Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000314.7 | RefSeq Select | 8514 nt | 845–2056 |
| NM_000314.5 | Alternative | 8719 nt | 1032–2243 |
| NM_000314.4 | Alternative | 5572 nt | 1032–2243 |
| NM_000314.3 | Alternative | 3416 nt | 1032–2243 |
| NM_000314.6 | Alternative | 8718 nt | 1032–2243 |
| NM_000314.8 | MANE Select | 8515 nt | 846–2057 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Open""
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
The PTEN T321* variant is a truncating mutation that results in a premature stop codon, leading to the production of a C-terminally truncated PTEN protein. Functional studies indicate that this truncation results in a loss of PTEN phosphatase activity, as demonstrated by reduced activity in a yeast assay. Additionally, expression of similar truncating mutations in mouse embryonic fibroblasts has shown oncogenic properties, including increased genome fragility and impaired regulation of the PI3K/AKT pathway. These findings support a damaging effect of the PTEN T321* variant.
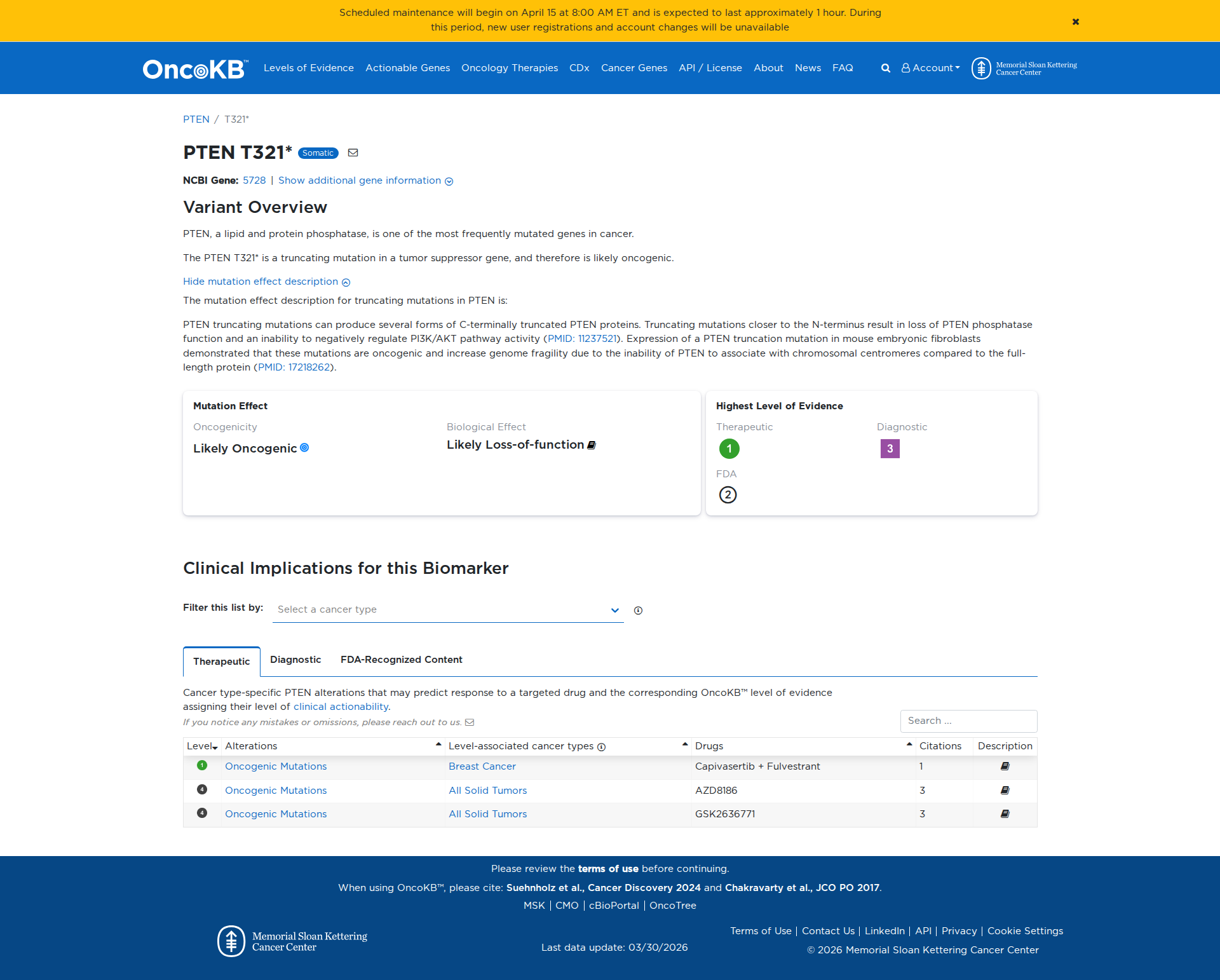
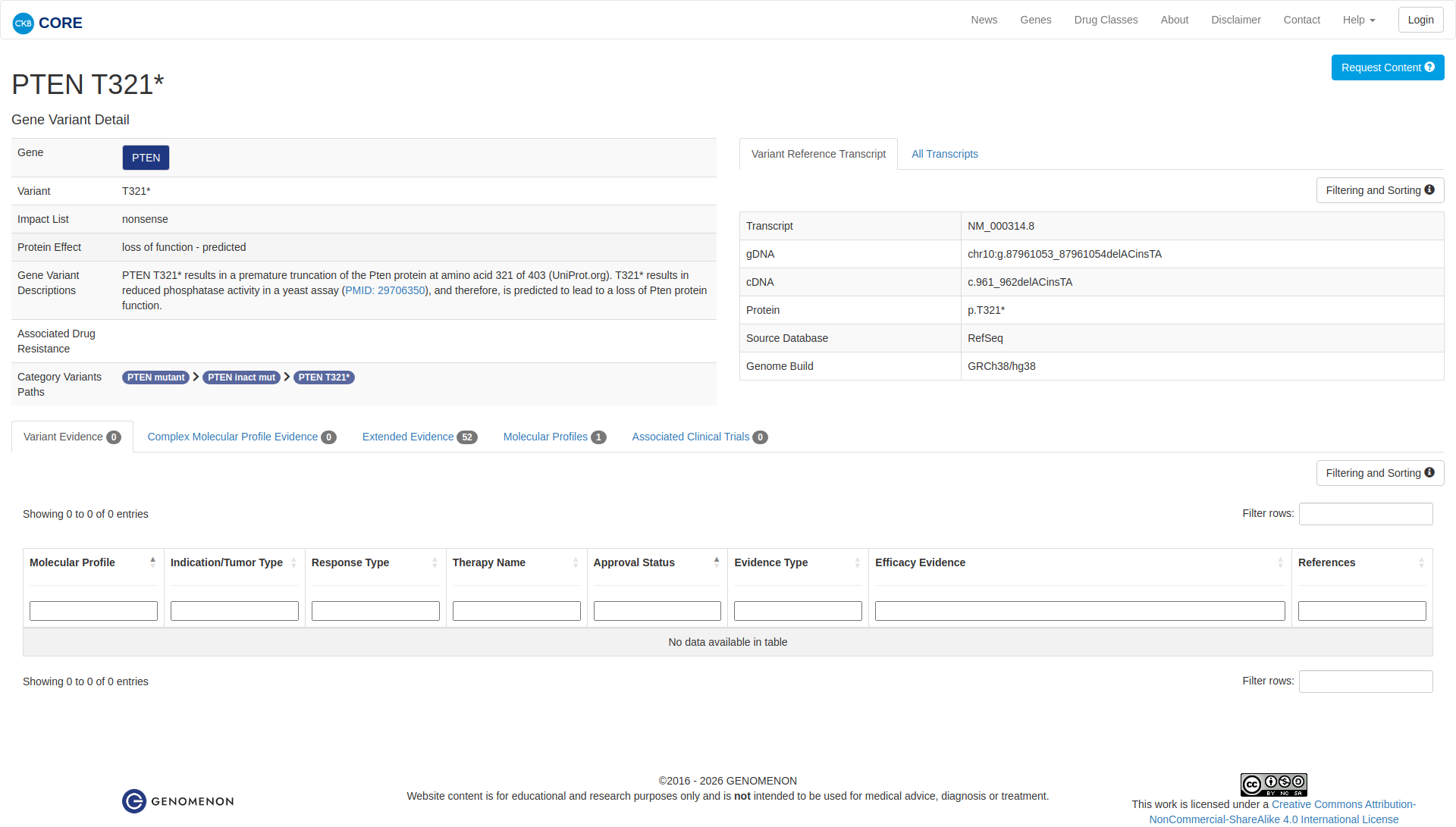
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.01 | -109 bp |
| Donor Loss (DL) | 0.0 | 72 bp |
| Acceptor Gain (AG) | 0.0 | 140 bp |
| Donor Gain (DG) | 0.01 | -132 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Very Strong)
According to VCEP guidelines, the rule for PVS1 is: "Very Strong Strength: Very Strong Use PTEN PVS1 decision tree." The evidence for this variant shows: c.955_956dup (p.T321*) introduces a premature stop codon resulting in loss of PTEN function. Therefore, this criterion is applied at Very Strong strength because the variant is a null variant in PTEN, a gene where loss-of-function is a known mechanism of disease.
PS1 (Not Applied)
According to VCEP guidelines, the rule for PS1 is: "Strong Same amino acid change as a previously established pathogenic variant regardless of nucleotide change OR different variant at same nucleotide position as a pathogenic splicing variant..." The evidence for this variant shows: it is a novel truncating variant not matching any known amino acid change. Therefore, this criterion is not applied.
PS2 (Not Applied)
According to VCEP guidelines, the rule for PS2 is: "Very Strong Two proven OR four assumed OR one proven + two assumed de novo observations..." The evidence for this variant shows: no de novo occurrence data are available. Therefore, this criterion is not applied.
PS3 (Moderate)
According to PTEN Pre-processing guidelines, the finding for PS3 is: "PS3_Moderate evidence added based on high-confidence functional score (-3.5048) < threshold (-1.11)." The evidence for this variant shows: functional studies demonstrate loss of PTEN phosphatase activity consistent with a score of -3.5048. Therefore, this criterion is applied at Moderate strength.
PS4 (Not Applied)
According to VCEP guidelines, the rule for PS4 is: "Very Strong Probands with specificity score ≥16 OR prevalence significantly increased in affected compared to controls." The evidence for this variant shows: no case-level prevalence or specificity scoring data. Therefore, this criterion is not applied.
PM1 (Not Applied)
According to VCEP guidelines, the rule for PM1 is: "Moderate Located in a mutational hot spot and/or critical and well-established functional domain (residues 90-94, 123-130, 166-168)." The evidence for this variant shows: p.T321* lies outside these domains. Therefore, this criterion is not applied.
PM2 (Supporting)
According to VCEP guidelines, the rule for PM2 is: "Supporting Strength: Absent in population Databases present at <0.00001 allele frequency in gnomAD." The evidence for this variant shows: absent from gnomAD. Therefore, this criterion is applied at Supporting strength.
PM3 (Not Applied)
According to standard ACMG guidelines, the rule for PM3 is: "Detected in trans with a pathogenic variant in a gene for a recessive disorder." The evidence for this variant shows: PTEN disorders are autosomal dominant and no trans data exist. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to VCEP guidelines, the rule for PM4 is: "Moderate Protein length changes due to in-frame deletions/insertions in a non-repeat region or stop-loss variants." The evidence for this variant shows: it is a frameshift/nonsense, not an in-frame indel. Therefore, this criterion is not applied.
PM5 (Not Applied)
According to VCEP guidelines, the rule for PM5 is: "Moderate Missense change at an amino acid residue where a different missense change determined to be pathogenic has been seen before." The evidence for this variant shows: it is a truncating variant, not missense. Therefore, this criterion is not applied.
PM6 (Not Applied)
According to VCEP guidelines, the rule for PM6 is: "Very Strong Two proven OR four assumed OR one proven + two assumed de novo observations..." The evidence for this variant shows: no assumed de novo data are available. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to VCEP guidelines, the rule for PP1 is: "Supporting Co-segregation with disease in multiple affected family members, with 3 or 4 meioses observed." The evidence for this variant shows: no segregation data. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, the rule for PP2 is: "Supporting Missense variant in a gene that has a low rate of benign missense variation and where missense variants are a common mechanism of disease." The evidence for this variant shows: it is a truncating variant, not missense. Therefore, this criterion is not applied.
PP3 (Not Applied)
According to VCEP guidelines, the rule for PP3 is: "Supporting Multiple lines of computational evidence support a deleterious effect on the gene or gene product." The evidence for this variant shows: SpliceAI score 0.01 and no deleterious in silico predictions; computational data are insufficient. Therefore, this criterion is not applied.
PP4 (Not Applied)
According to standard ACMG guidelines, the rule for PP4 is: "Supporting Patient's phenotype or family history is highly specific for a disease with a single genetic etiology." The evidence for this variant shows: no phenotype data. Therefore, this criterion is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines, the rule for PP5 is: "Supporting Reputable source reports variant as pathogenic." The evidence for this variant shows: not found in ClinVar or other databases. Therefore, this criterion is not applied.
BA1 (Not Applied)
According to VCEP guidelines, the rule for BA1 is: "Stand Alone gnomAD Filtering allele frequency >0.00056." The evidence for this variant shows: absent from gnomAD. Therefore, this criterion is not applied.
BS1 (Not Applied)
According to VCEP guidelines, the rule for BS1 is: "Strong gnomAD Filtering allele frequency from 0.000043 up to 0.00056." The evidence for this variant shows: absent from gnomAD. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to VCEP guidelines, the rule for BS2 is: "Strong Observed in the homozygous state in a healthy or PHTS-unaffected individual..." The evidence for this variant shows: no homozygous observations. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to VCEP guidelines, the rule for BS3 is: "Strong Well-established functional studies shows no damaging effect on protein function." The evidence for this variant shows: functional studies demonstrate damaging effect. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to VCEP guidelines, the rule for BS4 is: "Strong Lack of segregation in affected members of two or more families." The evidence for this variant shows: no segregation studies. Therefore, this criterion is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, the rule for BP1 is: "Supporting Missense variant in a gene for which primarily truncating variants are known to cause disease." The evidence for this variant shows: it is truncating. Therefore, this criterion is not applied.
BP2 (Not Applied)
According to VCEP guidelines, the rule for BP2 is: "Supporting Observed in trans with a pathogenic or likely pathogenic PTEN variant..." The evidence for this variant shows: no trans observations. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, the rule for BP3 is: "Supporting In-frame deletions/insertions in a repetitive region without a known function." The evidence for this variant shows: it is a frameshift/nonsense. Therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP guidelines, the rule for BP4 is: "Supporting Multiple lines of computational evidence suggest no impact on gene or gene product." The evidence for this variant shows: computational data indicate no splicing impact but the variant is truncating. Therefore, this criterion is not applied.
BP5 (Not Applied)
According to VCEP guidelines, the rule for BP5 is: "Supporting Variant found in a case with an alternate molecular basis for disease." The evidence for this variant shows: no alternate molecular basis data. Therefore, this criterion is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, the rule for BP6 is: "Supporting Reputable source reports variant as benign." The evidence for this variant shows: not found in databases. Therefore, this criterion is not applied.
BP7 (Not Applied)
According to VCEP guidelines, the rule for BP7 is: "Supporting A synonymous or intronic variant with no splicing impact." The evidence for this variant shows: it is a truncating variant. Therefore, this criterion is not applied.