Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenVariant summary: TP53 c.461G>A (p.Gly154Asp) results in a non-conservative amino acid change located in the p53, DNA-binding domain of the encoded protein sequence. Four of five in-silico tools predict a damaging effect of the variant on protein function. The variant allele was found at a frequency of 8e-06 in 251272 control chromosomes (gnomAD). The available data on variant occurrences in the general population are insufficient to allow any conclusion about variant significance. c.461G>A has been reported in the literature in an individual with a personal and family history of breast cancer (Zanti_2020). This report does not provide unequivocal conclusions about association of the variant with Li-Fraumeni Syndrome. Publications report experimental evidence evaluating an impact on protein function, showing either partially reduced function or no deleterious effect (e.g. Kato_2003, Giacomelli_2018, Kotler_2018). The following publications have been ascertained in the context of this evaluation (PMID: 33120919, 12826609, 30224644, 29979965). ClinVar contains an entry for this variant (Variation ID: 237950), including an evaluation from a ClinGen expert panel which classified the variant as uncertain significance. Based on the evidence outlined above, the variant was classified as uncertain significance.
This missense variant replaces glycine with aspartic acid at codon 154 of the TP53 protein. Computational prediction suggests that this variant may have deleterious impact on protein structure and function (internally defined REVEL score threshold >= 0.7, PMID: 27666373). Functional studies have shown that this variant functional in transactivation and cell growth and proliferation assays (PMID: 12826609, 329979965, 0224644). This variant has been reported in an individuals affected with triple negative breast cancer s in the literature (PMID: 33120919). This variant has been identified in 2/251272 chromosomes in the general population by the Genome Aggregation Database (gnomAD). The available evidence is insufficient to determine the role of this variant in disease conclusively. Therefore, this variant is classified as a Variant of Uncertain Significance.
This submission and the accompanying classification are no longer maintained by the submitter. For more information on current observations and classification, please contact variantquestions@myriad.com.
The p.G154D variant (also known as c.461G>A), located in coding exon 4 of the TP53 gene, results from a G to A substitution at nucleotide position 461. The glycine at codon 154 is replaced by aspartic acid, an amino acid with similar properties. This variant has been reported in a cohort of triple negative breast cancer patients from Cyprus (Zanti M et al. Cancers (Basel), 2020 Oct;12:). In another study, this variant was reported in 0/60,466 breast cancer cases and in 2/53,461 controls (Dorling et al. N Engl J Med. 2021 02;384:428-439). This variant is in the DNA binding domain of the TP53 protein and is reported to have partially functional transactivation in yeast based assays (Kato S et al. Proc. Natl. Acad. Sci. USA. 2003 Jul;100:8424-9). Studies conducted in human cell lines indicate this alteration is proficient at growth suppression and has no dominant negative effect (Kotler E et al. Mol.Cell. 2018 Jul;71:178-190.e8; Giacomelli AO et al. Nat. Genet. 2018 Oct;50:1381-1387). Based on internal structural analysis, this variant is not anticipated to result in a significant decrease in structural stability (Cho Y, Science 1994 Jul; 265(5170):346-55). In addition, this alteration did not segregate with disease in one family tested in our laboratory (Ambry internal data). This amino acid position is well conserved in available vertebrate species. In addition, this alteration is predicted to be deleterious by in silico analysis. Since supporting evidence is limited at this time, the clinical significance of this alteration remains unclear.
This sequence change replaces glycine, which is neutral and non-polar, with aspartic acid, which is acidic and polar, at codon 154 of the TP53 protein (p.Gly154Asp). This variant is present in population databases (rs762846821, gnomAD 0.002%). This missense change has been observed in individual(s) with acute myeloid leukemia and/or triple negative breast cancer (PMID: 27276561, 33120919). ClinVar contains an entry for this variant (Variation ID: 237950). Invitae Evidence Modeling incorporating data from in vitro experimental studies (PMID: 12826609, 29979965, 30224644) indicates that this missense variant is not expected to disrupt TP53 function with a negative predictive value of 97.5%. Experimental studies have shown that this missense change does not substantially affect TP53 function (PMID: 12826609, 29979965, 30224644). In summary, the available evidence is currently insufficient to determine the role of this variant in disease. Therefore, it has been classified as a Variant of Uncertain Significance.
"This variant has been reported in ClinVar as Uncertain significance (9 clinical laboratories) and as Likely pathogenic (1 clinical laboratories) and as Uncertain Significance by ClinGen TP53 Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open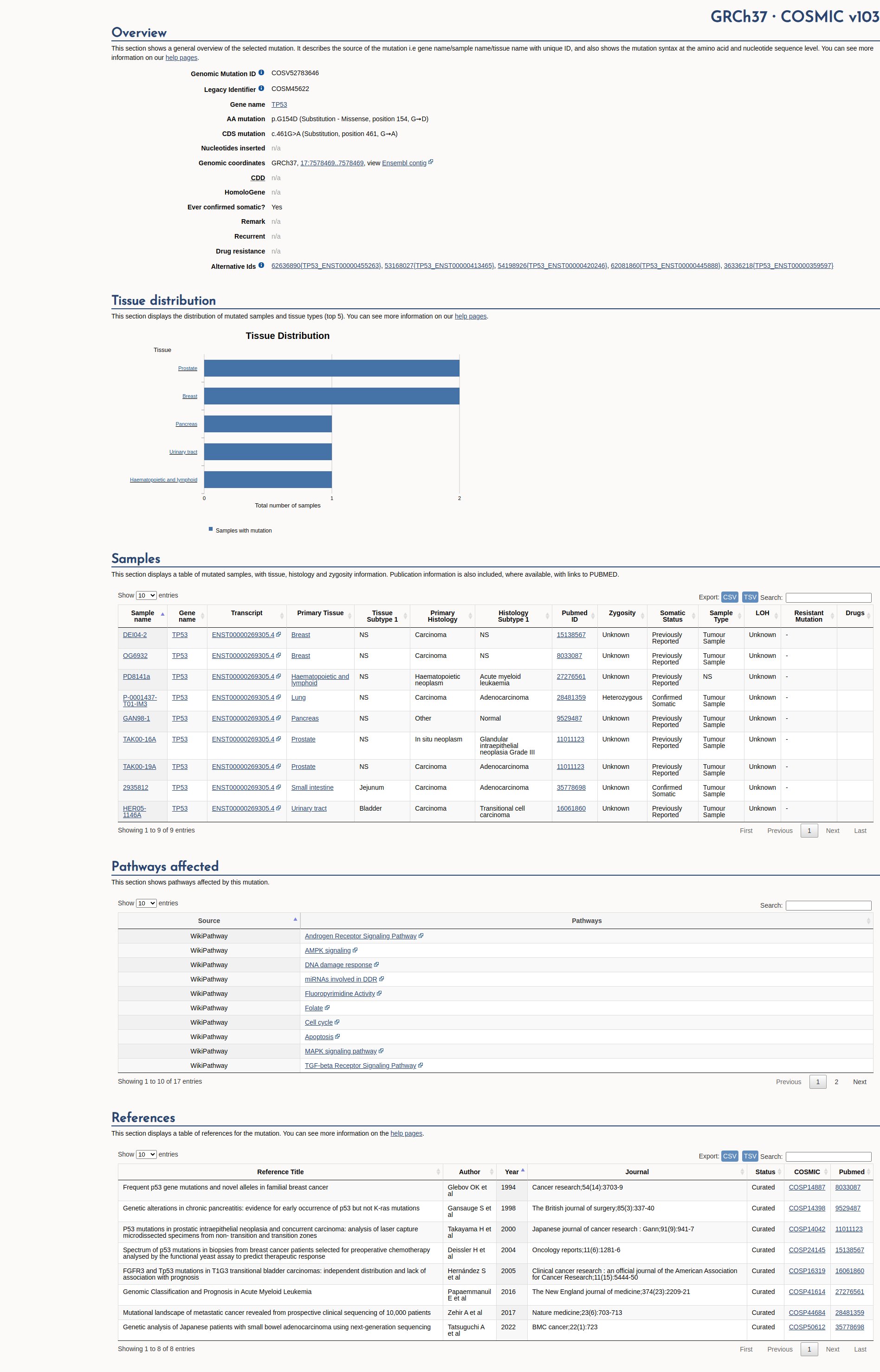
Functional Impact & Domains
Functional Domain
The TP53 G154D variant has been functionally characterized and demonstrated to be inactivating. In vivo studies using yeast models have shown that this mutation results in a partial loss of transactivational activity compared to the wildtype, indicating a damaging effect on the protein's function.
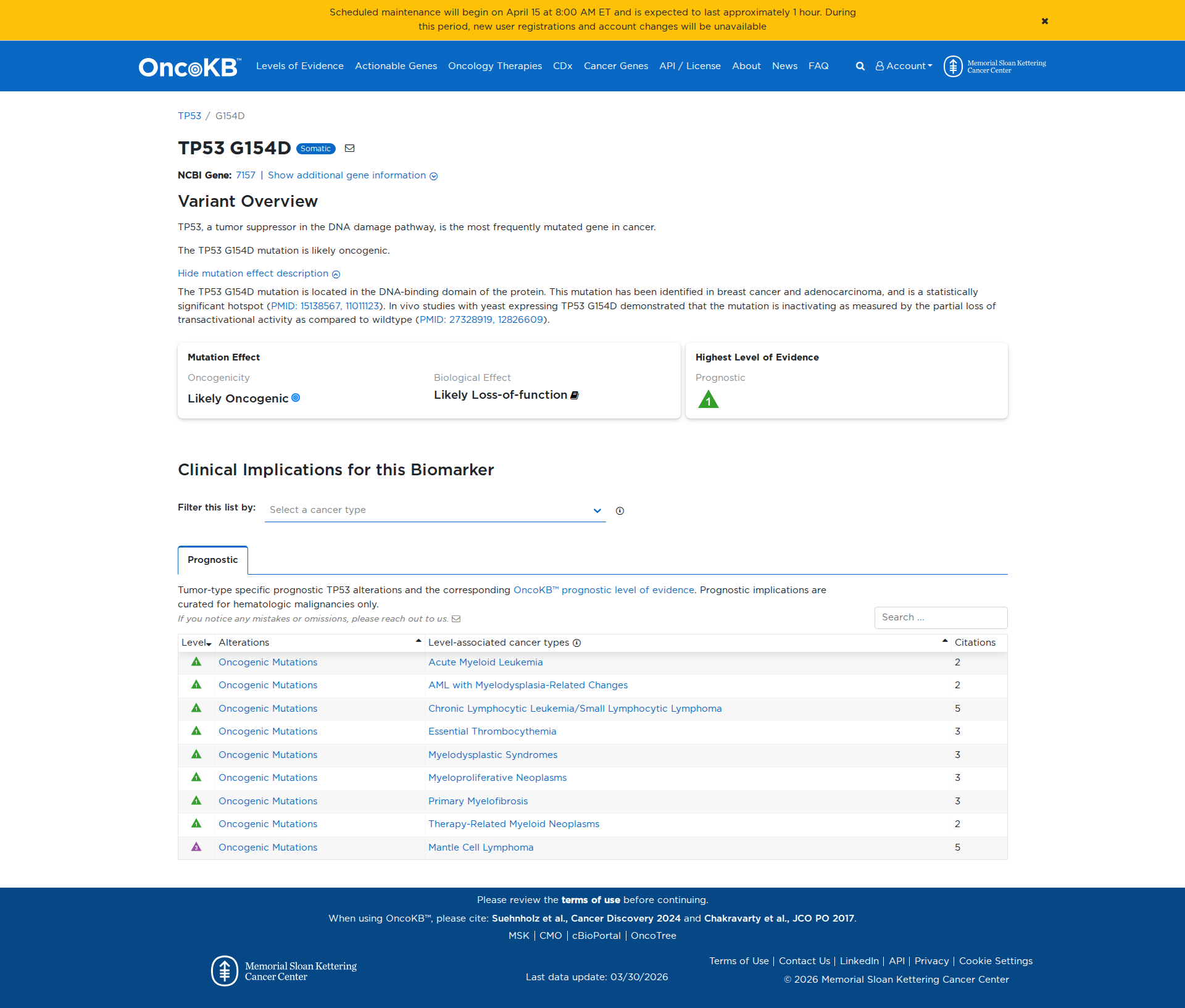
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.01 | -197 bp |
| Donor Loss (DL) | 0.01 | -297 bp |
| Acceptor Gain (AG) | 0.0 | 85 bp |
| Donor Gain (DG) | 0.0 | 185 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, "PVS1 not applicable or not evaluated." The evidence for this variant shows it is a missense change (G154D), not a null variant. Therefore, this criterion is not applied.
PS1 (Not Applied)
According to VCEP guidelines, "PS1 can be applied to variants asserted as Pathogenic following the TP53 VCEP’s specifications." The evidence for this variant shows no other variant at codon 154 classified as Pathogenic. Therefore, PS1 is not applied.
PS2 (Not Applied)
According to VCEP guidelines, "PS2 requires de novo confirmation with points-based system (Very Strong ≥8 points, Strong 4–7 points, etc.)." No de novo data are available for this variant. Therefore, PS2 is not applied.
PS3 (Not Applied)
According to VCEP guidelines, "PS3_Strong: Non-functional on Kato et al. data AND loss of function on another assay. PS3_Moderate: Partially functional on Kato et al. data AND LOF on Giacomelli et al. data AND/OR another assay." The evidence shows a single yeast model assay with partial loss of transactivational activity, but no Kato or Giacomelli data. Therefore, PS3 is not applied.
PS4 (Not Applied)
According to VCEP guidelines, "PS4 requires case points (Very Strong ≥8 points, Strong 4–7.5 points, etc.)." No proband or case-level data are available. Therefore, PS4 is not applied.
PM1 (Not Applied)
According to VCEP guidelines, "PM1: Moderate for missense variants within hotspot codons 175, 245, 248, 249, 273, 282." Codon 154 is outside these hotspots. Therefore, PM1 is not applied.
PM2 (Supporting)
According to VCEP guidelines, "PM2: Supporting. Apply when allele frequency <0.00003 in gnomAD or another large population database." The evidence shows MAF=0.00000796 (0.000796%), which is below 0.00003. Therefore, PM2 is applied at Supporting strength.
PM3 (Not Applied)
According to standard ACMG guidelines, PM3 applies to recessive disorders with variants detected in trans. TP53-associated disease is dominant and no trans data are available. Therefore, PM3 is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, PM4 applies to protein length changes such as in-frame indels. This is a missense variant without indel or splicing alteration. Therefore, PM4 is not applied.
PM5 (Not Applied)
According to VCEP guidelines, "PM5: Moderate for missense variants at a residue with ≥1 other pathogenic variant." No other pathogenic missense variants at codon 154 are documented. Therefore, PM5 is not applied.
PM6 (Not Applied)
According to VCEP guidelines, "PM6: De novo without confirmation of paternity/maternity (points-based)." No de novo data are available. Therefore, PM6 is not applied.
PP1 (Not Applied)
According to VCEP guidelines, "PP1 requires cosegregation evidence across meioses (Supporting 3–4, Moderate 5–6, Strong ≥7)." No family segregation data are available. Therefore, PP1 is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, PP2 applies to genes with low rate of benign missense variation and where missense is common mechanism. TP53 has many benign and pathogenic missense variants. Therefore, PP2 is not applied.
PP3 (Not Applied)
According to VCEP guidelines, "PP3: Moderate when BayesDel ≥0.16 and no predicted splice effect; Supporting at lower thresholds." BayesDel score is not provided and computational predictions are mixed. Therefore, PP3 is not applied.
PP4 (Not Applied)
According to VCEP guidelines, "PP4 requires disease-specific phenotype observations." No phenotype or tumor VAF data are provided. Therefore, PP4 is not applied.
BA1 (Not Applied)
According to VCEP guidelines, "BA1: Stand Alone when allele frequency ≥0.001 in gnomAD subpopulation." MAF is 0.00000796, below threshold. Therefore, BA1 is not applied.
BS1 (Not Applied)
According to VCEP guidelines, "BS1: Strong when allele frequency ≥0.0003 but <0.001." MAF is 0.00000796, below 0.0003. Therefore, BS1 is not applied.
BS2 (Not Applied)
According to VCEP guidelines, "BS2: Strong when ≥8 unaffected older females without cancer are observed." No such data are available. Therefore, BS2 is not applied.
BS3 (Not Applied)
According to VCEP guidelines, "BS3 applies when functional assays show preserved function on Kato and another assay." The variant shows partial loss of function. Therefore, BS3 is not applied.
BS4 (Not Applied)
According to VCEP guidelines, "BS4: Strong when lack of segregation in affected family members." No segregation data exist. Therefore, BS4 is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines, BP1 applies to missense variants in a gene where only truncating variants are pathogenic. TP53 has many pathogenic missense variants. Therefore, BP1 is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, BP2 applies when variant is in trans with a pathogenic variant in recessive disorder. TP53 disease is dominant. Therefore, BP2 is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, BP3 applies to in-frame indels in repetitive regions. This is a missense variant. Therefore, BP3 is not applied.
BP4 (Not Applied)
According to VCEP guidelines, "BP4: Moderate when BayesDel ≤ –0.008 and no predicted splicing impact; Supporting for BayesDel <0.16 and no splicing effect." BayesDel score is not available and evidence is mixed. Therefore, BP4 is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, BP5 applies when variant found in case with other pathogenic variant. No such evidence is provided. Therefore, BP5 is not applied.
BP6 (Not Applied)
According to ACMG guidelines, BP6 applies to reputable source assertion of benign without evidence. ClinVar entries are conflicting (uncertain and likely pathogenic). Therefore, BP6 is not applied.
BP7 (Not Applied)
According to VCEP guidelines, BP7 applies to synonymous or intronic variants with no splicing impact. This variant is missense. Therefore, BP7 is not applied.