Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Open""
COSMIC Somatic Evidence
Open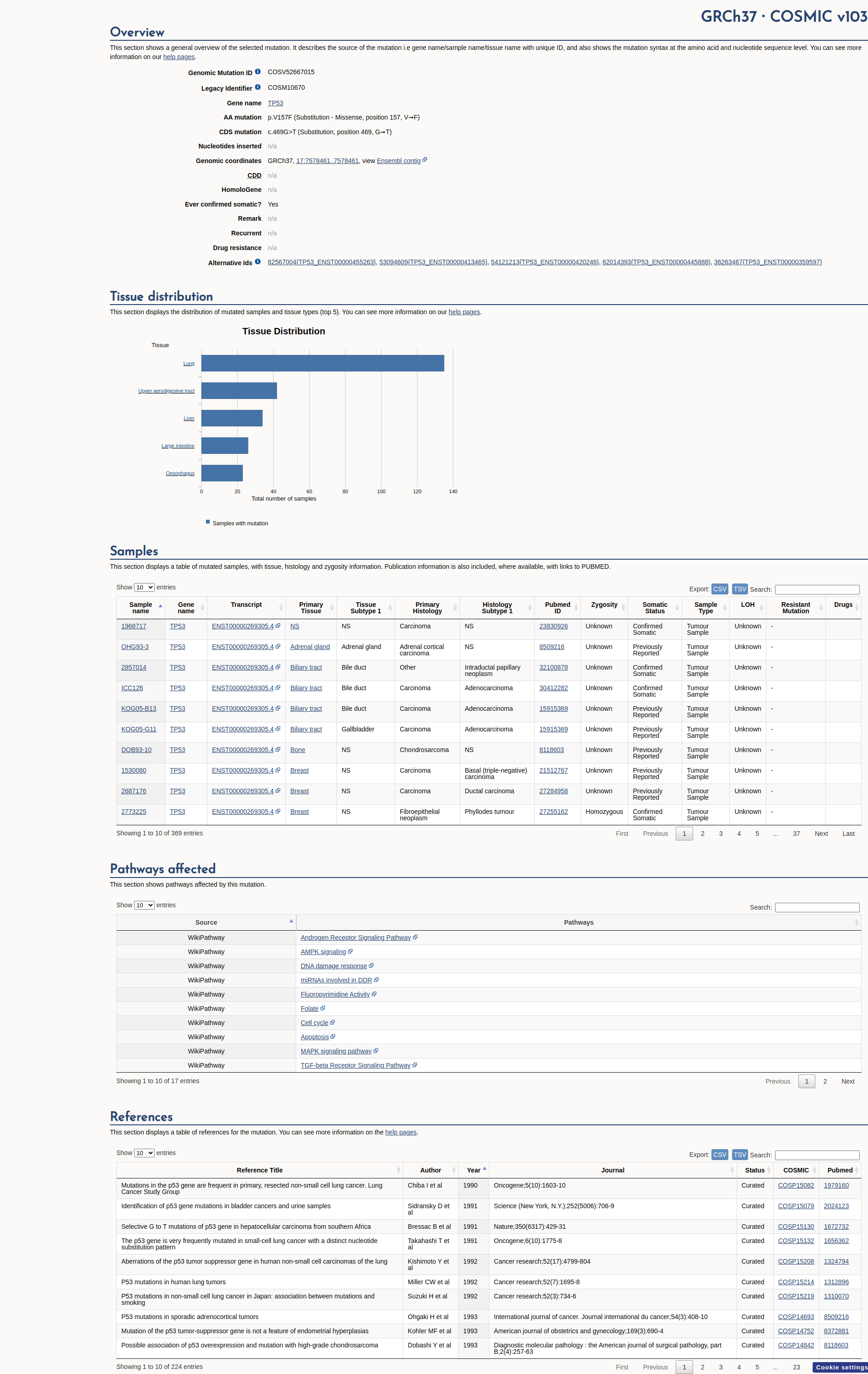
Functional Impact & Domains
Functional Domain
The TP53 V157F variant has been functionally characterized and is likely damaging. It is located in the DNA-binding domain of the TP53 protein. In vitro and structural studies have shown that this mutation leads to a rearranged DNA-binding surface and reduced DNA-binding activity compared to the wildtype. It also results in decreased expression of pro-apoptotic proteins, a reduced apoptotic response, and impaired transcriptional activity. Additionally, the variant confers a gain of function, causing aberrant gene regulation, including suppression of uPA in cell culture.
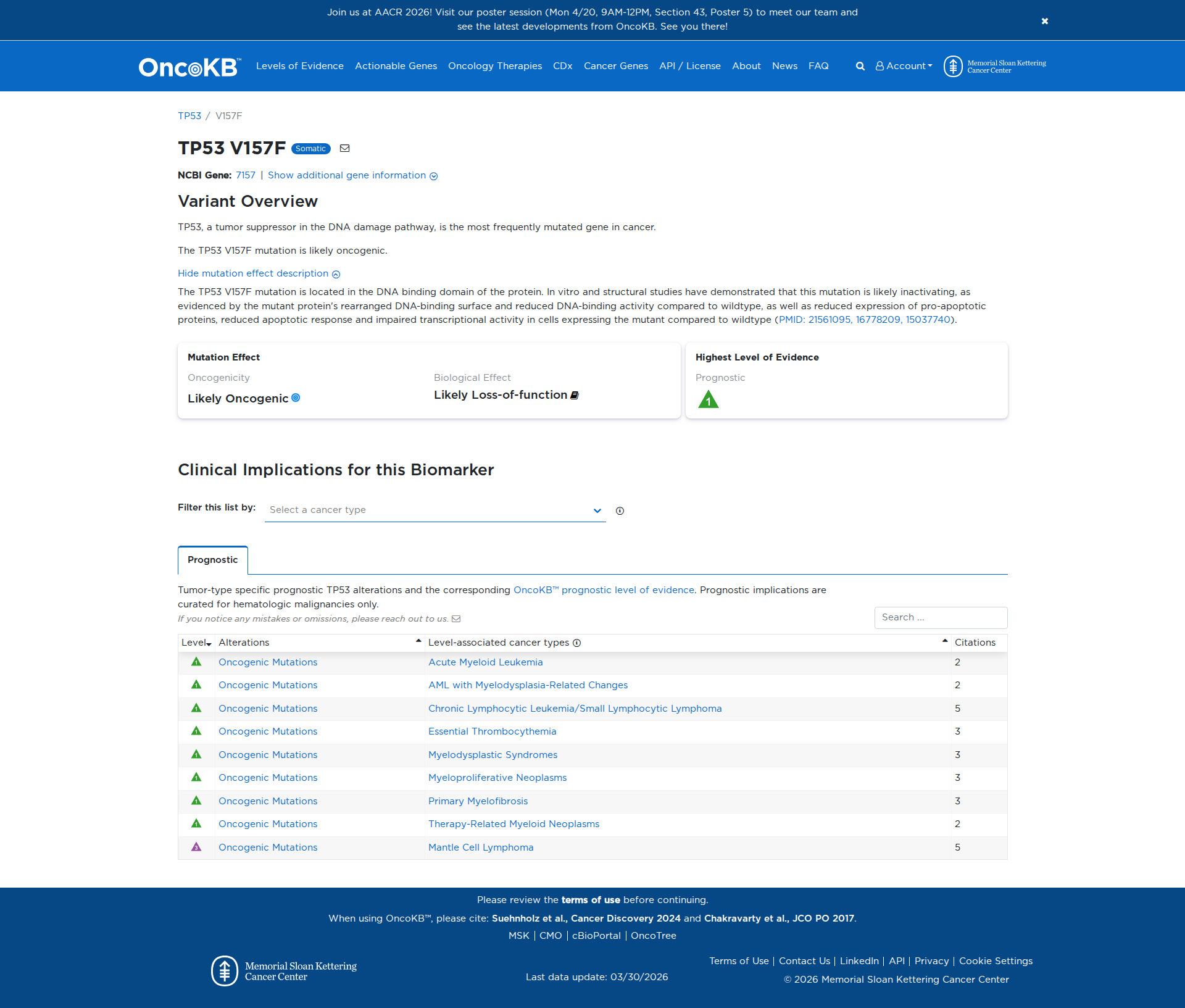
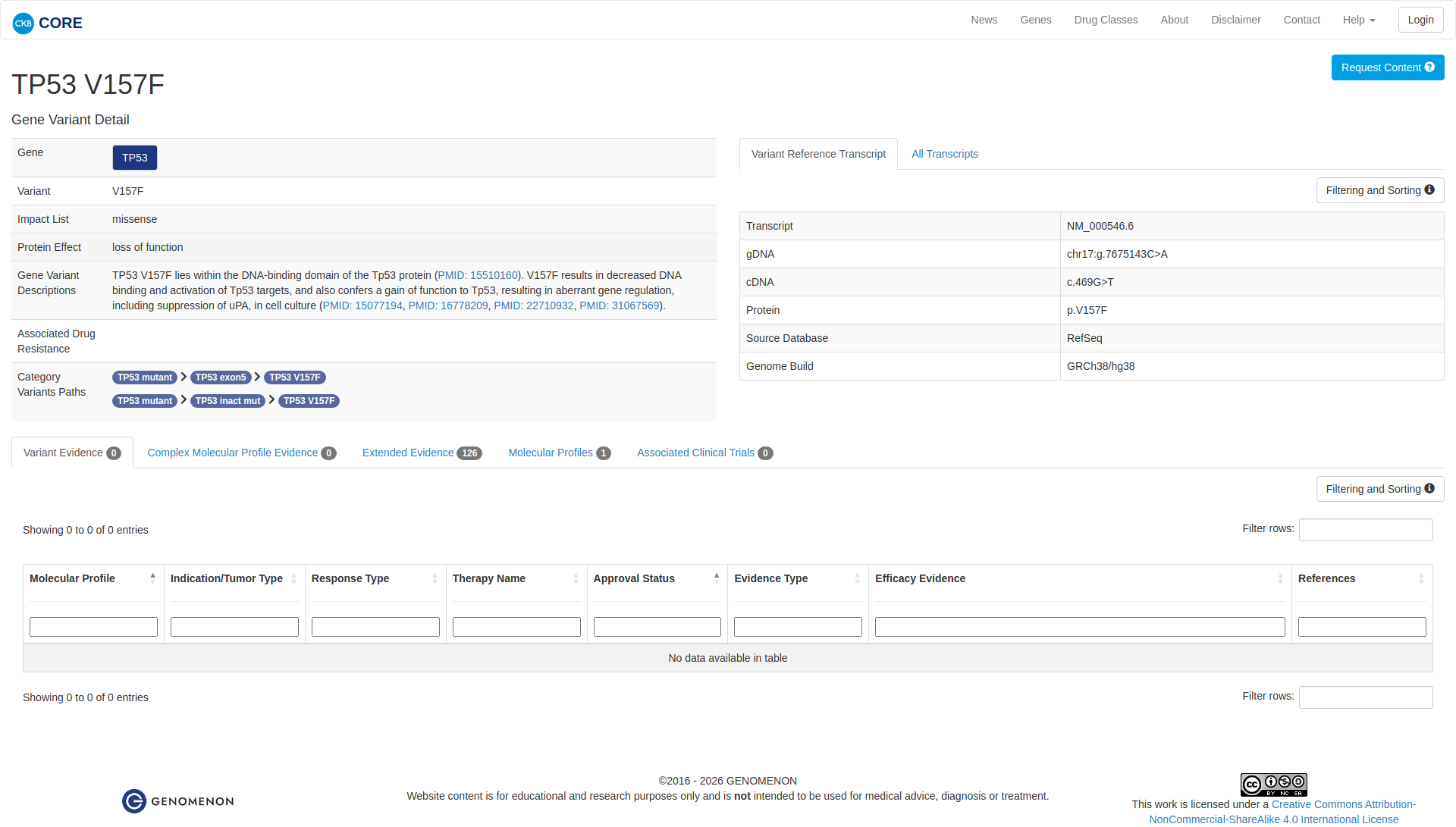
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -143 bp |
| Donor Loss (DL) | 0.0 | -44 bp |
| Acceptor Gain (AG) | 0.0 | -189 bp |
| Donor Gain (DG) | 0.0 | 35 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines (PVS1 Decision Tree for TP53): “PVS1 applies to null variants (nonsense, frameshift, canonical ±1,2 splice) predicted to undergo NMD.” The variant is a missense change (p.V157F), not a null variant. Therefore, PVS1 is not applied.
PS1 (Not Applied)
According to VCEP guidelines (PS1—Disease-specific Strength): “Strong if same amino acid change as a previously established pathogenic variant.” No other TP53 variant yielding V157F has been classified pathogenic by TP53 VCEP. Therefore, PS1 is not applied.
PS2 (Not Applied)
According to VCEP guidelines (PS2—Disease-specific Strength): “Very Strong: ≥8 de novo points; Strong: 4–7 points; Moderate: 2–3 points; Supporting: 1 point for de novo (with paternity/maternity confirmed).” No de novo data are available. Therefore, PS2 is not applied.
PS3 (Not Applied)
According to VCEP guidelines (PS3—Functional Assays): “PS3_Strong: Non-functional on Kato AND LOF on another assay; PS3_Moderate/Supp: specific combinations of Kato/Giacomelli/Kotler data.” The available functional studies are general in vitro/structural data without specific Kato, Giacomelli or Kotler assay results. Therefore, PS3 is not applied.
PS4 (Not Applied)
According to VCEP guidelines (PS4—Disease-specific Strength): “Very Strong: ≥8 proband points; Strong: 4–7.5; Moderate: 2–3.5; Supporting: 1–1.5.” No case or proband data are provided. Therefore, PS4 is not applied.
PM1 (Not Applied)
According to VCEP guidelines (PM1—Disease-specific Strength): “Moderate for missense in TP53 hotspot codons 175, 245, 248, 249, 273, 282.” Codon 157 is outside these hotspots. Therefore, PM1 is not applied.
PM2 (Supporting)
According to VCEP guidelines (PM2_Supporting): “Apply at supporting level if allele frequency <0.00003 in gnomAD.” The variant is absent from gnomAD (frequency 0%). Therefore, PM2 is applied at Supporting strength.
PM3 (Not Applied)
According to standard ACMG guidelines (PM3 applies to recessive conditions when in trans with a pathogenic variant). TP53‐associated Li-Fraumeni is autosomal dominant. Therefore, PM3 is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines (PM4 for protein length changes such as in-frame indels). This is a missense variant. Therefore, PM4 is not applied.
PM5 (Not Applied)
According to VCEP guidelines (PM5—Disease-specific Strength): “Moderate if one other pathogenic missense at same residue; Strong if two or more.” No other TP53 pathogenic variant at residue 157 is reported. Therefore, PM5 is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines (PM6 for presumed de novo without confirmation). No de novo data are available. Therefore, PM6 is not applied.
PP1 (Not Applied)
According to VCEP guidelines (PP1—Cosegregation): “Supporting: 3–4 meioses; Moderate: 5–6; Strong: ≥7.” No family segregation data are provided. Therefore, PP1 is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines (PP2 for missense in genes with low benign variation and where missense is disease mechanism). TP53 is intolerant to missense but this rule is not specified in VCEP TP53. Therefore, PP2 is not applied.
PP3 (Not Applied)
According to VCEP guidelines (PP3—Disease-specific Strength): “Moderate if BayesDel ≥0.16 and no splice impact; Supporting if BayesDel ≥0.16 (C25–C55) and SpliceAI ≥0.2).” BayesDel scores are not provided and predictions are mixed. Therefore, PP3 is not applied.
PP4 (Not Applied)
According to VCEP guidelines (PP4—Tumor VAF observations): “Supporting: VAF 5–35%; Moderate: ≥2 observations VAF 5–25%.” No tumor VAF data are available. Therefore, PP4 is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines (PP5 for reputable source classification). The variant is not in ClinVar. Therefore, PP5 is not applied.
BA1 (Not Applied)
According to VCEP guidelines (BA1: filtering allele frequency ≥0.001). The variant frequency is 0%. Therefore, BA1 is not applied.
BS1 (Not Applied)
According to VCEP guidelines (BS1: filtering allele frequency ≥0.0003 but <0.001). The variant is absent. Therefore, BS1 is not applied.
BS2 (Not Applied)
According to VCEP guidelines (BS2—unaffected elderly females). No such data are available. Therefore, BS2 is not applied.
BS3 (Not Applied)
According to VCEP guidelines (BS3—functional assays supporting benign): requires preserved function on Kato AND other assays. The functional data show impaired canonical TP53 activity. Therefore, BS3 is not applied.
BS4 (Not Applied)
According to VCEP guidelines (BS4—lack of segregation). No segregation data are provided. Therefore, BS4 is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines (BP1 for missense in gene where only truncating variants cause disease). TP53 disease mechanism is often missense. Therefore, BP1 is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines (BP2 for observed in trans with pathogenic variant in recessive disease). Not applicable to dominant TP53. Therefore, BP2 is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines (BP3 for in-frame indels in repetitive region). This is a missense variant. Therefore, BP3 is not applied.
BP4 (Not Applied)
According to VCEP guidelines (BP4—Disease-specific Strength): “Moderate if BayesDel ≤ -0.008 and SpliceAI <0.2.” BayesDel score is not provided. Although SpliceAI predicts no splice impact, BayesDel is unknown. Therefore, BP4 is not applied.
BP5 (Not Applied)
According to standard ACMG guidelines (BP5 for variant found in cis with pathogenic variant). No such data. Therefore, BP5 is not applied.
BP6 (Not Applied)
According to standard ACMG guidelines (BP6 for reputable source reporting benign). Not present in ClinVar. Therefore, BP6 is not applied.
BP7 (Not Applied)
According to VCEP guidelines (BP7 for synonymous/intronic variants). This is missense. Therefore, BP7 is not applied.