, the rule for PS3 is: "Moderate Strength: Use when a variant fails to rescue both an ATM specific feature AND radiosensitivity". The evidence for this variant shows no functional studies characterizing ATM G204R. Therefore, this criterion is not applied at Not Applied strength because functional impact has not been assessed.
PS4 (Not Applied)
According to VCEP guidelines, the rule for PS4 is: "Strong Strength: Case-control studies; p-value ≤.05 AND (Odds ratio ≥2 OR lower 95% CI ≥1.5)". The evidence for this variant shows no case-control or cohort data. Therefore, this criterion is not applied at Not Applied strength because no statistical association data are available.
PM1 (Not Applied)
According to standard ACMG guidelines, the rule for PM1 is: "Located in a mutational hot spot and/or critical and well-established functional domain without benign variation". The evidence for this variant shows p.G204 is not in a recognized hotspot or critical functional domain for ATM. Therefore, this criterion is not applied at Not Applied strength.
PM2 (Not Applied)
According to VCEP guidelines, the rule for PM2 is: "Supporting Strength: Frequency ≤.001% if n=1 in a single subpopulation". The evidence for this variant shows a gnomAD MAF of 0.0103% (29/282440 alleles). Therefore, this criterion is not applied at Not Applied strength because the allele frequency exceeds the VCEP threshold for PM2_supporting.
PM3 (Not Applied)
According to VCEP guidelines, the rule for PM3 is: "Very Strong/Strong/Moderate/Supporting Strength: Use ATM PM3/BP2 table". The evidence for this variant shows no data on trans configuration with a pathogenic allele in recessive disease. Therefore, this criterion is not applied at Not Applied strength.
PM4 (Not Applied)
According to VCEP guidelines, the rule for PM4 is: "Moderate Strength: Use for stop-loss variants". The evidence for this variant shows it is a missense change, not a stop-loss. Therefore, this criterion is not applied at Not Applied strength.
PM5 (Not Applied)
According to VCEP guidelines, the rule for PM5 is: "Supporting Strength: Use for genomic frameshift and truncating variants with PTC upstream of p.R3047". The evidence for this variant shows it is a missense change and does not create a novel stop codon. Therefore, this criterion is not applied at Not Applied strength.
PM6 (Not Applied)
According to standard ACMG guidelines, the rule for PM6 is: "Unconfirmed de novo (without confirmation of paternity and maternity)". The evidence for this variant shows no de novo information at all. Therefore, this criterion is not applied at Not Applied strength because de novo status cannot be evaluated.
PP1 (Not Applied)
According to standard ACMG guidelines, the rule for PP1 is: "Co-segregation with disease in multiple affected family members in a gene definitively known to cause the disease". The evidence for this variant shows no segregation data. Therefore, this criterion is not applied at Not Applied strength.
PP2 (Not Applied)
According to standard ACMG guidelines, the rule for PP2 is: "Missense variant in a gene with low rate of benign missense variation and where missense is a common mechanism of disease". The evidence for ATM shows both loss-of-function and missense may underlie disease, and benign missense are also observed. Therefore, this criterion is not applied at Not Applied strength.
PP3 (Not Applied)
According to VCEP guidelines, the rule for PP3 is: "Supporting Strength: REVEL >.7333 or RNA predictor impact". The evidence for this variant shows mixed in silico results and no REVEL score >0.7333, with SpliceAI score 0.24 indicating low splicing impact. Therefore, this criterion is not applied at Not Applied strength.
PP4 (Not Applied)
According to standard ACMG guidelines, the rule for PP4 is: "Patient’s phenotype or family history is highly specific for a disease with a single genetic etiology". The evidence for this variant shows no clinical phenotype data provided. Therefore, this criterion is not applied at Not Applied strength.
PP5 (Not Applied)
According to standard ACMG guidelines, the rule for PP5 is: "Reputable source recently reports variant as pathogenic but the evidence is not available to the laboratory to perform an independent evaluation". The evidence for this variant shows ClinVar reports VUS and likely benign, not pathogenic. Therefore, this criterion is not applied at Not Applied strength.
BA1 (Not Applied)
According to VCEP guidelines, the rule for BA1 is: "Stand Alone Strength: Filtering Allele Frequency >.5%". The evidence for this variant shows a frequency of 0.0103%, which is below 0.5%. Therefore, this criterion is not applied at Not Applied strength.
BS1 (Not Applied)
According to VCEP guidelines, the rule for BS1 is: "Strong Strength: Filtering Allele Frequency >.05%". The evidence for this variant shows a frequency of 0.0103%, which is below 0.05%. Therefore, this criterion is not applied at Not Applied strength.
BS2 (Not Applied)
According to standard ACMG guidelines, the rule for BS2 is: "Observed in a healthy adult individual for a dominant disorder with full penetrance expected at an early age". The evidence for this variant shows no data on observation in healthy adults. Therefore, this criterion is not applied at Not Applied strength.
BS3 (Not Applied)
According to VCEP guidelines, the rule for BS3 is: "Moderate Strength: Use when a variant rescues both an ATM specific feature AND radiosensitivity". The evidence for this variant shows no functional rescue data. Therefore, this criterion is not applied at Not Applied strength.
BS4 (Not Applied)
According to standard ACMG guidelines, the rule for BS4 is: "Lack of segregation in affected members of a family". The evidence for this variant shows no segregation analysis. Therefore, this criterion is not applied at Not Applied strength.
BP1 (Not Applied)
According to standard ACMG guidelines, the rule for BP1 is: "Missense variant in a gene for which primarily truncating variants are known to cause disease". The evidence for ATM shows both missense and truncating variants can be pathogenic. Therefore, this criterion is not applied at Not Applied strength.
BP2 (Not Applied)
According to VCEP guidelines, the rule for BP2 is: "Strong/Moderate/Supporting Strength: Use ATM PM3/BP2 table". The evidence for this variant shows no trans configuration data with another pathogenic variant. Therefore, this criterion is not applied at Not Applied strength.
BP3 (Not Applied)
According to standard ACMG guidelines, the rule for BP3 is: "In-frame deletions/insertions in a repetitive region without a known function". The evidence for this variant shows it is a missense change. Therefore, this criterion is not applied at Not Applied strength.
BP4 (Not Applied)
According to VCEP guidelines, the rule for BP4 is: "Supporting Strength: REVEL ≤.249 or at least one RNA predictor shows impact". The evidence for this variant shows mixed protein predictions and no strong splicing prediction. Therefore, this criterion is not applied at Not Applied strength.
BP5 (Not Applied)
According to standard ACMG guidelines, the rule for BP5 is: "Variant found in a case with an alternate molecular basis for disease". The evidence for this variant shows no alternate molecular diagnosis. Therefore, this criterion is not applied at Not Applied strength.
BP6 (Not Applied)
According to standard ACMG guidelines, the rule for BP6 is: "Reputable source reports variant as benign but evidence unavailable". The evidence for this variant shows ClinVar submissions as VUS and likely benign, not unequivocally benign. Therefore, this criterion is not applied at Not Applied strength.
BP7 (Not Applied)
According to standard ACMG guidelines, the rule for BP7 is: "Synonymous variant with no predicted splicing impact". The evidence for this variant shows it is a missense change. Therefore, this criterion is not applied at Not Applied strength.