Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000314.7 | RefSeq Select | 8514 nt | 845–2056 |
| NM_000314.5 | Alternative | 8719 nt | 1032–2243 |
| NM_000314.4 | Alternative | 5572 nt | 1032–2243 |
| NM_000314.3 | Alternative | 3416 nt | 1032–2243 |
| NM_000314.6 | Alternative | 8718 nt | 1032–2243 |
| NM_000314.8 | MANE Select | 8515 nt | 846–2057 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenFor these reasons, this variant has been classified as Pathogenic. While this particular variant has not been reported in the literature, loss-of-function variants in PTEN are known to be pathogenic (PMID: 9467011, 21194675). This sequence change deletes 1 nucleotide from exon 6 of the PTEN mRNA (c.525delG), causing a frameshift at codon 176. This creates a premature translational stop signal (p.Tyr176Ilefs*7) and is expected to result in an absent or disrupted protein product.
"This variant has been reported in ClinVar as Pathogenic (1 clinical laboratories)."
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: PTENY176Ifs*7PTENY176Ifs*7SomaticNCBI Gene:5728|Show additional gene information Variant OverviewPTEN, a lipid and protein phosphatase, is one of the most frequently mutated genes in cancer.The PTEN Y176Ifs*7 is a truncating mutation in a tumor suppressor gene, and therefore is likely oncogenic.Hide mutation effect description The mutation effect description for truncating mutations in PTEN is: PTEN truncating mutations can produce several forms of C-terminally truncated PTEN proteins. Truncating mutations closer to the N-terminus result in loss of PTEN phosphatase function and an inability to negatively regulate PI3K/AKT pathway activity (PMID: 11237521). Expression of a PTEN truncation mutation in mouse embryonic fibroblasts demonstrated that these mutations are oncogenic and increase genome fragility due to the inability of PTEN to associate with chromosomal centromeres compared to the full-length protein (PMID: 17218262). JAX-CKB: No results found
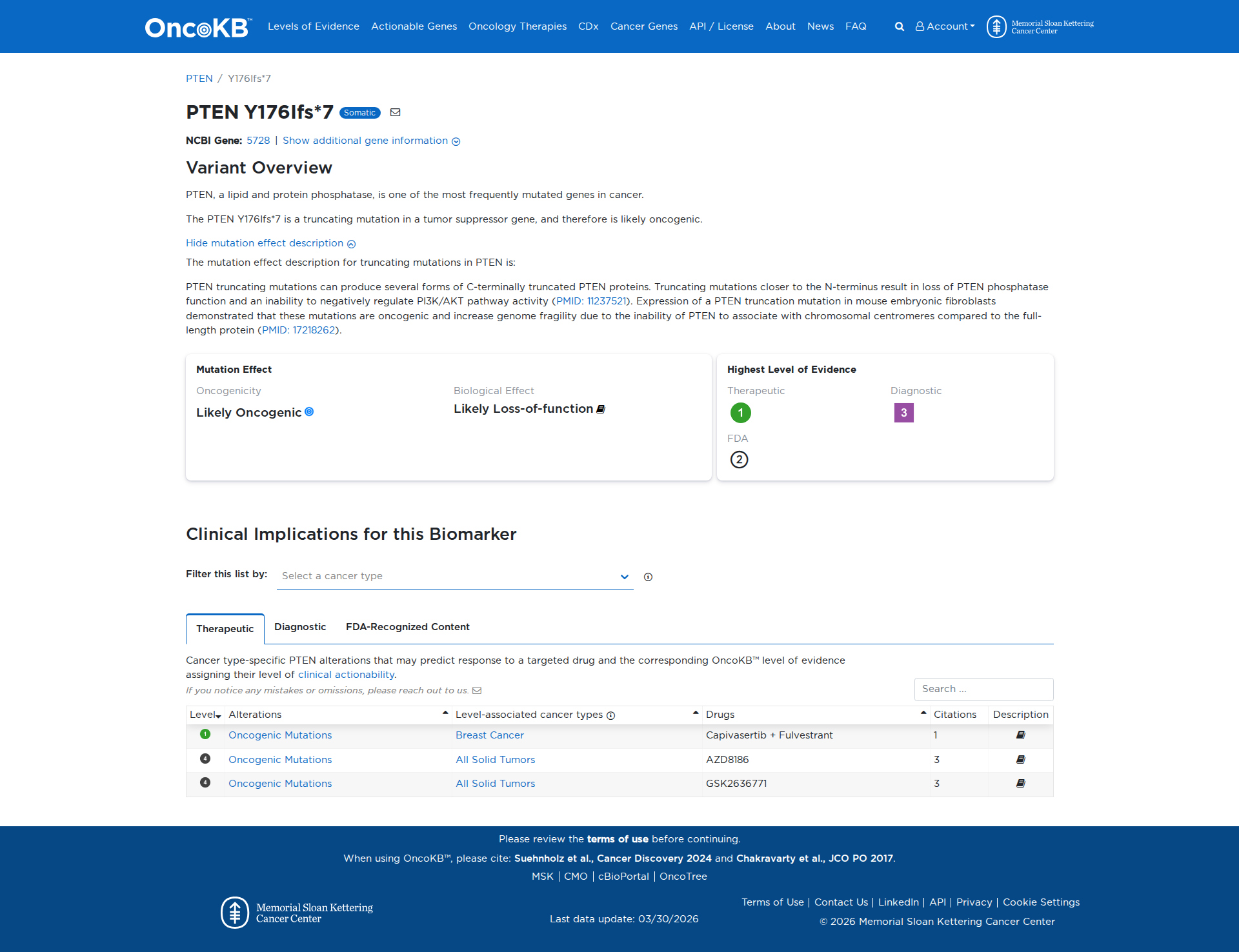
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.02 | -31 bp |
| Donor Loss (DL) | 0.01 | 110 bp |
| Acceptor Gain (AG) | 0.0 | 13 bp |
| Donor Gain (DG) | 0.0 | -31 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)