Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_002755.4 | MANE Select | 2547 nt | 437–1618 |
| NM_002755.3 | RefSeq Select | 2603 nt | 476–1657 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenVariant summary: MAP2K1 c.158T>C (p.Phe53Ser) results in a non-conservative amino acid change in the encoded protein sequence. Four of five in-silico tools predict a damaging effect of the variant on protein function. The variant was absent in 251374 control chromosomes. c.158T>C has been reported in the literature in individuals affected with Cardiofaciocutaneous Syndrome (example, Rodrigues-Viciana_2006, Yoon_2007, Siegel_2011, Bessis_2019). These data indicate that the variant is likely to be associated with disease. At least one publication reports experimental evidence evaluating an impact on protein function. The most pronounced variant effect results in significant stimulation of ERK phosphorylation relative to the wild-type control (example, Rodriguez-Viciana_2008). One clinical diagnostic laboratory and an expert panel (ClinGen RASopathy Variant Curation Expert Panel) have submitted clinical-significance assessments for this variant to ClinVar after 2014 without evidence for independent evaluation. Both submitters classified the variant as pathogenic. Based on the evidence outlined above, the variant was classified as pathogenic.
"This variant has been reported in ClinVar as Pathogenic (2 clinical laboratories) and as Likely Pathogenic by ClinGen RASopathy Variant Curation Expert Panel expert panel."
COSMIC Somatic Evidence
Open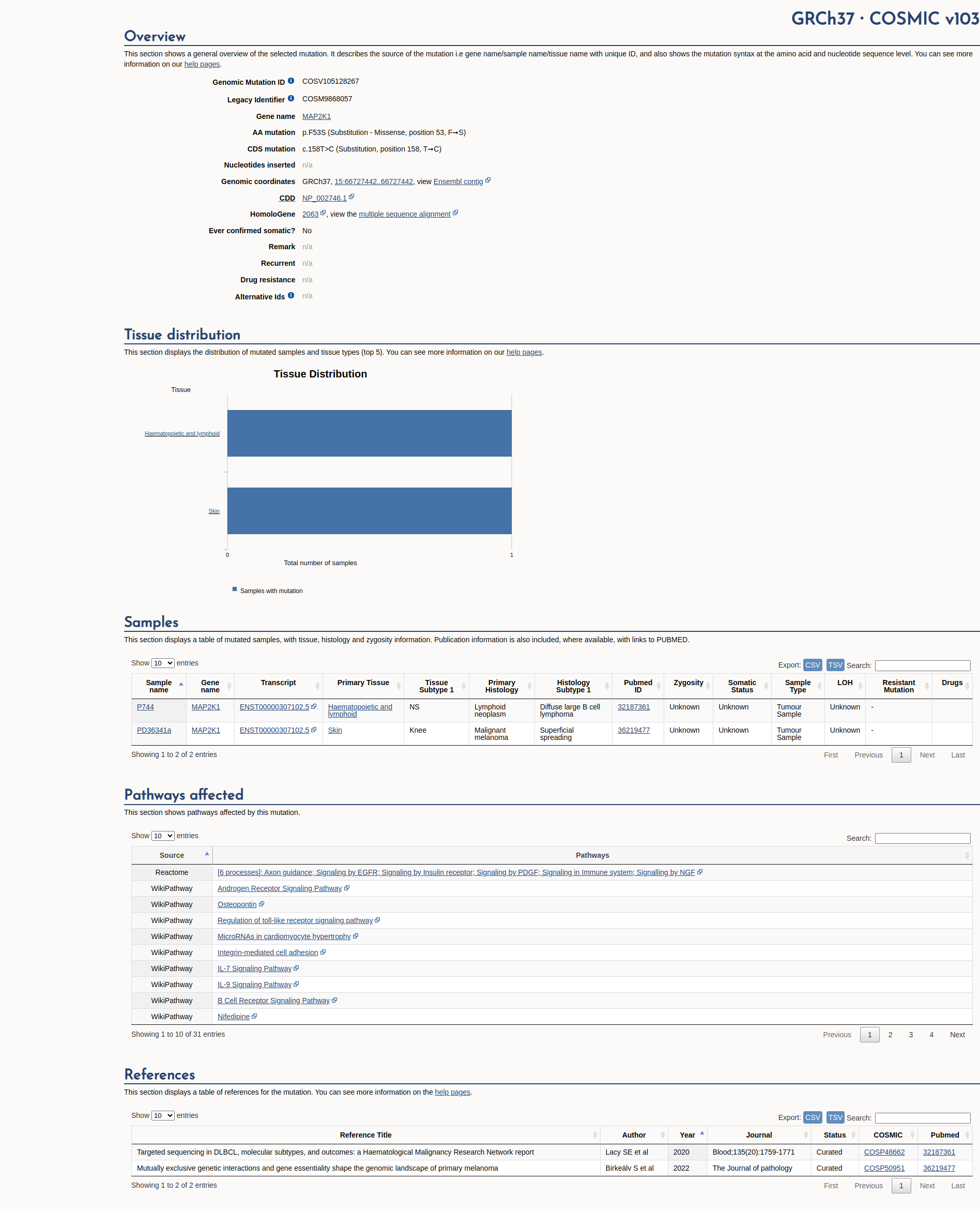
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: MAP2K1F53SMAP2K1F53SSomaticNCBI Gene:5604|Show additional gene information Variant OverviewMAP2K1, an intracellular kinase, is mutated at low frequencies in various cancer types including melanoma, colorectal and lung cancers.The MAP2K1 F53S mutation is likely oncogenic.Hide mutation effect description The MEK1 F53S mutation is located in the negative regulatory region of the protein and occurs as a result of a mutation in exon 2 of the MAP2K1 gene. Expression of the MEK1 F53S mutation in 293T and HEK293 cells demonstrated that it is activating as measured by increased downstream pathway activation associated with mutant MEK1 compared to wildtype MEK1 that is dependent on upstream activation by BRAF (PMID: 16439621, 17981815). JAX-CKB: MAP2K1 F53S lies within the negative regulatory region of the Map2k1 protein (PMID: 24241536). F53S demonstrates phosphorylation profile similar to wild-type Map2k1 (PMID: 29753091), and results in increased Erk phosphorylation in cell culture (PMID: 16439621, PMID: 12370306).


Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 11 bp |
| Donor Loss (DL) | 0.0 | 155 bp |
| Acceptor Gain (AG) | 0.0 | -141 bp |
| Donor Gain (DG) | 0.0 | 19 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)