Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000059.4 | MANE Select | 11954 nt | 200–10456 |
| NM_000059.2 | Alternative | 11386 nt | 228–10484 |
| NM_000059.3 | RefSeq Select | 11386 nt | 228–10484 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe c.7977-1G>C intronic pathogenic mutation results from a G to C substitution one nucleotide upstream from coding exon 17 of the BRCA2 gene. A multifactorial likelihood ratio model that included co-segregation, pathology, co-occurrence, and family history data found this alteration to be pathogenic (Spurdle AB et al. Hum. Mutat. 2010 Feb;31:E1141-5; Parsons MT et al. Hum Mutat 2019 09;40(9):1557-1578). RNA studies have demonstrated that this alteration results in abnormal splicing in the set of samples tested (Ambry internal data; Fraile-Bethencourt E et al. PLoS Genet. 2017 Mar;13:e1006691). Note, this variant is also referred to as IVS17-1G>C in the literature. This nucleotide position is highly conserved in available vertebrate species. In silico splice site analysis predicts that this alteration will weaken the native splice acceptor site. Alterations that disrupt the canonical splice site are expected to cause aberrant splicing, resulting in an abnormal protein or a transcript that is subject to nonsense-mediated mRNA decay. As such, this alteration is classified as likely pathogenic.
Variant summary: BRCA2 c.7977-1G>C is located in a canonical splice-site and is predicted to affect mRNA splicing resulting in a significantly altered protein due to either exon skipping, shortening, or inclusion of intronic material. Several computational tools predict a significant impact on normal splicing: Four predict the variant abolishes a 3' acceptor site. At least one publication reported experimental evidence confirming that this variant affects mRNA splicing, by showing in a minigene assay several types of aberrant transcripts that exclude exon 18 (partly or fully), and no detectable canonical transcript (Fraile-Bethencourt_2017). The variant allele was found at a frequency of 4e-06 in 249640 control chromosomes (gnomAD). c.7977-1G>C has been reported in the literature in several individuals affected with breast, ovarian, and prostate cancer (e.g. Edwards_2003, Walsh_2011, Snape_2012, Jimenez_2013, Pritzlaff_2017, Dorling_2021). These data indicate that the variant is very likely to be associated with disease. 13 other submitters, including an expert panel (ENIGMA), have provided clinical-significance assessments for this variant in ClinVar after 2014 and all classified the variant as pathogenic. Based on the evidence outlined above, the variant was classified as pathogenic.
This variant disrupts a canonical splice-acceptor site and interferes with normal BRCA2 mRNA splicing. The frequency of this variant in the general population, 0.00004 (2/50340 chromosomes, http://gnomad.broadinstitute.org), is consistent with pathogenicity. In the published literature, the variant has been reported in individuals with breast, ovarian, peritoneal, and prostate cancer (PMIDs: 28008555 (2017), 25685387 (2015), 23569316 (2013), 22006311 (2011), 20736950 (2010), 12474142 (2003)). In addition, this variant has been reported to cause aberrant splicing and loss of exon 18 (PMIDs: 28339459 (2017), 16211554 (2005)). Based on the available information, this variant is classified as pathogenic.
This sequence change affects an acceptor splice site in intron 17 of the BRCA2 gene. RNA analysis indicates that disruption of this splice site induces altered splicing and may result in an absent or altered protein product. This variant is present in population databases (rs81002874, gnomAD 0.002%). Disruption of this splice site has been observed in individual(s) with prostate, ovarian, and breast cancer (PMID: 12474142, 16211554, 22006311, 23479189). This variant is also known as IVS17-1G>C or 8205-1G>C. ClinVar contains an entry for this variant (Variation ID: 38132). Based on a multifactorial likelihood algorithm using genetic, in silico, and/or statistical data, this variant has been determined to have a high probability of being pathogenic (PMID: 20020529). Studies have shown that disruption of this splice site results in skipping of exons 17-18 and skipping of exon 18, and produces a non-functional protein and/or introduces a premature termination codon (PMID: 16211554, 22006311; internal data). For these reasons, this variant has been classified as Pathogenic.
"This variant has been reported in ClinVar as Pathogenic (20 clinical laboratories) and as Pathogenic by ClinGen ENIGMA BRCA1 and BRCA2 Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open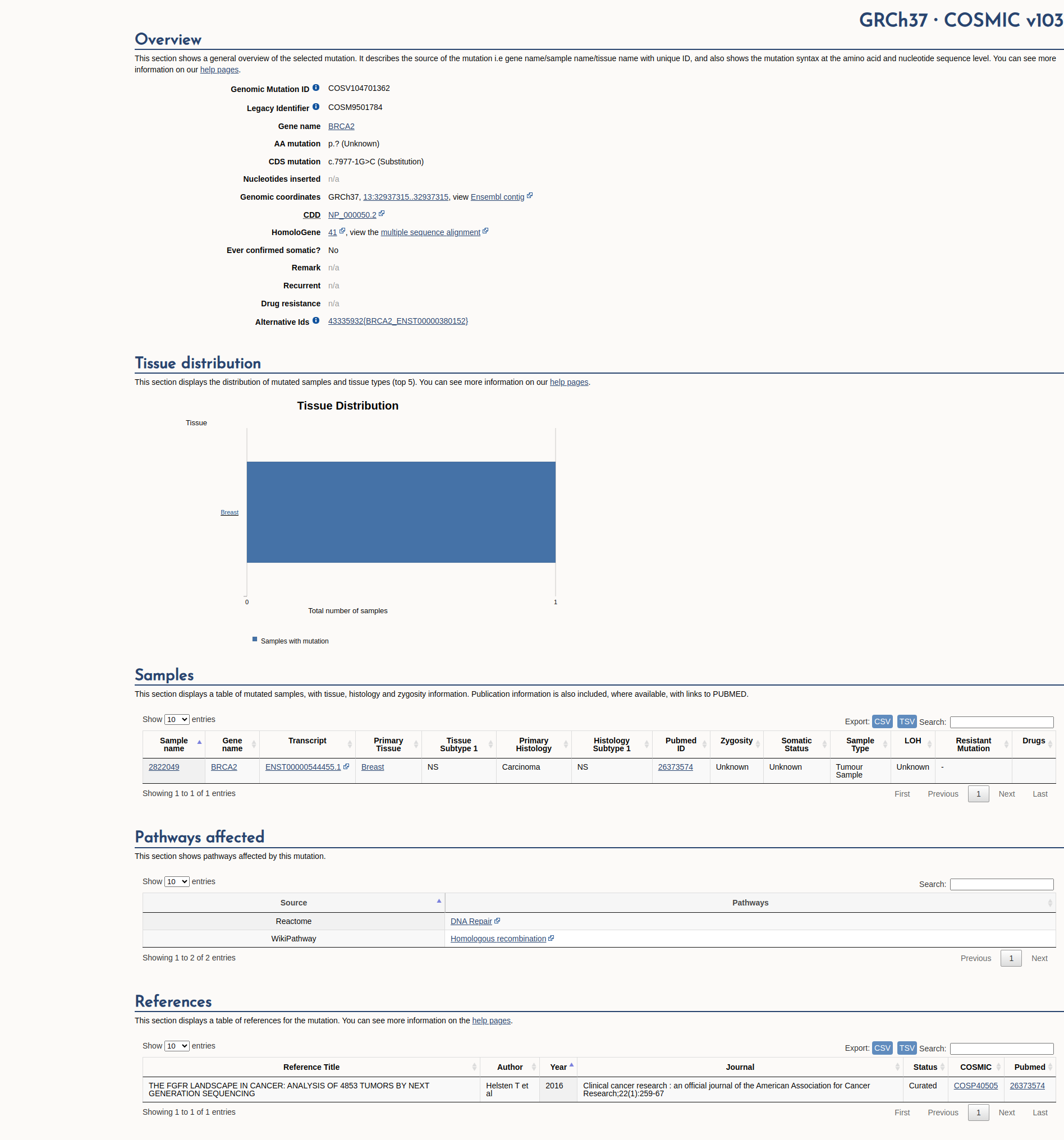
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: BRCA2SpliceBRCA2SpliceSomaticNCBI Gene:675|Show additional gene information Variant OverviewBRCA2, a tumor suppressor involved in the DNA damage response, is mutated in various cancer types.The BRCA2 splice alteration has not specifically been reviewed by the OncoKB team, and therefore its biological significance is unknown. JAX-CKB: No results found

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.95 | 1 bp |
| Donor Loss (DL) | 0.27 | 355 bp |
| Acceptor Gain (AG) | 0.11 | 22 bp |
| Donor Gain (DG) | 0.0 | 3 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)