Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_002834.5 | MANE Select | 6073 nt | 166–1947 |
| NM_002834.3 | Alternative | 6300 nt | 381–2162 |
| NM_002834.4 | RefSeq Select | 6139 nt | 215–1996 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis sequence change replaces aspartic acid, which is acidic and polar, with glycine, which is neutral and non-polar, at codon 61 of the PTPN11 protein (p.Asp61Gly). This variant is not present in population databases (gnomAD no frequency). This missense change has been observed in individual(s) with Noonan syndrome and is one of the most commonly reported variants in this condition. It has also been observed in an individual with juvenile myelomonocytic leukemia. (PMID: 11704759, 11992261, 12634870, 15928039, 16358218, 17020470, 22420426, 23321623, 26084119, 26242988). In at least one individual the variant was observed to be de novo. ClinVar contains an entry for this variant (Variation ID: 13330). Invitae Evidence Modeling incorporating data from in vitro experimental studies (internal data) indicates that this missense variant is expected to disrupt PTPN11 function with a positive predictive value of 95%. Experimental studies have shown that this missense change affects PTPN11 function (PMID: 15273746, 15987685, 16377799, 19008228). For these reasons, this variant has been classified as Pathogenic.
The p.Asp61Gly variant in PTPN11 has been previously reported in >30 individuals with Noonan syndrome with or without juvenile myelomonocytic leukemia (JMML) in cluding at least 5 de novo occurrences (Tartagila 2001, Kosaki 2002, Yoshida 200 4, Kratz 2005, Bertola 2006, Chan 2006, Shaw 2007, Noordam 2005, Strullu 2014, B ouchikhi 2015, LMM data). It was also identified as a somatic variant in 1 child with acute lymphoblastic leukemia (ALL) and 2 children with JMML (Yamamoto 2006 , Stullu 2014). It has not been identified in large population studies. Both in vivo animal models and in vitro studies provide evidence that this variant impac ts protein function (Araki 2004, Kontaridis 2006, Uhlen 2006, Eminaga 2008, Wang 2009, Xu 2010, De Rocca 2012, Bonetti 2014, Lee 2014). In summary, this variant meets our criteria to be classified as pathogenic for Noonan syndrome and JMML in an autosomal dominant manner.
Variant summary: PTPN11 c.182A>G (p.Asp61Gly) results in a non-conservative amino acid change located in the SH2 domain of the encoded protein sequence. Four of five in-silico tools predict a damaging effect of the variant on protein function. The variant was absent in 251804 control chromosomes. c.182A>G has been well reported in the literature in multiple individuals affected with Noonan Syndrome and Related Conditions (example, Tartaglia_2001, Musante_2003, Bertola_2006, Kosaki_2002, Ferreira_2008, Strullu_2014). These data indicate that the variant is very likely to be associated with disease. Several publications report experimental evidence evaluating an impact on protein function. The most pronounced variant effect results in a gain of function leading to activation of the Ras-ErK signaling pathway (example, Hu_2015). Five clinical diagnostic laboratories have submitted clinical-significance assessments for this variant to ClinVar after 2014 without evidence for independent evaluation. All laboratories classified the variant as pathogenic. Based on the evidence outlined above, the variant was classified as pathogenic.
The PTPN11 c.182A>G (p.Asp61Gly) missense variant results in the substitution of asparagine at amino acid position 61 with glycine. This variant is one of the most commonvPTPN11 variants reported in association with Noonan syndrome. Across a selection of the available literature, the c.182A>G variant has been reported in at least 28 individuals with Noonan syndrome, at least six of whom also showed features of juvenile myelomonocytic leukemia or myeloproliferative disorder (PMID: 11992261; PMID: 15928039; PMID: 25097206; PMID: 26084119). The c.182A>G variant has also been shown to occur de novo in at least two additional affected individuals (PMID: 23321623; PMID: 26242988). This variant is not found in version 2.1.1 or version 3.1.2 of the Genome Aggregation Database. Structural modeling has shown that asparagine 61 is located at the N-SH2/PTP interaction surface, which is a mutational hotspot (PMID: 11992261), and functional assays have demonstrated that the variant enhances basal protein activity (gain of function) (PMID: 15987685). A heterozygous knock-in mouse model with the p.Asp61Gly amino acid change exhibits decreased viability and recapitulates clinical features of Noonan syndrome, including short stature, craniofacial anomalies, and myeloproliferative disease (PMID: 15273746). This variant was also identified in a de novo state. Based on the available evidence, the c.182A>G (p.Asp61Gly) variant is classified as pathogenic for Noonan syndrome.
"This variant has been reported in ClinVar as Pathogenic (25 clinical laboratories)."
COSMIC Somatic Evidence
Open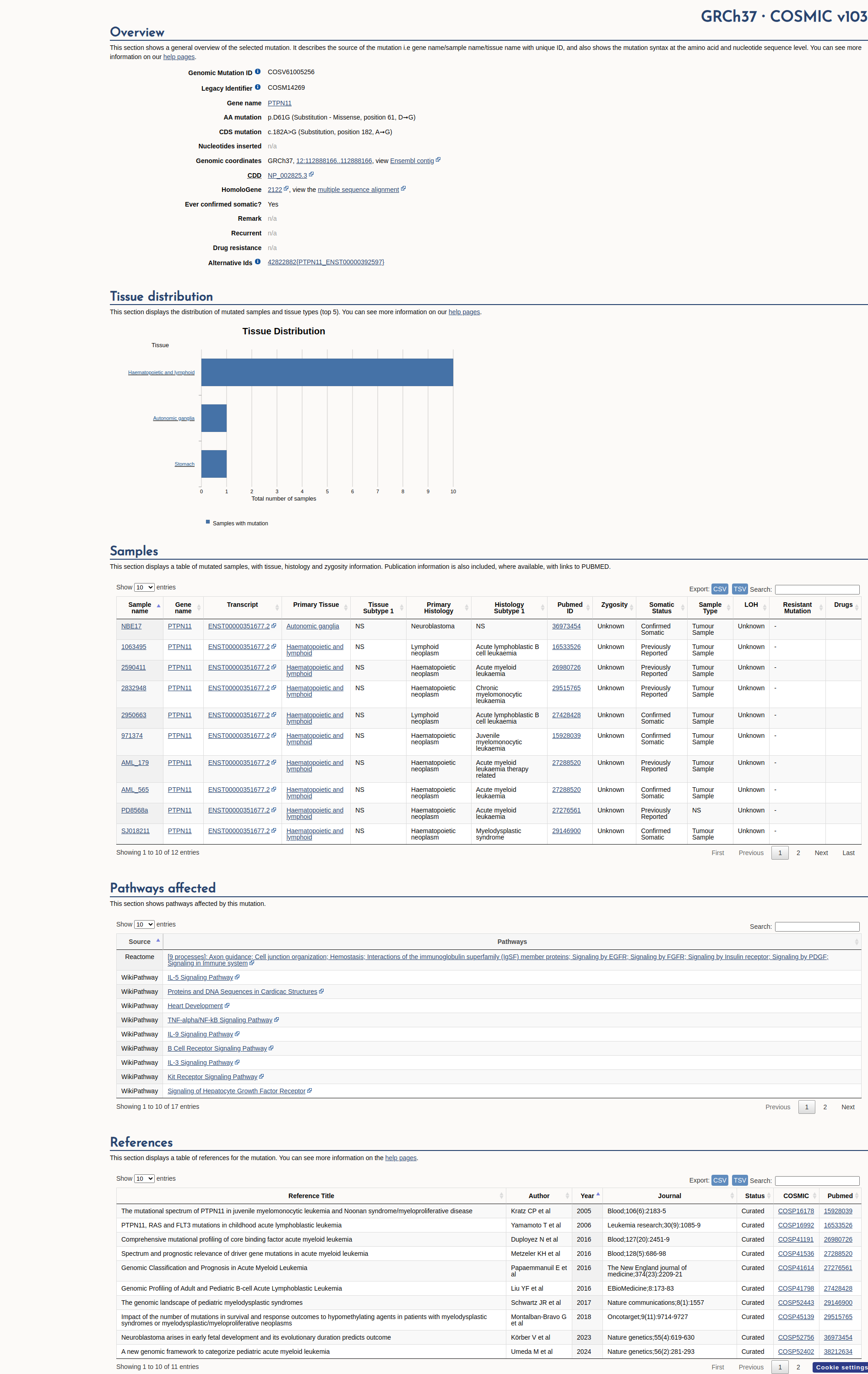
Functional Impact & Domains
Functional Domain
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.04 | -133 bp |
| Donor Loss (DL) | 0.0 | -4 bp |
| Acceptor Gain (AG) | 0.0 | -340 bp |
| Donor Gain (DG) | 0.0 | -250 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to standard ACMG guidelines, the rule for PVS1 is: "Null variant (nonsense, frameshift, start codon, ±1/2 splice sites) in a gene where loss of function is a known mechanism of disease." The evidence for this variant shows: NM_002834.3:c.182A>G is a missense change (p.D61G), not a null variant. Therefore, this criterion is not applied.
PS1 (Not Applied)
According to VCEP guidelines, the rule for PS1 (Strong) is: "Same amino acid change as a previously established pathogenic variant in PTPN11 regardless of nucleotide change." The evidence for this variant shows: p.D61G is the established change but no different nucleotide change producing the same amino acid substitution is reported. Therefore, this criterion is not applied.
PS2 (Not Applied)
According to VCEP guidelines, the rule for PS2 (Very Strong/Strong/Moderate strengths) concerns confirmed de novo occurrence. The evidence for this variant shows: no de novo data are available. Therefore, this criterion is not applied.
PS3 (Not Applied)
According to VCEP guidelines, the rule for PS3 is: "Moderate strength for two or more different approved assays; Supporting strength for one approved assay." The evidence for this variant shows: no functional assay data are available. Therefore, this criterion is not applied.
PS4 (Not Applied)
According to VCEP guidelines, the rule for PS4 (Strong ≥5 points; Moderate ≥3 points; Supporting ≥1 point) is based on case counts. The evidence for this variant shows: no quantified proband/case data are available. Therefore, this criterion is not applied.
PM1 (Moderate)
According to VCEP guidelines, the rule for PM1 (Moderate) is: "Applicable to critical and well-established functional domains (directly interacting residues between N-SH2 and PTPN domains: AA 58-63)." The evidence for this variant shows: p.D61G lies within residues 58-63, a critical PTPN11 functional domain. Therefore, this criterion is applied at Moderate strength.
PM2 (Supporting)
According to VCEP guidelines, the rule for PM2 (Supporting) is: "The variant must be absent from controls (gnomAD)." The evidence for this variant shows: NM_002834.3:c.182A>G is not present in gnomAD. Therefore, this criterion is applied at Supporting strength.
PM3 (Not Applied)
According to standard ACMG guidelines, the rule for PM3 concerns detection in trans with a pathogenic variant for recessive disorders. The evidence for this variant shows: PTPN11 disorders are autosomal dominant and no trans data exist. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, the rule for PM4 is: "Protein length changes due to in-frame indels or stop-loss variants." The evidence for this variant shows: p.D61G is a missense substitution without length change. Therefore, this criterion is not applied.
PM5 (Moderate)
According to VCEP guidelines, the rule for PM5 (Moderate) is: "One [likely] pathogenic residue change at the same codon." The evidence for this variant shows: other missense changes at codon 61 (e.g., D61H, D61N) are established pathogenic. Therefore, this criterion is applied at Moderate strength.
PM6 (Not Applied)
According to standard ACMG guidelines, the rule for PM6 concerns assumed de novo occurrence without confirmation. The evidence for this variant shows: no de novo information is available. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to VCEP guidelines, the rule for PP1 (Supporting/Moderate/Strong) requires segregation data. The evidence for this variant shows: no familial segregation data are available. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to VCEP guidelines, the rule for PP2 (Supporting) is: "Missense Z-score >3.09 in gnomAD indicates missense constraint." The evidence for this variant shows: PTPN11 missense constraint Z-score is not provided. Therefore, this criterion is not applied.
PP3 (Not Applied)
According to VCEP guidelines, the rule for PP3 (Supporting) is: "For missense variants, REVEL ≥0.7." The evidence for this variant shows: mixed in silico predictions and REVEL score unavailable or below threshold. Therefore, this criterion is not applied.
PP4 (Not Applied)
According to standard ACMG guidelines, the rule for PP4 concerns phenotype specificity. The evidence for this variant shows: no detailed phenotype correlation provided. Therefore, this criterion is not applied.
PP5 (Supporting)
According to standard ACMG guidelines, the rule for PP5 is: "Reputable source reports variant as pathogenic but evidence is unavailable." The evidence for this variant shows: ClinVar lists p.D61G as Pathogenic from 25 submitters without accessible underlying data. Therefore, this criterion is applied at Supporting strength.
BA1 (Not Applied)
According to VCEP guidelines, the rule for BA1 (Stand Alone) is: "GnomAD filtering allele frequency ≥0.05%." The evidence for this variant shows: absent from population databases. Therefore, this criterion is not applied.
BS1 (Not Applied)
According to VCEP guidelines, the rule for BS1 (Strong) is: "GnomAD filtering allele frequency ≥0.025%." The evidence for this variant shows: absent from population databases. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to VCEP guidelines, the rule for BS2 (Strong/Supporting) concerns observation in healthy adults. The evidence for this variant shows: no such observations. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to standard ACMG guidelines, the rule for BS3 is: "Well-established functional studies show no deleterious effect." The evidence for this variant shows: no functional studies performed. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to standard ACMG guidelines, the rule for BS4 is: "Lack of segregation in affected family members." The evidence for this variant shows: no segregation data. Therefore, this criterion is not applied.
BP1 (Not Applied)
According to VCEP guidelines, the rule for BP1 (Supporting) applies to truncating variants in genes without established loss-of-function disease mechanism. The evidence for this variant shows: p.D61G is missense. Therefore, this criterion is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, the rule for BP2 concerns observation in cis/trans with a pathogenic variant for dominant/recessive patterns. The evidence for this variant shows: no such data. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, the rule for BP3 is: "In-frame deletions/insertions in repetitive regions." The evidence for this variant shows: it is a missense substitution, not an in-frame indel. Therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP guidelines, the rule for BP4 (Supporting) is: "For missense variants, REVEL ≤0.3." The evidence for this variant shows: REVEL score unavailable or above threshold. Therefore, this criterion is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, the rule for BP5 is: "Variant found in a case with an alternate molecular cause." The evidence for this variant shows: no alternate diagnosis described. Therefore, this criterion is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, the rule for BP6 is: "Reputable source reports variant as benign but evidence is unavailable." The evidence for this variant shows: ClinVar reports as pathogenic, not benign. Therefore, this criterion is not applied.
BP7 (Not Applied)
According to standard ACMG guidelines, the rule for BP7 is: "Synonymous variant with no predicted splicing impact and low conservation." The evidence for this variant shows: p.D61G is missense, not synonymous. Therefore, this criterion is not applied.