Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_007294.4 | MANE Select | 7088 nt | 114–5705 |
| NM_007294.2 | Alternative | 7191 nt | 201–5792 |
| NM_007294.3 | RefSeq Select | 7224 nt | 233–5824 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis alteration is classified as likely benign based on a combination of the following: seen in unaffected individuals, population frequency, intact protein function, lack of segregation with disease, co-occurrence, RNA analysis, in silico models, amino acid conservation, lack of disease association in case-control studies, and/or the mechanism of disease or impacted region is inconsistent with a known cause of pathogenicity.
This submission and the accompanying classification are no longer maintained by the submitter. For more information on current observations and classification, please contact variantquestions@myriad.com.
Variant summary: BRCA1 c.1534C>T (p.Leu512Phe) results in a non-conservative amino acid change in the encoded protein sequence. Four of five in-silico tools predict a damaging effect of the variant on protein function. The variant allele was found at a frequency of 2.4e-05 in 251232 control chromosomes. The available data on variant occurrences in the general population are insufficient to allow any conclusion about variant significance. c.1534C>T has been reported in the literature in individuals affected with Hereditary Breast And Ovarian Cancer Syndrome (examples- Meindl_2002, Judkins_2005, Jimenez_2009, Plascocinska_2016). These data do not allow any conclusion about variant significance. Co-occurrences with other pathogenic variants have been reported [BRCA1 c.81_4986del , p.Cys27X; BRCA2 c.8023A>G , p.Ile2675Val; and BRCA2 c.5909C>A, p.Ser1970X (BIC database); BRCA1 c.1570delG, p.Ala524Glnfs (Jimenez_2009); BRCA2 c.1813dupA, p.Ile605AsnfsX11 (internal sample)], providing supporting evidence for a benign role. To our knowledge, no experimental evidence demonstrating an impact on protein function has been reported. Ten ClinVar submitters (evaluation after 2014) reported the variant with conflicting assessments, citing the variant as uncertain significance (n=2), likely benign (n=7), and benign (n=1, expert panel). Based on the evidence outlined above, the variant was classified as benign.
IARC class based on posterior probability from multifactorial likelihood analysis, thresholds for class as per Plon et al. 2008 (PMID: 18951446). Class 1 based on posterior probability = 0.000143
"This variant has been reported in ClinVar as Likely benign (8 clinical laboratories) and as Likely Benign (1 clinical laboratories) and as Uncertain significance (4 clinical laboratories) and as Benign (3 clinical laboratories) and as likely benign (1 clinical laboratories) and as Benign by Evidence-based Network for the Interpretation of Germline Mutant Alleles (ENIGMA) expert panel."
COSMIC Somatic Evidence
Open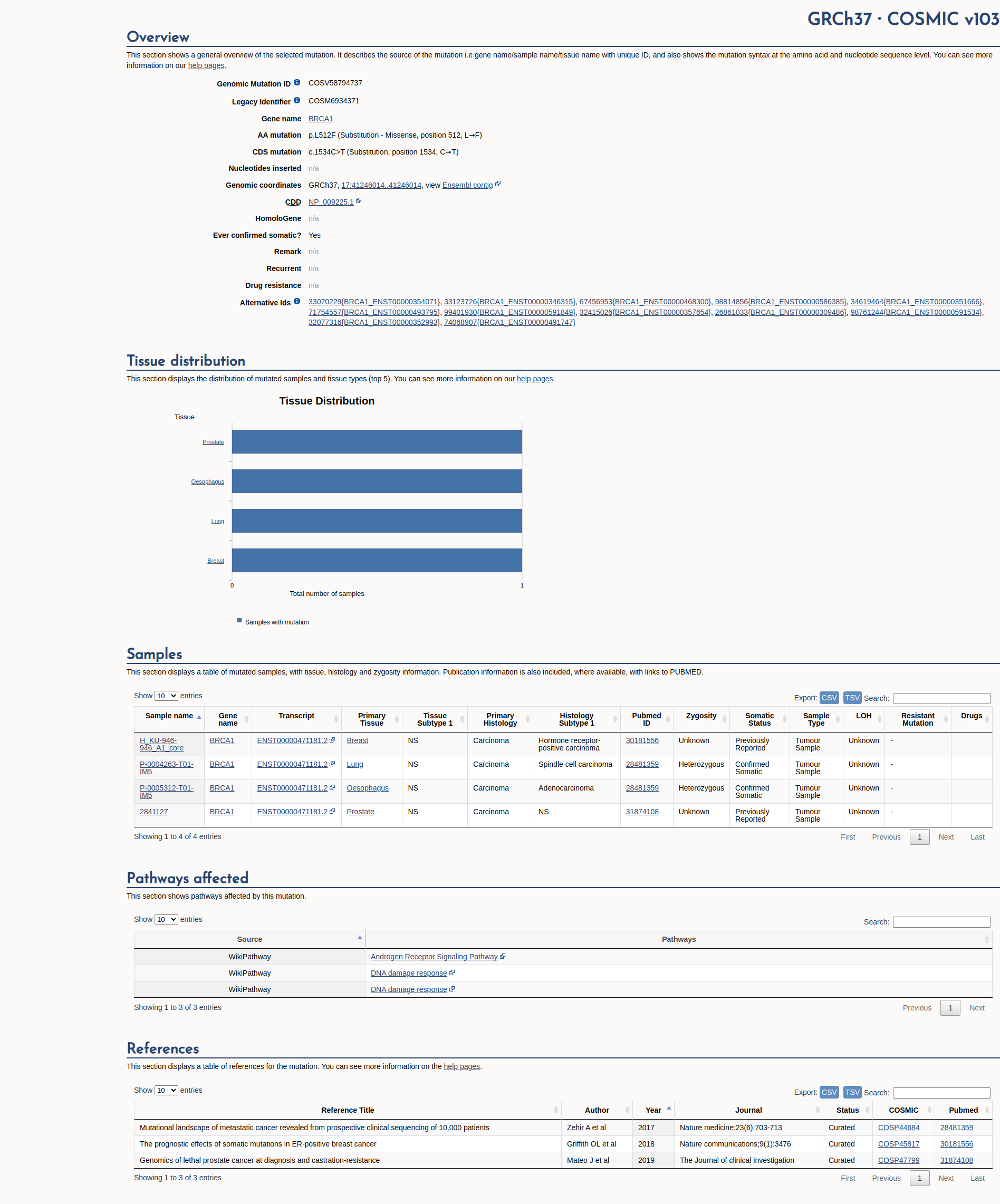
Functional Impact & Domains
Functional Domain
The BRCA1 L512F variant has not been functionally characterized, and its biological significance remains unknown.

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 83 bp |
| Donor Loss (DL) | 0.0 | -477 bp |
| Acceptor Gain (AG) | 0.0 | -395 bp |
| Donor Gain (DG) | 0.0 | -344 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines: 'Very Strong Null variant (nonsense, frameshift, splice site...) in a gene where loss of function is a known mechanism of disease.' The evidence for this variant shows it is a missense change (L512F), not a null variant. Therefore, this criterion is not applied.
PS1 (Not Applied)
According to VCEP guidelines: PS1 applies where a previously classified pathogenic variant yields the same amino acid change. The evidence for this variant shows no known pathogenic variant at residue L512 producing the same change. Therefore, this criterion is not applied.
PS2 (Not Applied)
Standard ACMG: PS2 applies for confirmed de novo occurrence. There is no information on de novo status. Therefore, this criterion is not applied.
PS3 (Not Applied)
Standard ACMG: PS3 requires well‐established functional studies demonstrating a damaging effect. No such functional data exist for this variant. Therefore, this criterion is not applied.
PS4 (Not Applied)
Standard ACMG: PS4 requires significant case‐control evidence of increased prevalence in affected individuals. No case‐control or cohort data are available. Therefore, this criterion is not applied.
PM1 (Not Applied)
According to VCEP guidelines: PM1 applies to variants in clinically important functional domains (RING 2‐101; coiled‐coil 1391‐1424; BRCT 1650‐1857). L512F lies outside these domains. Therefore, this criterion is not applied.
PM2 (Supporting)
According to VCEP guidelines: 'PM2 Supporting: Absent from controls in an outbred population.' The evidence for this variant shows a gnomAD MAF of 0.00354%, which meets the rarity threshold. Therefore, PM2 is applied at Supporting strength.
PM3 (Not Applied)
According to VCEP guidelines: PM3 applies for variants in trans with a pathogenic variant in BRCA1/2 in Fanconi anemia cases. There is no such phenotype or co‐occurrence data. Therefore, this criterion is not applied.
PM4 (Not Applied)
Standard ACMG: PM4 applies for protein length changes such as in‐frame indels. This is a missense substitution; no length change. Therefore, this criterion is not applied.
PM5 (Not Applied)
Standard ACMG: PM5 applies when a different pathogenic missense change occurs at the same residue. No known pathogenic variant at codon 512. Therefore, this criterion is not applied.
PM6 (Not Applied)
Standard ACMG: PM6 applies for assumed de novo without confirmation. No de novo data. Therefore, this criterion is not applied.
PP1 (Not Applied)
Standard ACMG: PP1 requires segregation data in families. No segregation information is available. Therefore, this criterion is not applied.
PP2 (Not Applied)
Standard ACMG: PP2 applies to missense variants in genes with low rate of benign missense. BRCA1 has known pathogenic missense in specific domains; this position is outside. No guideline‐endorsed application. Therefore, this criterion is not applied.
PP3 (Not Applied)
According to VCEP guidelines: PP3 applies to missense variants inside functional domains with damaging protein predictions. L512F is outside domains and in silico evidence is mixed. Therefore, this criterion is not applied.
PP4 (Not Applied)
Standard ACMG: PP4 applies for highly specific phenotype. No individual clinical phenotype provided. Therefore, this criterion is not applied.
PP5 (Not Applied)
Standard ACMG: PP5 applies when multiple reputable sources classify as pathogenic. ClinVar entries are mixed benign/VUS; no consensus pathogenic. Therefore, this criterion is not applied.
BA1 (Not Applied)
According to VCEP guidelines: BA1 applies if allele frequency >0.1%. The variant MAF of 0.00354% is below threshold. Therefore, this criterion is not applied.
BS1 (Not Applied)
According to VCEP guidelines: BS1 applies if filter AF >0.01%. The variant AF is 0.00354%, below threshold. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to VCEP guidelines: BS2 applies when variant is observed in healthy adults without FA features. No such observations reported. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to VCEP guidelines: BS3 requires well‐established functional studies showing no damaging effect. No such studies exist. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to VCEP guidelines: BS4 applies for lack of segregation. No family segregation data. Therefore, this criterion is not applied.
BP1 (Strong)
According to VCEP guidelines: 'BP1_Strong: missense variants outside clinically important domains AND no splicing predicted (SpliceAI ≤0.1).' L512F lies outside defined domains and SpliceAI predicts no splicing impact. Therefore, BP1 is applied at Strong strength.
BP2 (Not Applied)
Standard ACMG: BP2 applies for observation in trans with a pathogenic variant without phenotype. No such co‐observation data. Therefore, this criterion is not applied.
BP3 (Not Applied)
Standard ACMG: BP3 applies for in‐frame indels in repetitive regions. This is a missense substitution. Therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP guidelines: BP4 applies only to variants inside functional domains with no predicted impact. L512F is outside domains. Therefore, this criterion is not applied.
BP5 (Not Applied)
According to VCEP guidelines: BP5 applies to co‐occurrence with pathogenic variants in genes giving no specific phenotype. No such co‐occurrence reported. Therefore, this criterion is not applied.
BP6 (Not Applied)
Standard ACMG: BP6 applies for reputable source classification as benign without evidence. ClinVar submissions are mixed; no unanimous benign. Therefore, this criterion is not applied.
BP7 (Not Applied)
According to VCEP guidelines: BP7 applies to silent/intronic variants with no predicted splicing impact. This is a missense variant. Therefore, this criterion is not applied.