Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_007294.4 | MANE Select | 7088 nt | 114–5705 |
| NM_007294.2 | Alternative | 7191 nt | 201–5792 |
| NM_007294.3 | RefSeq Select | 7224 nt | 233–5824 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe BRCA1 c.131G>A (p.Cys44Tyr) variant has been reported in the published literature in individuals with ovarian cancer (PMID: 36367610 (2023), 31825140 (2019)) and breast cancer (PMID: 27083775 (2016)). Assessment of experimental evidence suggests this variant results in abnormal protein function (PMID: 21922593 (2011), 25823446 (2015), 27272900 (2016), 30209399 (2018)). This variant has not been reported in large, multi-ethnic general populations (Genome Aggregation Database, http://gnomad.broadinstitute.org). Analysis of this variant using bioinformatics tools for the prediction of the effect of amino acid changes on protein structure and function yielded predictions that this variant is damaging. Based on the available information, this variant is classified as pathogenic.
This submission and the accompanying classification are no longer maintained by the submitter. For more information on current observations and classification, please contact variantquestions@myriad.com.
IARC class based on posterior probability from multifactorial likelihood analysis, thresholds for class as per Plon et al. 2008 (PMID: 18951446). Class 5 based on posterior probability = 1
The p.C44Y variant (also known as c.131G>A), located in coding exon 2 of the BRCA1 gene, results from a G to A substitution at nucleotide position 131. The cysteine at codon 44 is replaced by tyrosine, an amino acid with highly dissimilar properties. This alteration, along with another alteration, p.C44S, at the same position, has been classified as definitely pathogenic (p>0.99) by multifactorial analysis, which integrates the following lines of evidence to produce a quantitative likelihood of pathogenicity: in silico prediction models, segregation with disease, tumor characteristics, mutation co-occurrence (Easton D et al. Am J Hum Genet. 2007;81:873-883; Vallee M et al. Hum Mutat. 2012 Jan;33(1):22-8; Lindor NM et al. Hum. Mutat., 2012 Jan;33:8-21). This alteration was shown to disrupt the function of the protein in multiple functional studies using different assays (Millot GA et al. Hum. Mutat., 2011 Dec;32:1470-80; Starita LM et al. Genetics, 2015 Jun;200:413-22; Thouvenot P et al. PLoS Genet., 2016 Jun;12:e1006096). In addition, a recent study found that this nucleotide substitution is deleterious in a high throughput genome editing haploid cell survival assay (Findlay GM et al. Nature. 2018 10;562(7726):217-222). This amino acid position is highly conserved in available vertebrate species. In addition, this alteration is predicted to be deleterious by BayesDel in silico analysis. Based on the supporting evidence, this alteration is interpreted as a disease-causing mutation.
"This variant has been reported in ClinVar as Pathogenic (9 clinical laboratories) and as pathogenic (1 clinical laboratories) and as Uncertain significance (1 clinical laboratories) and as Pathogenic by Evidence-based Network for the Interpretation of Germline Mutant Alleles (ENIGMA) expert panel."
COSMIC Somatic Evidence
Open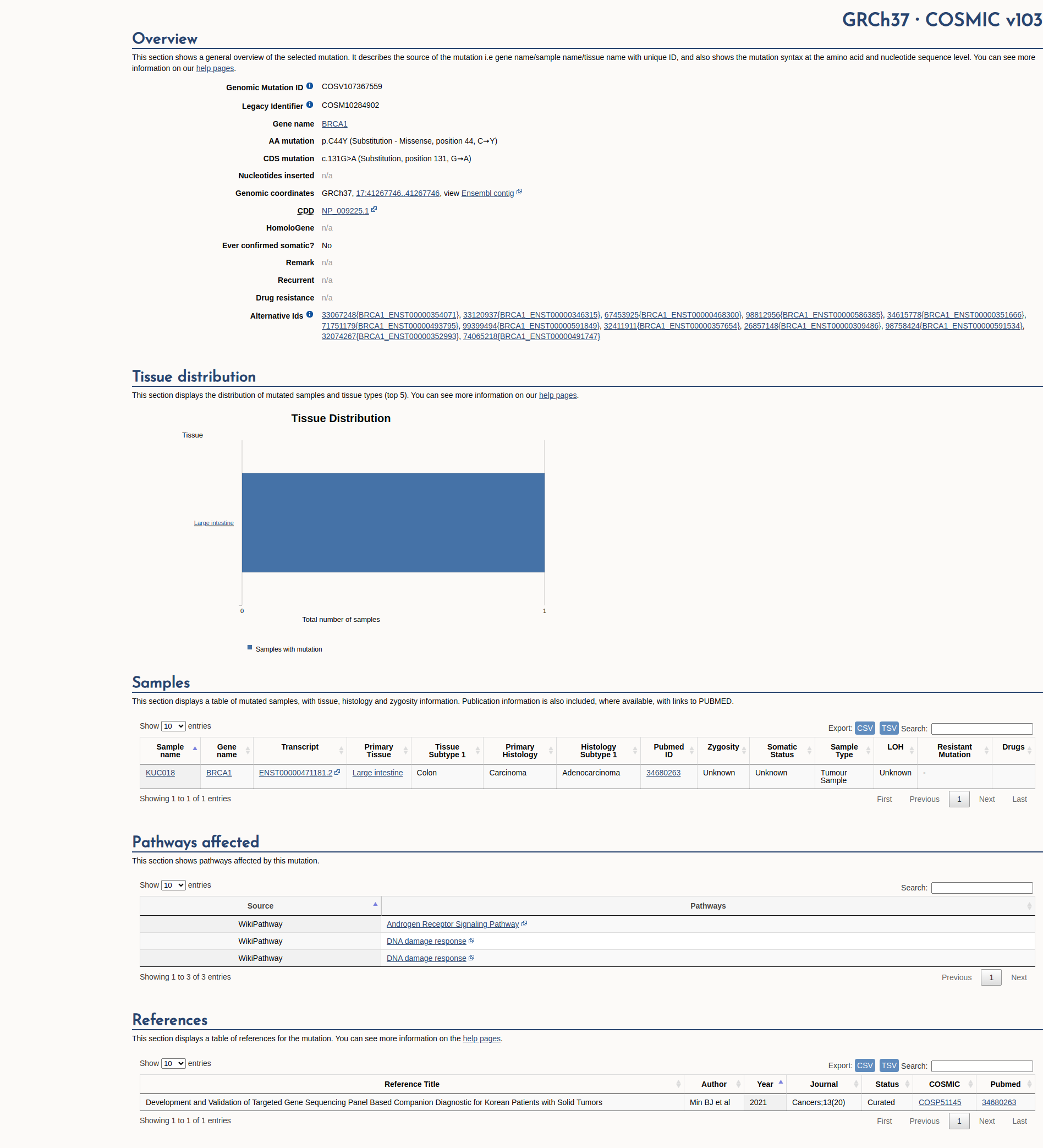
Functional Impact & Domains
Functional Domain
The BRCA1 C44Y variant has been functionally characterized and shown to have a damaging effect. It promotes cell colony growth in a yeast-based assay and disrupts the binding of BRCA1 to the E2 ubiquitin-conjugating enzyme UbcH5a, indicating a potential oncogenic role. However, the requirement of E3 ligase activity for BRCA1 tumor suppression remains unclear.
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 34 bp |
| Donor Loss (DL) | 0.0 | 1 bp |
| Acceptor Gain (AG) | 0.01 | 50 bp |
| Donor Gain (DG) | 0.0 | -3 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines, PVS1 applies to null variants in a gene where loss of function is a known mechanism (‘Very Strong Null variant…’). This variant is a missense change (C44Y), not a null variant. Therefore, PVS1 is not applied.
PS1 (Not Applied)
According to VCEP guidelines, PS1 applies when a variant results in the same amino acid change as a previously established pathogenic variant. There is no record of another nucleotide change producing C44Y. Therefore, PS1 is not applied.
PS2 (Not Applied)
According to standard ACMG guidelines, PS2 applies for confirmed de novo variants. No parental testing or de novo data are available for this variant. Therefore, PS2 is not applied.
PS3 (Not Applied)
According to VCEP guidelines, PS3 requires well-established functional assays calibrated per Specification Table 9. The available yeast-based assay is not a VCEP-calibrated assay. Therefore, PS3 is not applied.
PS4 (Not Applied)
According to VCEP guidelines, PS4 applies when prevalence in affected individuals is significantly increased over controls. No case-control or cohort data are available. Therefore, PS4 is not applied.
PM1 (Not Applied)
According to VCEP guidelines, PM1 is not specified for BRCA1 missense in functional domains. Although C44Y lies in the RING domain, VCEP does not designate PM1 for this context. Therefore, PM1 is not applied.
PM2 (Supporting)
According to VCEP guidelines, PM2_Supporting: ‘Absent from controls in gnomAD v2.1 and v3.1…’. The variant is not observed in gnomAD (MAF = 0%). Therefore, PM2 is applied at Supporting strength.
PM3 (Not Applied)
According to VCEP guidelines, PM3 applies to compound heterozygous or homozygous variants in Fanconi anemia phenotype. No FA phenotype or second variant data are available. Therefore, PM3 is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines, PM4 applies to protein length changes. This is a missense substitution without protein length change. Therefore, PM4 is not applied.
PM5 (Not Applied)
According to standard ACMG guidelines, PM5 applies when a different pathogenic missense at the same codon is known. No other pathogenic variant at codon 44 is reported. Therefore, PM5 is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines, PM6 applies to assumed de novo variants without confirmation. There is no de novo evidence. Therefore, PM6 is not applied.
PP1 (Not Applied)
According to VCEP guidelines, PP1 applies for co-segregation evidence. No family segregation data are available. Therefore, PP1 is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines, PP2 applies for missense variants in genes with low rate of benign missense variation. BRCA1 has both benign and pathogenic missense. Therefore, PP2 is not applied.
PP3 (Not Applied)
According to VCEP guidelines, PP3 applies when missense inside a functional domain has BayesDel no-AF ≥0.28 or SpliceAI ≥0.2. Computational evidence here shows low CADD, SpliceAI 0.01, and no BayesDel support. Therefore, PP3 is not applied.
PP4 (Not Applied)
According to VCEP guidelines, PP4 applies to multifactorial clinical evidence for breast cancer. No likelihood ratio‐based clinical data are available. Therefore, PP4 is not applied.
PP5 (Not Applied)
According to ClinGen recommendations (standard ACMG), PP5 (reputable source pathogenic without evidence) is discouraged. Although ClinVar reports pathogenic assertions, VCEP does not use PP5. Therefore, PP5 is not applied.
BA1 (Not Applied)
According to VCEP guidelines, BA1 applies to allele frequency >0.001 (0.1%) in gnomAD. The variant frequency is 0%. Therefore, BA1 is not applied.
BS1 (Not Applied)
According to VCEP guidelines, BS1 applies to allele frequency >0.0001. The variant frequency is 0%. Therefore, BS1 is not applied.
BS2 (Not Applied)
According to VCEP guidelines, BS2 applies when variant is observed in healthy individuals absent FA phenotype. No such data exist. Therefore, BS2 is not applied.
BS3 (Not Applied)
According to VCEP guidelines, BS3 requires well-established functional studies showing no damaging effect. Available assays show a damaging effect; no validated benign functional data. Therefore, BS3 is not applied.
BS4 (Not Applied)
According to VCEP guidelines, BS4 applies for lack of segregation. No segregation data are available. Therefore, BS4 is not applied.
BP1 (Not Applied)
According to VCEP guidelines, BP1 applies for missense outside functional domains with no splicing impact. C44Y is inside the RING domain. Therefore, BP1 is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines, BP2 applies for co-occurrence with a pathogenic variant without phenotype. No such data exist. Therefore, BP2 is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines, BP3 applies to in-frame repeats. Not applicable to this missense variant. Therefore, BP3 is not applied.
BP4 (Supporting)
According to VCEP guidelines, BP4_Supporting: ‘Missense inside a clinically important domain, no predicted impact (BayesDel no-AF ≤0.15 AND SpliceAI ≤0.1)’. Computational evidence shows CADD 5.51, SpliceAI 0.01, and no predicted protein impact. Therefore, BP4 is applied at Supporting strength.
BP5 (Not Applied)
According to VCEP guidelines, BP5 applies to multifactorial clinical data against pathogenicity. No co-occurrence or clinical likelihood data exist. Therefore, BP5 is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines, BP6 applies to reputable source benign assertions without evidence. No such assertions exist for this variant. Therefore, BP6 is not applied.
BP7 (Not Applied)
According to VCEP guidelines, BP7 applies to silent or intronic variants with no impact. This is a missense variant. Therefore, BP7 is not applied.