Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_002880.4 | MANE Select | 3191 nt | 332–2278 |
| NM_002880.3 | Alternative | 3291 nt | 416–2362 |
| NM_002880.2 | Alternative | 3245 nt | 394–2340 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis sequence change replaces arginine, which is basic and polar, with leucine, which is neutral and non-polar, at codon 398 of the RAF1 protein (p.Arg398Leu). This variant also falls at the last nucleotide of exon 11, which is part of the consensus splice site for this exon. In summary, the currently available evidence indicates that the variant is pathogenic, but additional data are needed to prove that conclusively. Therefore, this variant has been classified as Likely Pathogenic. Variants that disrupt the consensus splice site are a relatively common cause of aberrant splicing (PMID: 17576681, 9536098). An algorithm developed to predict the effect of missense changes on protein structure and function (PolyPhen-2) suggests that this variant is likely to be disruptive. ClinVar contains an entry for this variant (Variation ID: 40614). This missense change has been observed in individuals with clinical features of RAF1-related RASopathy and/or clinical features of RASopathy spectrum disorders (Invitae). This variant is not present in population databases (gnomAD no frequency).
"This variant has been reported in ClinVar as Uncertain significance (3 clinical laboratories) and as Likely pathogenic (2 clinical laboratories) and as Uncertain Significance by ClinGen RASopathy Variant Curation Expert Panel expert panel."
COSMIC Somatic Evidence
Open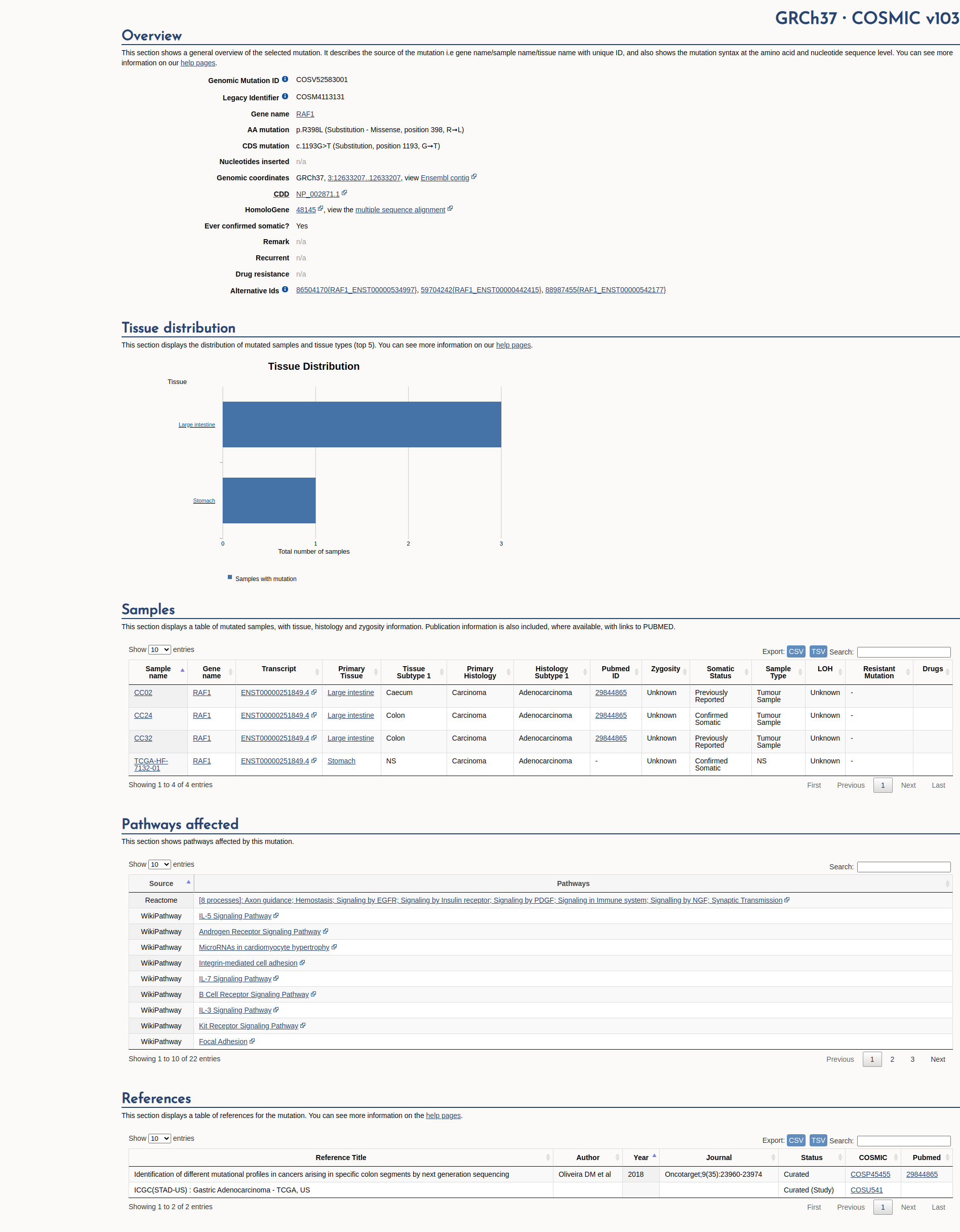
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: RAF1R398LRAF1R398LSomaticNCBI Gene:5894|Show additional gene information Variant OverviewRAF1 (CRAF), an intracellular kinase and component of the pro-oncogenic MAP-kinase signaling pathway, is infrequently mutated in cancer. Germline mutations of RAF1 are associated with Noonan and LEOPARD syndrome.There is no available functional data about the RAF1 R398L mutation (last reviewed on 10/16/2018), and therefore its biological significance is unknown. JAX-CKB: No results found


Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.1 | 84 bp |
| Donor Loss (DL) | 0.08 | 0 bp |
| Acceptor Gain (AG) | 0.0 | 72 bp |
| Donor Gain (DG) | 0.02 | 14 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)