Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_024675.4 | MANE Select | 4008 nt | 154–3714 |
| NM_024675.3 | RefSeq Select | 4069 nt | 201–3761 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe PALB2 c.3113G>A (p.Trp1038Ter) variant is a stop-gained variant that is predicted to result in a premature termination of the protein. Across a selection of the available literature, the c.3113G>A (p.Trp1038Ter) variant has been identified in a heterozygous state in at least 21 probands from 13 unrelated families (Rahman et al. 2007; Southey et al. 2010; Teo et al. 2013; Hartley et al. 2014). Affected individuals were found to have a variety of cancers including breast, ovarian, and pancreatic. The p.Trp1038Ter variant was absent from 1724 healthy control subjects and is reported at a frequency of 0.000125 in the African population of the Genome Aggregation Database. Experimental studies on RNA isolated from patient derived lymphoblastoid cell lines have shown that the p.Trp1038Ter variant caused altered splicing (Casadei et al. 2011; Teo et al. 2013). Although this variant has not been reported in any probands with Fanconi anemia, it is known that PALB2 heterozygous variants can also confer carrier status for Fanconi anemia. Due to the potential impact of stop-gained variants and the evidence from the literature, this variant is classified as pathogenic for PALB2-related disorders. This variant was observed by ICSL as part of a predisposition screen in an ostensibly healthy population.
Curators: Marc Tischkowitz, Arleen D. Auerbach. Submitters to LOVD: James Whitworth, kConFab - Heather Thorne, Marc Tischkowitz.
"This variant has been reported in ClinVar as Pathogenic (31 clinical laboratories) and as pathogenic (1 clinical laboratories) and as Pathogenic by ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open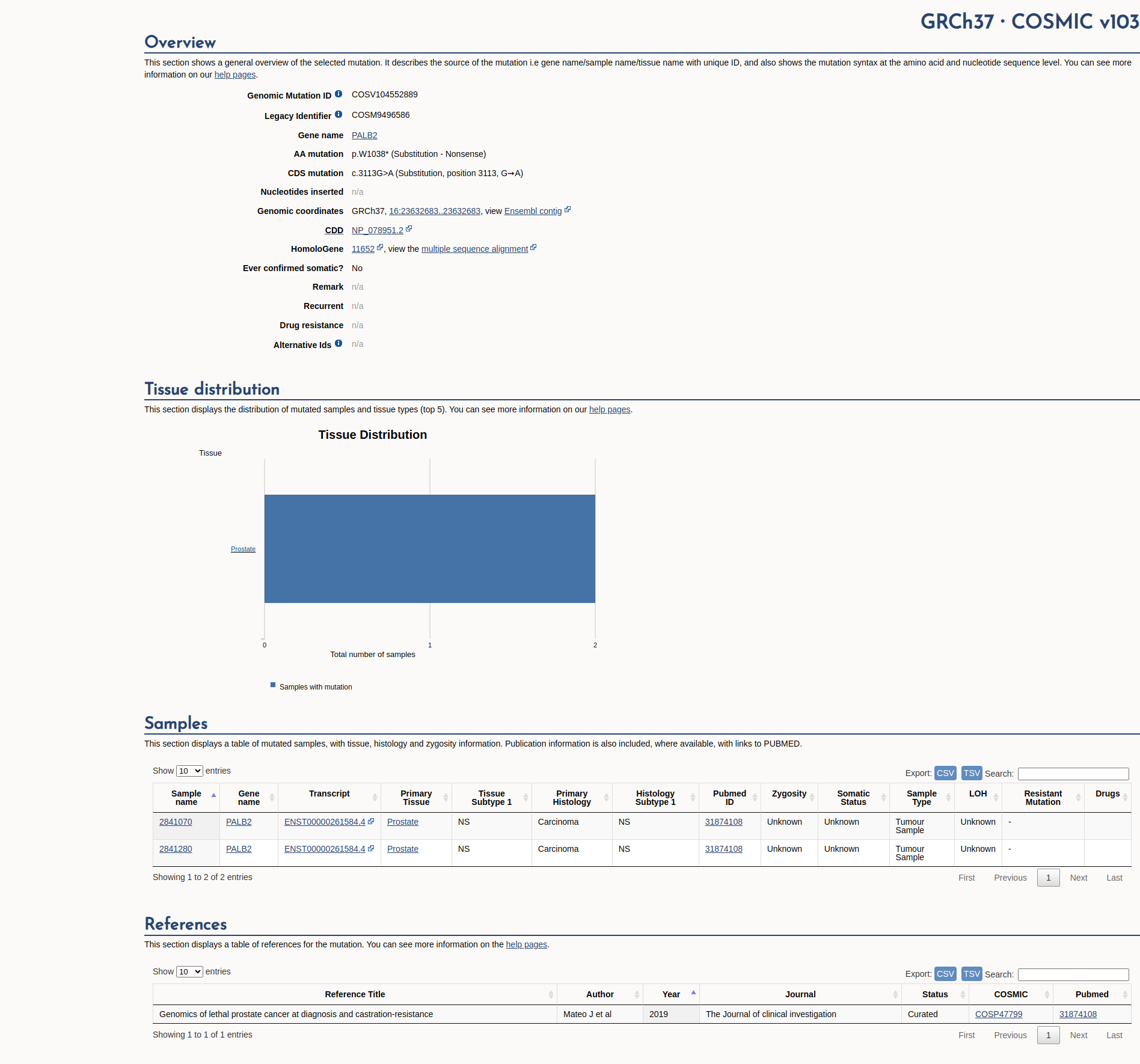
Functional Impact & Domains
Functional Domain
The PALB2 W1038* variant results in a premature truncation of the protein, leading to a loss of function. Functional studies demonstrate decreased protein stability, aberrant cytosolic accumulation, reduced BRCA2 binding, impaired RAD51 foci formation, and compromised homologous recombination. These findings support a damaging effect of the variant.
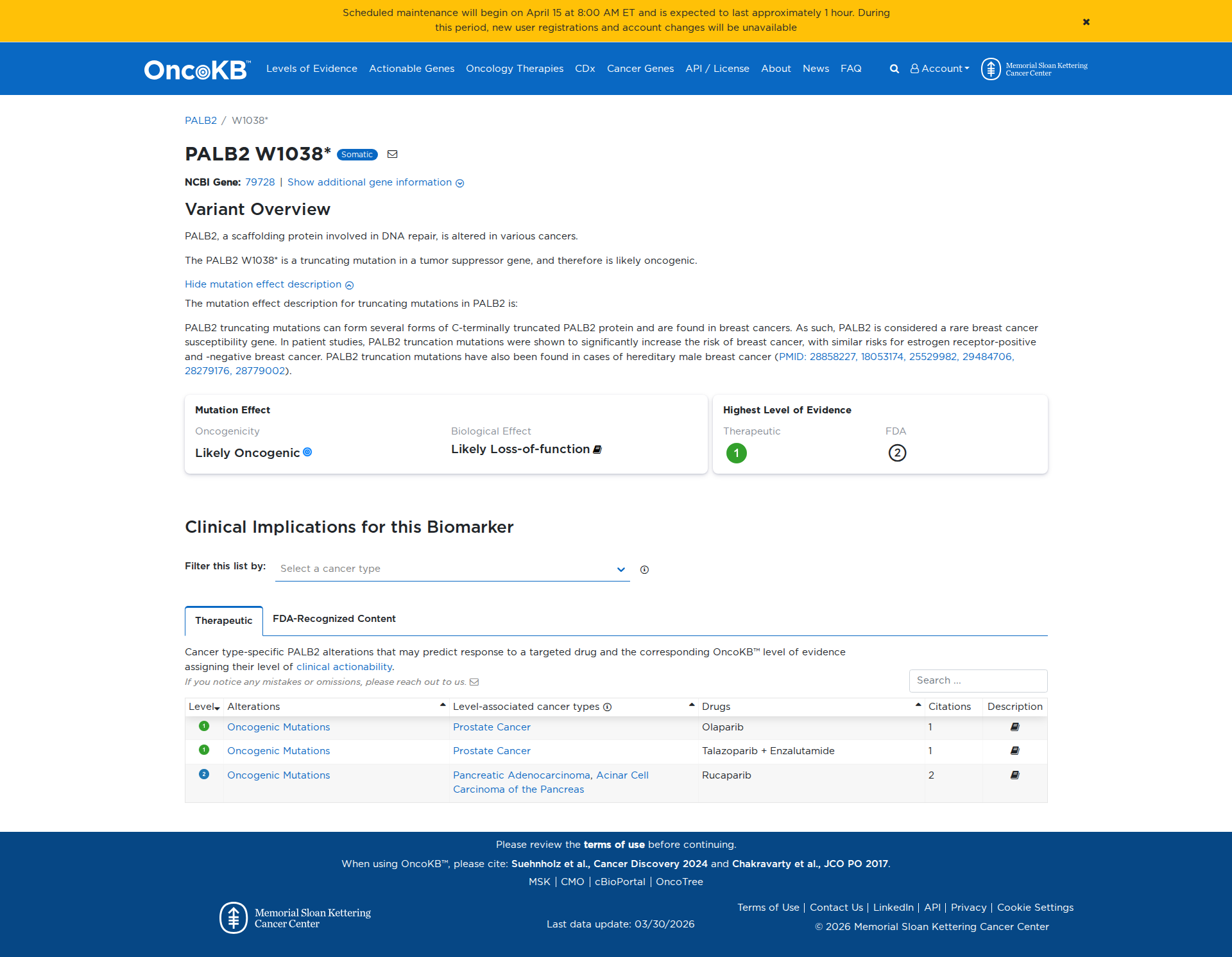
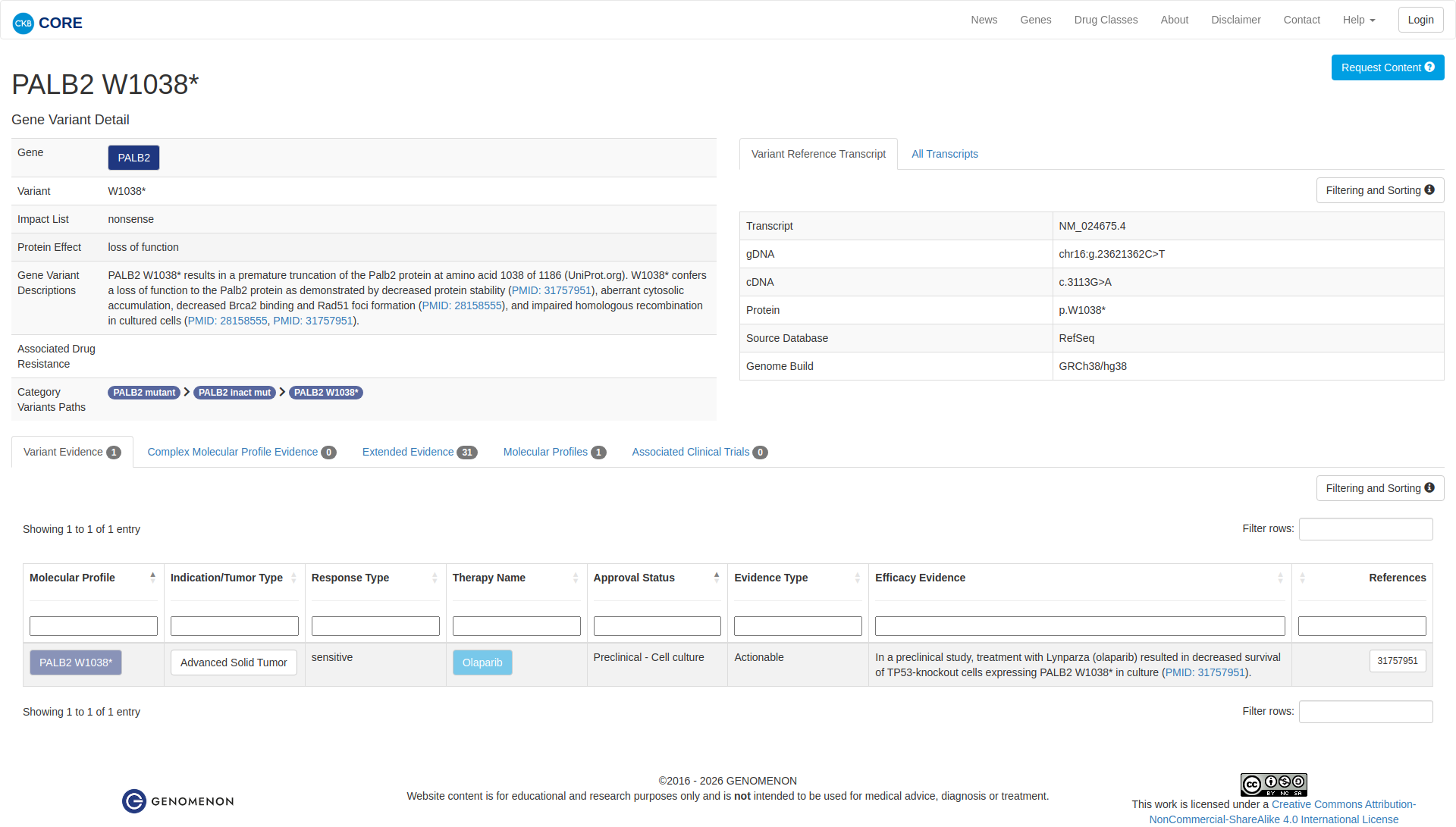
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.63 | 116 bp |
| Donor Loss (DL) | 0.74 | 0 bp |
| Acceptor Gain (AG) | 0.0 | 460 bp |
| Donor Gain (DG) | 0.17 | 31 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Very Strong)
According to VCEP guidelines, the rule/finding for PVS1 is: "Very Strong Strength: Very Strong Use PALB2 PVS1 Decision Tree Modification Type: Gene-specific,Strength". The evidence for this variant shows: NM_024675.4:c.3113G>A is a nonsense variant (W1038*) predicted to lead to loss of function in a gene where LOF is a known mechanism. Therefore, this criterion is applied at Very Strong strength because the variant is a truncating null variant not in the last exon, fitting the VCEP PVS1 decision tree.
PS1 (Not Applied)
According to VCEP guidelines for PS1 (splicing table), there is no matching previously established pathogenic variation at the same amino acid. The evidence for this variant shows: it is a novel nonsense change, not a recapitulation of a known amino acid substitution. Therefore, this criterion is not applied.
PS2 (Not Applied)
According to standard ACMG guidelines, the rule for PS2 is: "De novo (both maternity and paternity confirmed) in a patient with the disease and no family history." The evidence for this variant shows: no confirmed de novo occurrence data. Therefore, this criterion is not applied.
PS3 (Strong)
According to standard ACMG guidelines, the rule/finding for PS3 is: "Well-established in vitro or in vivo functional studies supportive of a damaging effect on the gene or gene product." The evidence for this variant shows: functional studies demonstrate decreased protein stability, aberrant cytosolic accumulation, reduced BRCA2 binding, impaired RAD51 foci formation, and compromised homologous recombination for W1038*. Therefore, this criterion is applied at Strong strength because multiple well-established assays show a damaging effect.
PS4 (Not Applied)
According to VCEP guidelines, the rule for PS4 is: "Strong Strength: Case-control studies; p-value ≤ .05 AND (Odds ratio, hazard ratio, or relative risk ≥3 OR lower 95% CI ≥1.5)." The evidence for this variant shows: no published case-control data. Therefore, this criterion is not applied.
PM1 (Not Applied)
According to standard ACMG guidelines, the rule for PM1 is: "Located in a mutational hot spot and/or critical and well-established functional domain without benign variation." The evidence for this variant shows: W1038* falls outside of defined mutational hot spots or critical domains with no established benign variation. Therefore, this criterion is not applied.
PM2 (Not Applied)
According to VCEP guidelines, the rule/finding for PM2 is: "Supporting Strength: Supporting Variant absent in gnomAD or present in ≤1/300,000 alleles Modification Type: Gene-specific,Strength". The evidence for this variant shows: MAF=0.00601% (17/282,756), which exceeds the VCEP threshold of ≤1/300,000 alleles. Therefore, this criterion is not applied.
PM3 (Not Applied)
According to VCEP guidelines for PM3 (Fanconi Anemia tables), the rule requires detection in trans with a pathogenic variant in a recessive disorder. The evidence for this variant shows: no phasing or biallelic data. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, the rule for PM4 is: "Protein length changes due to in-frame deletions/insertions or stop-loss variants are not directly applicable to truncating variants causing LOF." The evidence for this variant shows: it is a nonsense LOF variant, not an in-frame indel. Therefore, this criterion is not applied.
PM5 (Supporting)
According to VCEP guidelines, the rule/finding for PM5 is: "Supporting Strength: Apply to frameshifting or truncating variants with premature termination codons upstream of p.Tyr1183, based on location of the most C-terminal known pathogenic variant, p.Tyr1183* Modification Type: Gene-specific,Strength". The evidence for this variant shows: W1038* introduces a PTC upstream of Tyr1183. Therefore, this criterion is applied at Supporting strength because it meets the VCEP truncating variant location requirement.
PM6 (Not Applied)
According to standard ACMG guidelines, the rule for PM6 is: "Assumed de novo, but without confirmation of paternity and maternity." The evidence for this variant shows: no de novo assumption data. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to VCEP guidelines for PP1 (segregation), the rule requires evidence of segregation with disease in multiple affected family members. The evidence for this variant shows: no segregation data. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, the rule for PP2 is: "Missense variant in a gene with a low rate of benign missense variation and where missense variants are a common mechanism of disease." The evidence for this variant shows: it is a nonsense variant. Therefore, this criterion is not applied.
PP3 (Supporting)
According to VCEP guidelines, the rule/finding for PP3 is: "Supporting Strength: Protein: Do not use. RNA: At least one well-established in silico predictor (e.g. SpliceAI) shows impact on splicing Modification Type: General recommendation". The evidence for this variant shows: SpliceAI predicts a moderate splicing impact (donor loss score 0.74). Therefore, this criterion is applied at Supporting strength because well-established in silico prediction indicates splicing disruption.
PP4 (Not Applied)
According to standard ACMG guidelines, the rule for PP4 is: "Patient’s phenotype or family history is highly specific for a disease with a single genetic etiology." The evidence for this variant shows: no detailed phenotype/family history provided. Therefore, this criterion is not applied.
PP5 (Supporting)
According to standard ACMG guidelines, the rule/finding for PP5 is: "Reputable source recently reports variant as pathogenic, but the evidence is not available to the laboratory to perform an independent evaluation." The evidence for this variant shows: ClinVar entries from 31 clinical labs and ClinGen expert panel classify it as Pathogenic. Therefore, this criterion is applied at Supporting strength because multiple reputable sources report pathogenicity without publicly available primary evidence.
BA1 (Not Applied)
According to VCEP guidelines, the rule for BA1 is: "Stand Alone Strength: Allele frequency >0.1%". The evidence for this variant shows: MAF=0.00601%, below 0.1%. Therefore, this criterion is not applied.
BS1 (Not Applied)
According to VCEP guidelines, the rule for BS1 is: "Strong Strength: GnomAD Filtering Allele Frequency greater than expected for disease >0.01% Modification Type: Disease-specific,Gene-specific". The evidence for this variant shows: MAF=0.00601%, below 0.01%. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to VCEP guidelines for BS2 (Fanconi Anemia tables), the rule requires observation in healthy adults for a recessive disorder. The evidence for this variant shows: no such data. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to standard ACMG guidelines, the rule for BS3 is: "Well-established functional studies show no damaging effect on protein function or splicing." The evidence for this variant shows: functional studies demonstrate damaging effects. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to VCEP guidelines, the rule for BS4 is: "Strong Strength: Lack of segregation (LOD ≤ -1.28 or LR ≤ 0.053:1)". The evidence for this variant shows: no segregation data. Therefore, this criterion is not applied.
BP1 (Not Applied)
According to VCEP guidelines, the rule for BP1 is: "Supporting Strength: Apply to all missense variants." The evidence for this variant shows: it is a nonsense variant. Therefore, this criterion is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, the rule for BP2 is: "Observed in trans with a pathogenic variant for a dominant disorder or in cis with a pathogenic variant in any inheritance pattern." The evidence for this variant shows: no such data. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, the rule for BP3 is: "In-frame deletions/insertions in a repetitive region without a known function." The evidence for this variant shows: a nonsense change, not an in-frame indel. Therefore, this criterion is not applied.
BP4 (Not Applied)
According to VCEP guidelines, the rule for BP4 is: "Supporting Strength: Protein: Do not use. RNA: At least one well-established in silico predictor (e.g. SpliceAI) shows no impact on splicing." The evidence for this variant shows: SpliceAI predicts a splice impact. Therefore, this criterion is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines, the rule for BP5 is: "Variant found in a case with an alternate molecular basis for disease." The evidence for this variant shows: no alternate molecular basis. Therefore, this criterion is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, the rule for BP6 is: "Reputable source reports benign, but the evidence is not available to the laboratory to perform an independent evaluation." The evidence for this variant shows: no reputable benign classification. Therefore, this criterion is not applied.
BP7 (Not Applied)
According to VCEP guidelines, the rule for BP7 is: "Supporting Strength: Synonymous and deep intronic variants with observed lack of aberrant RNA defect." The evidence for this variant shows: it is a nonsense coding variant. Therefore, this criterion is not applied.