Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_006231.4 | MANE Select | 7823 nt | 28–6888 |
| NM_006231.2 | Alternative | 7859 nt | 45–6905 |
| NM_006231.3 | RefSeq Select | 8024 nt | 210–7070 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Open""
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: POLES297PPOLES297PSomaticNCBI Gene:5426|Show additional gene information Variant OverviewPOLE, the catalytic subunit of DNA polymerase epsilon, is an enzyme involved in DNA replication and repair. Select POLE mutations lead to ultra-high mutation rates, most frequently in endometrial and colorectal cancer.The POLE S297P mutation has not specifically been reviewed by the OncoKB team. However, POLE S297F/Y are likely oncogenic, and therefore POLE S297P is considered likely oncogenic.Hide mutation effect description The POLE S297P mutation has not specifically been reviewed by the OncoKB team. However, we have mutation effect descriptions for POLE S297F/Y JAX-CKB: No results found
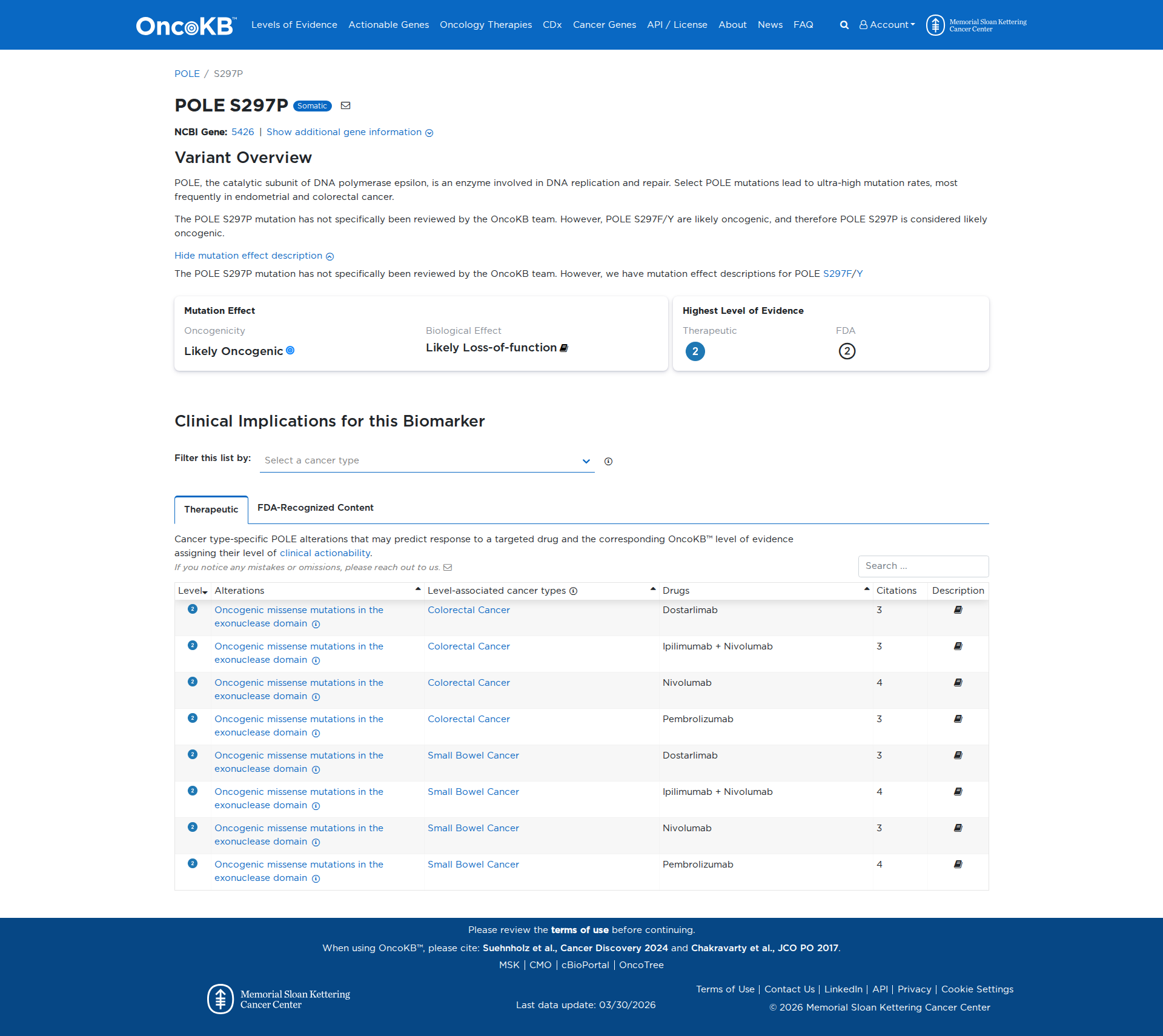
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 393 bp |
| Donor Loss (DL) | 0.0 | 284 bp |
| Acceptor Gain (AG) | 0.0 | -399 bp |
| Donor Gain (DG) | 0.0 | -28 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)