Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Open"This variant has been reported in ClinVar as Uncertain significance (1 clinical laboratories) and as Benign (25 clinical laboratories) and as Benign by ClinGen TP53 Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open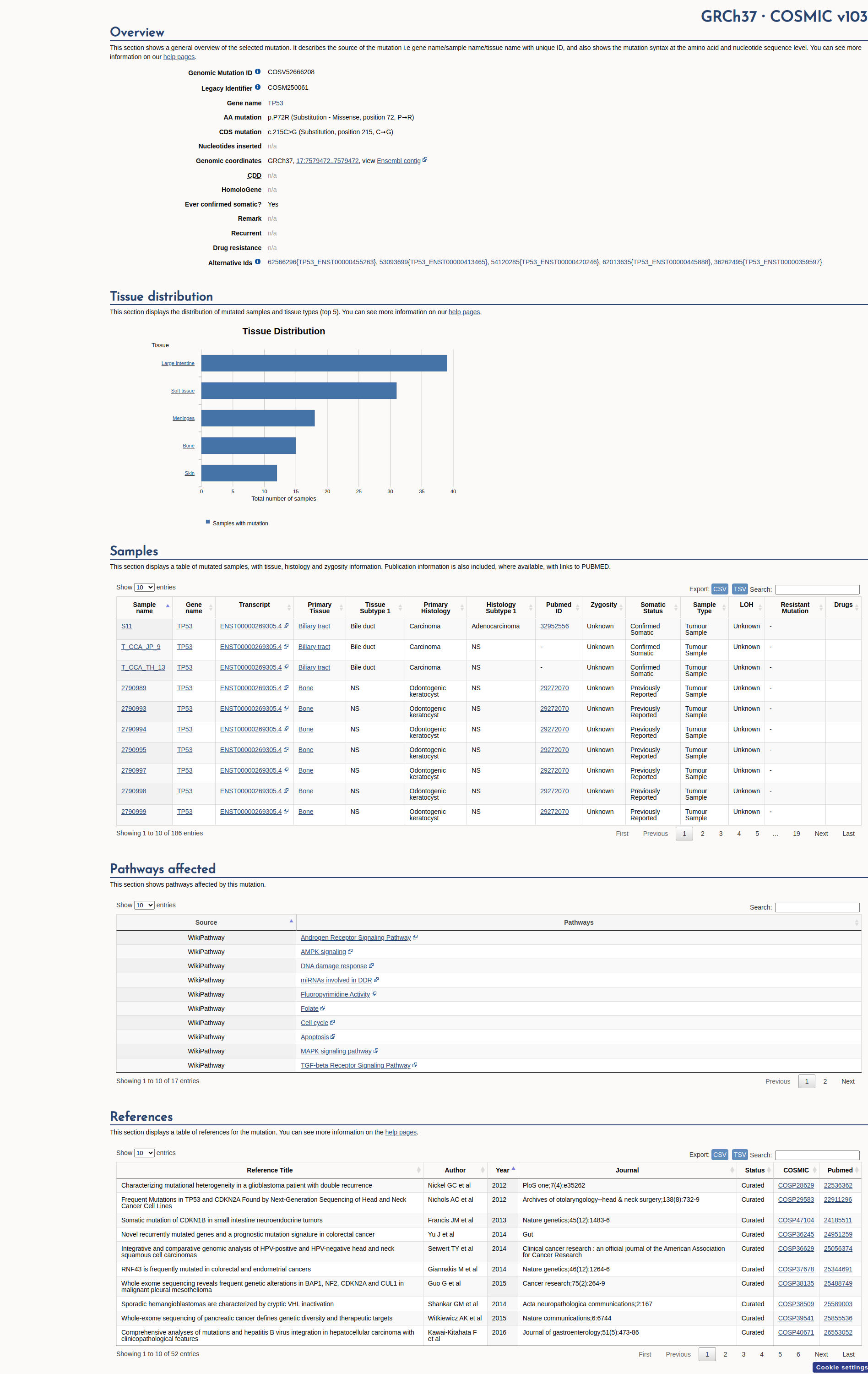
Functional Impact & Domains
Functional Domain
The TP53 P72R variant has been functionally characterized in several studies. It is a common polymorphism that decreases Pgc-1a binding and enhances mitochondrial function, migration, invasion, and tumor cell metastasis in cell culture and mouse models. However, it also transactivates downstream target genes and inhibits cell growth similarly to wild-type TP53 in culture. The overall effect of the P72R variant on TP53 protein function remains unclear.
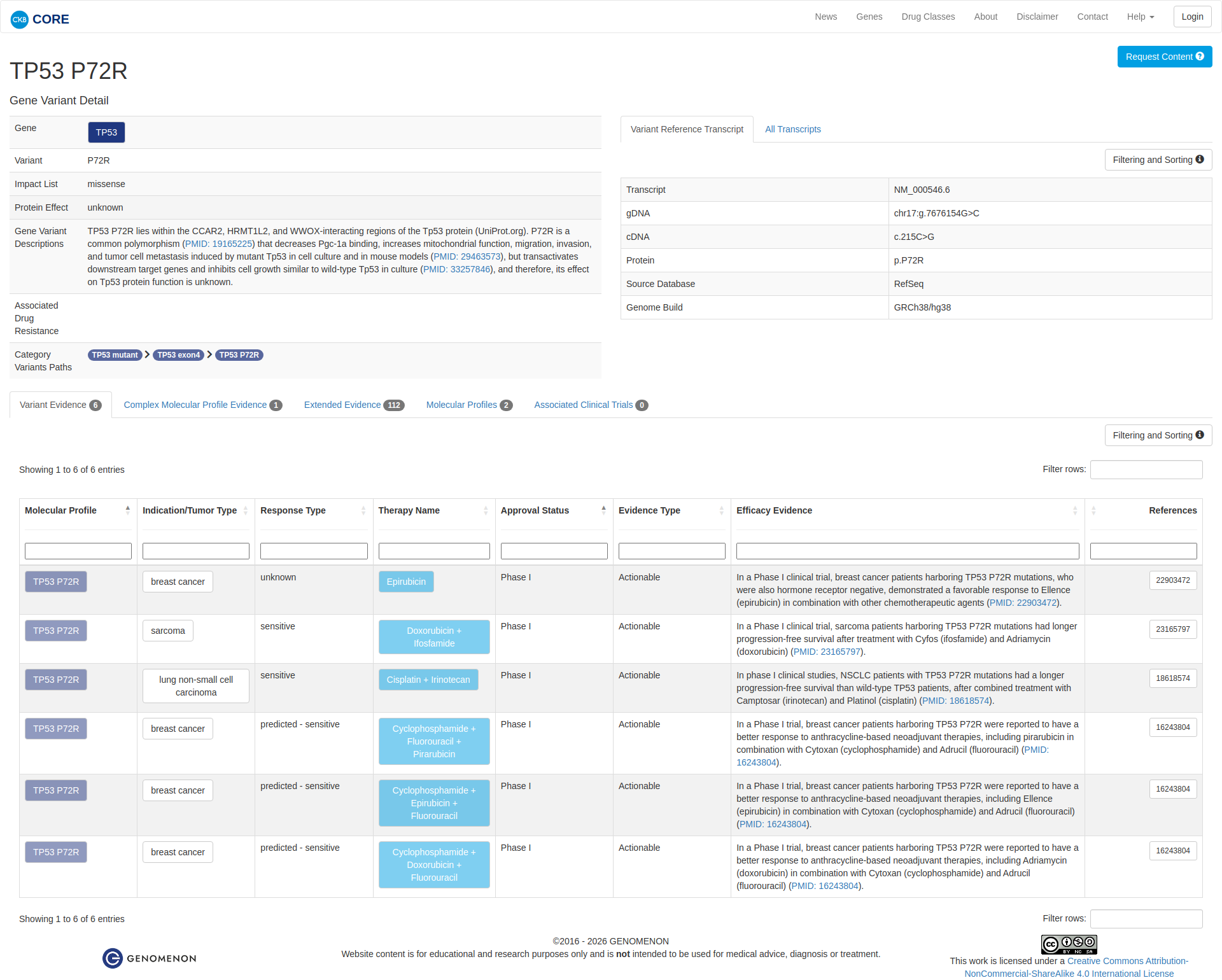
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -86 bp |
| Donor Loss (DL) | 0.05 | 40 bp |
| Acceptor Gain (AG) | 0.0 | 99 bp |
| Donor Gain (DG) | 0.0 | -160 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Not Applied)
According to VCEP guidelines: "PVS1 applies to null variants predicted to result in nonsense-mediated decay (NMD)". The evidence shows: this is a missense variant (P72R) with no introduction of a premature stop codon or disruption of canonical splice sites. Therefore, this criterion is not applied because the variant type does not meet the null variant requirement.
PS1 (Not Applied)
According to VCEP guidelines: "PS1 can be applied to variants asserted as Pathogenic following the TP53 VCEP’s specifications". The evidence shows: there is no previously established pathogenic variant resulting in the same amino acid change (Pro72Arg). Therefore, this criterion is not applied.
PS2 (Not Applied)
According to VCEP guidelines: "PS2 Very Strong strength: ≥8 points for confirmed de novo (both maternity and paternity)". The evidence shows: no de novo occurrence has been documented. Therefore, this criterion is not applied.
PS3 (Not Applied)
According to VCEP guidelines: "PS3 Strong: Non-functional on Kato et al. data AND loss of function on another assay". The evidence shows: functional studies of P72R are conflicting, with retained transactivation and enhanced mitochondrial/invasive effects. Therefore, this criterion is not applied due to lack of consistent loss-of-function evidence.
PS4 (Not Applied)
According to VCEP guidelines: "PS4 Very Strong strength: ≥8 points from proband observations". The evidence shows: no case–control or proband data have been provided. Therefore, this criterion is not applied.
PM1 (Not Applied)
According to VCEP guidelines: "PM1 Moderate strength: missense variants within TP53 codons 175, 245, 248, 249, 273, 282". The evidence shows: this variant affects codon 72, which is not a recognized hotspot. Therefore, this criterion is not applied.
PM2 (Not Applied)
According to VCEP guidelines: "PM2 Supporting strength: allele frequency <0.00003 in gnomAD". The evidence shows: the variant MAF is 66.3% overall (73.7% in European (non-Finnish)), far exceeding the threshold. Therefore, this criterion is not applied.
PM3 (Not Applied)
According to standard ACMG guidelines: "PM3 applies to variants observed in trans with a pathogenic variant for recessive disorders". The evidence shows: TP53-related disorders are autosomal dominant and no trans‐allele data are provided. Therefore, this criterion is not applied.
PM4 (Not Applied)
According to standard ACMG guidelines: "PM4 applies to protein length changes such as in-frame indels or stop-loss variants". The evidence shows: this is a missense variant with no change in protein length. Therefore, this criterion is not applied.
PM5 (Not Applied)
According to VCEP guidelines: "PM5 Moderate strength: missense variant at a residue where ≥1 different pathogenic missense has been seen". The evidence shows: no other pathogenic missense variants at codon 72 have been established. Therefore, this criterion is not applied.
PM6 (Not Applied)
According to standard ACMG guidelines: "PM6 applies to presumed de novo without confirmation of paternity and maternity". The evidence shows: no such data are available. Therefore, this criterion is not applied.
PP1 (Not Applied)
According to VCEP guidelines: "PP1 Supporting strength: cosegregation observed in 3–4 meioses". The evidence shows: no familial segregation data have been reported. Therefore, this criterion is not applied.
PP2 (Not Applied)
According to standard ACMG guidelines: "PP2 applies if a gene has a low rate of benign missense variation and missense variants are a common mechanism of disease". The evidence shows: TP53 tolerates missense variation and this specific change is a common polymorphism. Therefore, this criterion is not applied.
PP3 (Not Applied)
According to VCEP guidelines: "PP3 Supporting strength: BayesDel ≥0.16 and SpliceAI ≥0.2 for splicing". The evidence shows: mixed in silico predictions and SpliceAI predicts no significant splice impact (score 0.05). Therefore, this criterion is not applied.
PP4 (Not Applied)
According to standard ACMG guidelines: "PP4 applies when a specific phenotype or family history is strongly predictive of a single gene disorder". The evidence shows: no phenotype or family history data are provided. Therefore, this criterion is not applied.
PP5 (Not Applied)
According to standard ACMG guidelines: "PP5 applies when a reputable source reports the variant as pathogenic without evidence". The evidence shows: ClinVar entries are VUS or benign, not pathogenic. Therefore, this criterion is not applied.
BA1 (Stand Alone)
According to VCEP guidelines: "BA1 Stand Alone: Filtering allele frequency ≥0.001 in gnomAD continental subpopulations". The evidence shows: MAF = 66.3% overall and 73.7% in European (non-Finnish), far above 0.1%. Therefore, this criterion is applied at Stand Alone strength because the variant is too common to be disease-causing.
BS1 (Not Applied)
According to VCEP guidelines: "BS1 Strong: Filtering allele frequency ≥0.0003 but <0.001". The evidence shows: allele frequency is 66.3%, which exceeds the upper limit of 0.001 for BS1. Therefore, this criterion is not applied.
BS2 (Not Applied)
According to VCEP guidelines: "BS2 Strong: ≥8 unrelated unaffected individuals ≥60 years old (females)". The evidence shows: no such cohort data are available. Therefore, this criterion is not applied.
BS3 (Not Applied)
According to VCEP guidelines: "BS3 Strong: functional on Kato et al. data AND no loss of function on another assay". The evidence shows: functional data are conflicting with some assays showing gain of function. Therefore, this criterion is not applied.
BS4 (Not Applied)
According to VCEP guidelines: "BS4 Strong: lack of segregation in affected family members". The evidence shows: no segregation data are provided. Therefore, this criterion is not applied.
BP1 (Not Applied)
According to standard ACMG guidelines: "BP1 applies to missense variants in a gene where only truncating variants cause disease". The evidence shows: TP53 disease is caused by missense variants. Therefore, this criterion is not applied.
BP2 (Not Applied)
According to standard ACMG guidelines: "BP2 applies when a variant is observed in trans with a pathogenic variant for a dominant disorder or in cis phase". The evidence shows: no such data. Therefore, this criterion is not applied.
BP3 (Not Applied)
According to standard ACMG guidelines: "BP3 applies to in-frame deletions/insertions in repetitive regions". The evidence shows: this is a single amino acid substitution. Therefore, this criterion is not applied.
BP4 (Supporting)
According to standard ACMG guidelines: "BP4: multiple lines of computational evidence suggest no impact on gene or gene product". The evidence shows: in silico tools (CADD, PolyPhen, MetaSVM, MetaLR, PrimateAI) predominantly predict benign effect and SpliceAI predicts no splicing impact. Therefore, this criterion is applied at Supporting strength.
BP5 (Not Applied)
According to standard ACMG guidelines: "BP5 applies when a variant is found in a case with an alternate molecular basis for disease". The evidence shows: no evidence of an alternate cause. Therefore, this criterion is not applied.
BP6 (Supporting)
According to standard ACMG guidelines: "BP6: reputable source reports variant as benign without available evidence". The evidence shows: ClinVar entries include 25 benign classifications and expert panel curation considers it benign. Therefore, this criterion is applied at Supporting strength.
BP7 (Not Applied)
According to VCEP guidelines: "BP7 applies to synonymous or intronic variants outside core splice motifs". The evidence shows: this is a missense variant. Therefore, this criterion is not applied.