Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_024675.4 | MANE Select | 4008 nt | 154–3714 |
| NM_024675.3 | RefSeq Select | 4069 nt | 201–3761 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenVariant summary: The PALB2 c.721A>G (p.Asn241Asp) variant involves the alteration of a non-conserved nucleotide and 5/5 in silico tools predict a benign outcome for this variant, although these predictions have yet to be functionally assessed. This variant was found in 75/121762 control chromosomes, predominantly observed in the African subpopulation at a frequency of 0.006935 (71/10238). This frequency is about 44 times the estimated maximal expected allele frequency of a pathogenic PALB2 variant (0.0001563), suggesting this is likely a benign polymorphism found primarily in the populations of African origin. Multiple publications have cited the variant in affected individuals, predominantly of African decent, although with limited information (ie, lack of co-occurrence and cosegregation data). In addition, multiple clinical diagnostic laboratories classified this variant as likely benign/benign. Taken together, this variant is classified as benign.
This alteration is classified as benign based on a combination of the following: seen in unaffected individuals, population frequency, intact protein function, lack of segregation with disease, co-occurrence, RNA analysis, in silico models, amino acid conservation, lack of disease association in case-control studies, and/or the mechanism of disease or impacted region is inconsistent with a known cause of pathogenicity.
Curators: Marc Tischkowitz, Arleen D. Auerbach. Submitters to LOVD: Marc Tischkowitz, Melissa DeRycke, Yukihide Momozawa.
This variant is considered benign. Homozygosity for this variant has been confirmed in one or more individuals lacking clinical features consistent with gene-specific recessive disease, indicating that this variant is unlikely to be pathogenic. This variant is strongly associated with less severe personal and family histories of cancer, typical for individuals without pathogenic variants in this gene [PMID: 25085752]. This variant has been observed at a population frequency that is significantly greater than expected given the associated disease prevalence and penetrance.
"This variant has been reported in ClinVar as Benign (9 clinical laboratories) and as Likely benign (8 clinical laboratories) and as Benign by ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open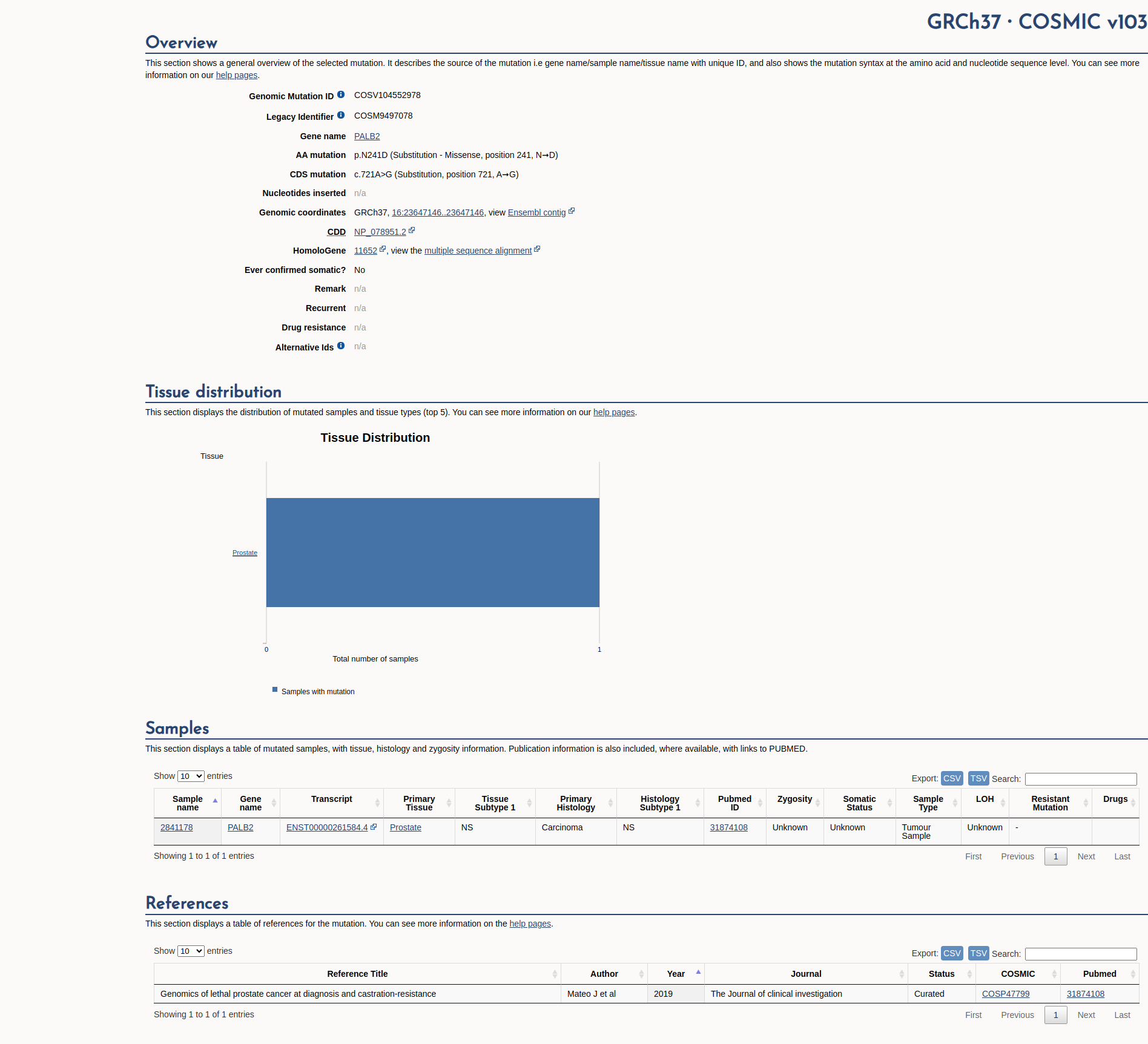
Functional Impact & Domains
Functional Domain
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | None bp |
| Donor Loss (DL) | 0.0 | None bp |
| Acceptor Gain (AG) | 0.0 | None bp |
| Donor Gain (DG) | 0.0 | None bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, the rule for PVS1 is: "Very Strong Use PALB2 PVS1 Decision Tree Modification Type: Gene-specific,Strength". The evidence for this variant shows: it is a missense change (N241D), not a null variant. Therefore, this criterion is not applied.
PS1 (Not Applied)
According to VCEP guidelines, the rule for PS1 is: "Strong Use PALB2 PS1 Splicing table Modification Type: General recommendation". The evidence for this variant shows: no previously established pathogenic variant results in the same amino acid change. Therefore, this criterion is not applied.
PS2 (Not Applied)
According to standard ACMG guidelines, the rule for PS2 is: "De novo (both maternity and paternity confirmed) in a patient with the disease and no family history". The evidence for this variant shows: no confirmed de novo occurrence. Therefore, this criterion is not applied.
PS3 (Not Applied)
According to standard ACMG guidelines, the rule for PS3 is: "Well‐established in vitro or in vivo functional studies supportive of a damaging effect". The evidence for this variant shows: no functional studies have been performed. Therefore, this criterion is not applied.
PS4 (Not Applied)
According to VCEP guidelines, the rule for PS4 is: "Strong Case-control studies; p-value ≤ .05 AND (Odds ratio … ≥3 or lower 95% CI ≥1.5)". The evidence for this variant shows: no case-control data. Therefore, this criterion is not applied.
PM1 (Not Applied)
According to standard ACMG guidelines, the rule for PM1 is: "Located in a mutational hot spot and/or critical functional domain without benign variation". The evidence for this variant shows: no data placing N241D in a known hotspot or critical domain. Therefore, this criterion is not applied.
PM2 (Not Applied)
According to VCEP guidelines, the rule for PM2 is: "Supporting Variant absent in gnomAD or present in ≤ 1/300,000 alleles Modification Type: Gene-specific,Strength". The evidence for this variant shows: MAF = 0.0616% (~1/1,622 alleles), above the ≤1/300,000 threshold. Therefore, this criterion is not applied.
PM3 (Not Applied)
According to VCEP guidelines, the rule for PM3 is: "Strong Use Fanconi Anemia PM3 tables Modification Type: Disease-specific,Strength". The evidence for this variant shows: no evidence of trans observations in recessive disease. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, the rule for PM4 is: "Protein length changes due to in-frame indels or stop-loss variants". The evidence for this variant shows: it is a missense substitution with no change in protein length. Therefore, this criterion is not applied.
PM5 (Not Applied)
According to VCEP guidelines, the rule for PM5 is: "Supporting Apply to frameshifting or truncating variants with premature termination codons upstream of p.Tyr1183". The evidence for this variant shows: it is missense, not a truncating variant. Therefore, this criterion is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines, the rule for PM6 is: "Assumed de novo, but without confirmation of paternity and maternity". The evidence for this variant shows: no de novo data. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to VCEP guidelines, the rule for PP1 is: "Supporting LOD ≥0.3 or Bayes Factor ≥2:1 Modification Type: Gene-specific". The evidence for this variant shows: no segregation data. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, the rule for PP2 is: "Missense variant in a gene with low rate of benign missense variation and where pathogenic missense variants are common". The evidence for this variant shows: PALB2 tolerates missense and has both benign and pathogenic missense reported; no clear enrichment. Therefore, this criterion is not applied.
PP3 (Not Applied)
According to VCEP guidelines, the rule for PP3 is: "Supporting Protein: Do not use. RNA: At least one well-established in silico predictor shows impact on splicing". The evidence for this variant shows: mixed in silico predictions and no clear splice impact. Therefore, this criterion is not applied.
PP4 (Not Applied)
According to standard ACMG guidelines, the rule for PP4 is: "Patient’s phenotype or family history highly specific for gene". The evidence for this variant shows: no phenotype or family history data. Therefore, this criterion is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, the rule for PP5 is: "Reputable source recently reports variant as pathogenic". The evidence for this variant shows: ClinVar reports it as benign/likely benign. Therefore, this criterion is not applied.
BA1 (Not Applied)
According to VCEP guidelines, the rule for BA1 is: "Stand Alone GnomAD Filtering Allele Frequency >0.1% Modification Type: Disease-specific,Gene-specific". The evidence for this variant shows: MAF = 0.0616%, below the 0.1% threshold. Therefore, this criterion is not applied.
BS1 (Strong)
According to VCEP guidelines, the rule for BS1 is: "Strong GnomAD Filtering Allele Frequency greater than expected for disease >0.01% Modification Type: Disease-specific,Gene-specific". The evidence for this variant shows: MAF = 0.0616%, which is above 0.01%. Therefore, this criterion is applied at Strong strength because the allele frequency exceeds the gene-specific threshold for a benign classification.
BS2 (Not Applied)
According to VCEP guidelines, the rule for BS2 is: "Strong Per Fanconi Anemia BS2 tables Modification Type: Disease-specific". The evidence for this variant shows: no observation in healthy adults. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to standard ACMG guidelines, the rule for BS3 is: "Well‐established functional studies show no damaging effect". The evidence for this variant shows: no functional studies. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to VCEP guidelines, the rule for BS4 is: "Strong LOD ≤ -1.28 or Bayes Factor ≤0.053:1 Modification Type: Gene-specific". The evidence for this variant shows: no segregation data. Therefore, this criterion is not applied.
BP1 (Supporting)
According to VCEP guidelines, the rule for BP1 is: "Supporting Apply to all missense variants. Modification Type: Gene-specific". The evidence for this variant shows: N241D is a missense change. Therefore, this criterion is applied at Supporting strength because it is a missense variant in PALB2.
BP2 (Not Applied)
According to standard ACMG guidelines, the rule for BP2 is: "Observed in trans with a pathogenic variant for a fully penetrant dominant gene". The evidence for this variant shows: no trans observations. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, the rule for BP3 is: "In-frame deletions/insertions in a repetitive region without a known function". The evidence for this variant shows: not an in-frame indel. Therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP guidelines, the rule for BP4 is: "Supporting RNA: At least one well-established in silico predictor shows no impact on splicing; Protein: do not use". The evidence for this variant shows: mixed computational predictions and no consistent splice result. Therefore, this criterion is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, the rule for BP5 is: "Variant found in a case with an alternate molecular basis for disease". The evidence for this variant shows: no such case data. Therefore, this criterion is not applied.
BP6 (Supporting)
According to standard ACMG guidelines, the rule for BP6 is: "Reputable source recently reports variant as benign". The evidence for this variant shows: ClinVar submissions classify it as benign or likely benign by multiple reputable laboratories. Therefore, this criterion is applied at Supporting strength because of the consensus benign reports.
BP7 (Not Applied)
According to VCEP guidelines, the rule for BP7 is: "Strong BP7_Strong(RNA): Observed lack of aberrant RNA defect for silent substitutions and intronic variants". The evidence for this variant shows: it is not silent or intronic. Therefore, this criterion is not applied.