Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_007294.4 | MANE Select | 7088 nt | 114–5705 |
| NM_007294.2 | Alternative | 7191 nt | 201–5792 |
| NM_007294.3 | RefSeq Select | 7224 nt | 233–5824 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenNot found in the total gnomAD dataset, and the data is high quality. Found in at least one patient with expected phenotype for this gene. Predicted to have a damaging effect on the protein. Located in potentially critical domain of the protein. Assessment of experimental evidence suggests this variant results in abnormal protein function.
This sequence change replaces cysteine, which is neutral and slightly polar, with arginine, which is basic and polar, at codon 1697 of the BRCA1 protein (p.Cys1697Arg). This variant is not present in population databases (gnomAD no frequency). This missense change has been observed in individual(s) with breast and/or ovarian cancer (PMID: 11157798, 11389159, 18465347, 36537080). This variant is also known as 5208T>C. ClinVar contains an entry for this variant (Variation ID: 55392). Advanced modeling performed at Invitae incorporating data from internal and/or published experimental studies (PMID: 30209399) indicates that this missense variant is expected to disrupt BRCA1 function with a positive predictive value of 95%. Experimental studies have shown that this missense change affects BRCA1 function (PMID: 11157798, 11389159, 14534301, 20516115, 28398198, 30209399, 30257991). In summary, the currently available evidence indicates that the variant is pathogenic, but additional data are needed to prove that conclusively. Therefore, this variant has been classified as Likely Pathogenic.
The c.5089T>C variant in BRCA1 is a missense variant predicted to cause substitution of Cysteine by Arginine at amino acid 1697 (p.Cys1697Arg). This variant is absent from gnomAD v2.1 (exomes only, non-cancer subset, read depth >=25) and gnomAD v3.1 (non-cancer subset, read depth >=25) (PM2_Supporting met). This BRCA1 missense variant is within a key functional domain and the computational predictor BayesDel (noAF) gives a score of 0.40, above the recommended threshold of 0.28 for prediction of impact on BRCA1 function via protein change. SpliceAI predictor score of 0.05 suggests that the variant has no impact on splicing (score threshold <0.10) (PP3 met). Reported by three calibrated studies to exhibit protein function similar to pathogenic control variants (PMIDs: 30209399, 30257991, 30765603) (PS3 met). Cosegregation analysis of family(ies) carrying this variant provided evidence towards pathogenicity, and has a Bayes Score of 5.24, within the thresholds for moderate pathogenic evidence (LR >4.3 & <=18.7) (PP1_Moderate met; internal lab contributor). Multifactorial likelihood ratio analysis using clinically calibrated data produced a combined LR for this variant of 2735.2 (based on Cosegregation LR=730.6; Pathology LR=2.41; Family History LR=1.56), above the thresholds for Very strong evidence towards pathogenicity (LR>350) (PP4_Very strong met; PMID: 31131967, 31853058). In summary, this variant meets the criteria to be classified as a Pathogenic variant for BRCA1-related cancer predisposition based on the ACMG/AMP criteria applied as specified by the ENIGMA BRCA1/2 VCEP (PM2_Supporting, PP3, PS3, PP1_Moderate, PP4_Very strong).
The p.C1697R variant (also known as c.5089T>C), located in coding exon 16 of the BRCA1 gene, results from a T to C substitution at nucleotide position 5089. The cysteine at codon 1697 is replaced by arginine, an amino acid with highly dissimilar properties. This alteration has been identified in several breast/ovarian cancer kindreds and has not been reported in control populations (Bergthorsson JT, J. Med. Genet. 2001 Jun; 38(6):361-8; Malander S, Eur. J. Cancer 2004 Feb; 40(3):422-8; Thomassen M, Acta Oncol 2008; 47(4):772-7; Biunno I et al. Fam Cancer, 2014 Sep;13:437-44; Machackova E et al. Klin Onkol, 2019;32:51-71; Shao D et al. Cancer Sci, 2020 Feb;111:647-657; Dorling et al. N Engl J Med. 2021 02;384:428-439; Kechin A et al. Breast Cancer Res Treat, 2023 Jan;197:387-395). Structural modeling and functional assays evaluating homology directed DNA repair, proteolytic degradation, protein stability, transcriptional activation, and peptide binding ability have suggested that this alteration results in reduced or loss of wild type function (Vallon-Christersson J, Hum. Mol. Genet. 2001 Feb; 10(4):353-60; Williams RS, J. Biol. Chem. 2003 Dec;278(52):53007-16; Clapperton JA et al. Nat. Struct. Mol. Biol., 2004 Jun;11:512-8; Glover JN, Fam. Cancer 2006; 5(1):89-93; Anantha RW et al. Elife, 2017 04;6; Findlay GM et al. Nature, 2018 10;562:217-222; Fernandes VC et al. J Biol Chem, 2019 04;294:5980-5992; Petitalot A et al. Mol Cancer Res, 2019 01;17:54-69). Multifactorial and computational likelihood models predict this variant to be deleterious (Karchin R, PLoS Comput. Biol. 2007 Feb; 3(2):e26; Iversen ES, Cancer Epidemiol. Biomarkers Prev. 2011 Jun; 20(6):1078-88; Lee JS et al. J Med Genet, 2018 12;55:794-802; Parsons MT et al. Hum Mutat, 2019 09;40:1557-1578). This amino acid position is highly conserved in available vertebrate species. In addition, this alteration is predicted to be deleterious by in silico analysis. This variant is considered to be rare based on population cohorts in the Genome Aggregation Database (gnomAD). Based on the majority of available evidence to date, this variant is likely to be pathogenic.
"This variant has been reported in ClinVar as Uncertain significance (2 clinical laboratories) and as Likely pathogenic (6 clinical laboratories) and as Pathogenic (5 clinical laboratories) and as Pathogenic by ClinGen ENIGMA BRCA1 and BRCA2 Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open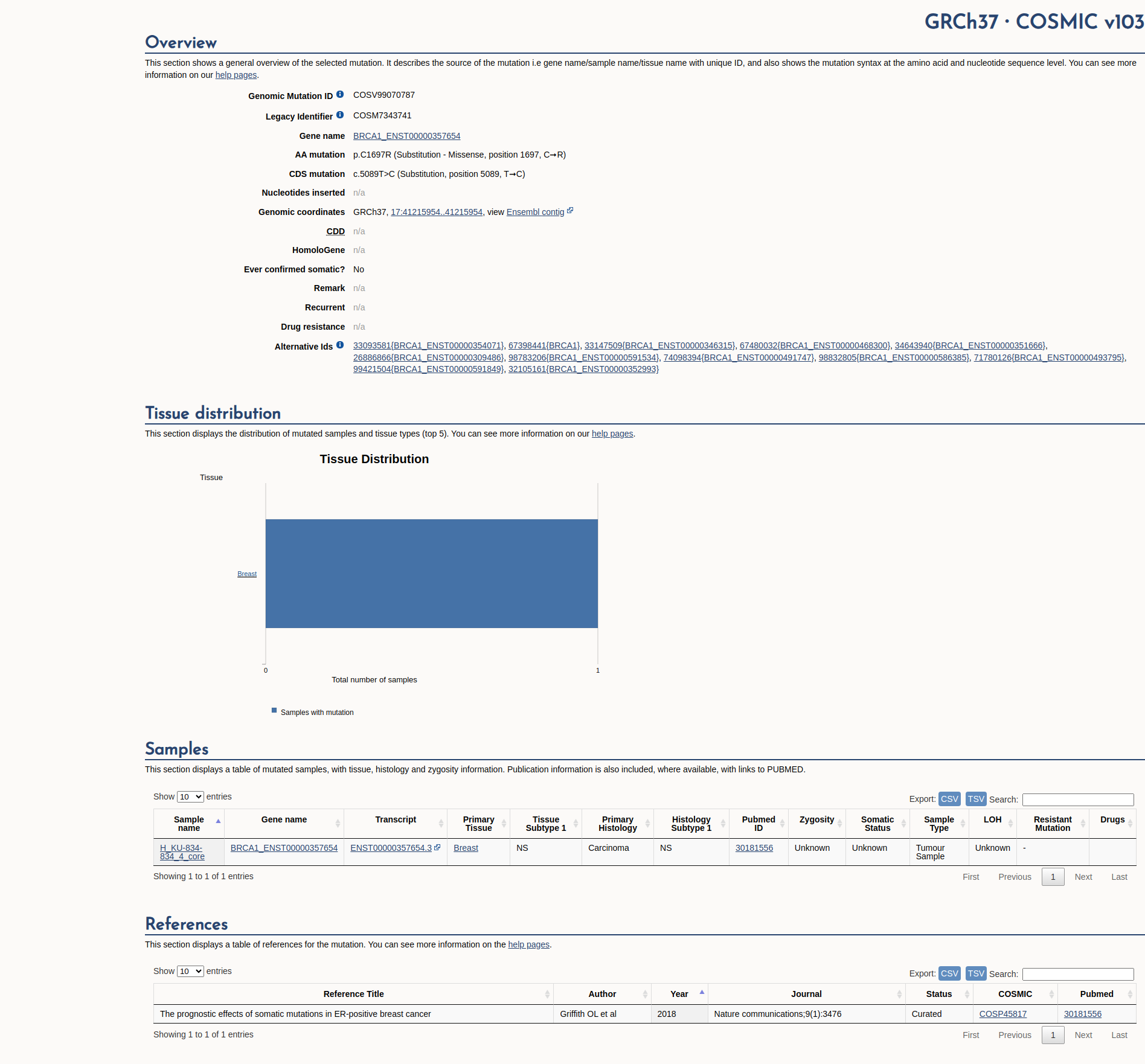
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: BRCA1C1697RBRCA1C1697RSomaticNCBI Gene:672|Show additional gene information Variant OverviewBRCA1, a tumor suppressor involved in the DNA damage response, is mutated in various cancer types.The BRCA1 C1697R mutation is likely oncogenic.Hide mutation effect description The BRCA1 C1697R mutation is located in the BRCT domain. In a study of 117 distinct BRCA1 missense variants identified in patients with a known family history of breast or ovarian cancer, the C1697R variant was demonstrated to have severe folding defect, uncertain binding activity, uncertain binding specificity and compromised transcriptional activity (PMID: 20516115). In a separate study of 8 BRCA1 variants identified in Scandinavian breast and ovarian cancer families, C1697R was shown to have loss of transcriptional activity compared to wild type BRCA1 (PMID: 11157798). Another study of 94 BRCA1 variants showed that C1697R had no transcriptional activation of BRCA1 and was cancer-associated (PMID: 15172985). JAX-CKB: No results found
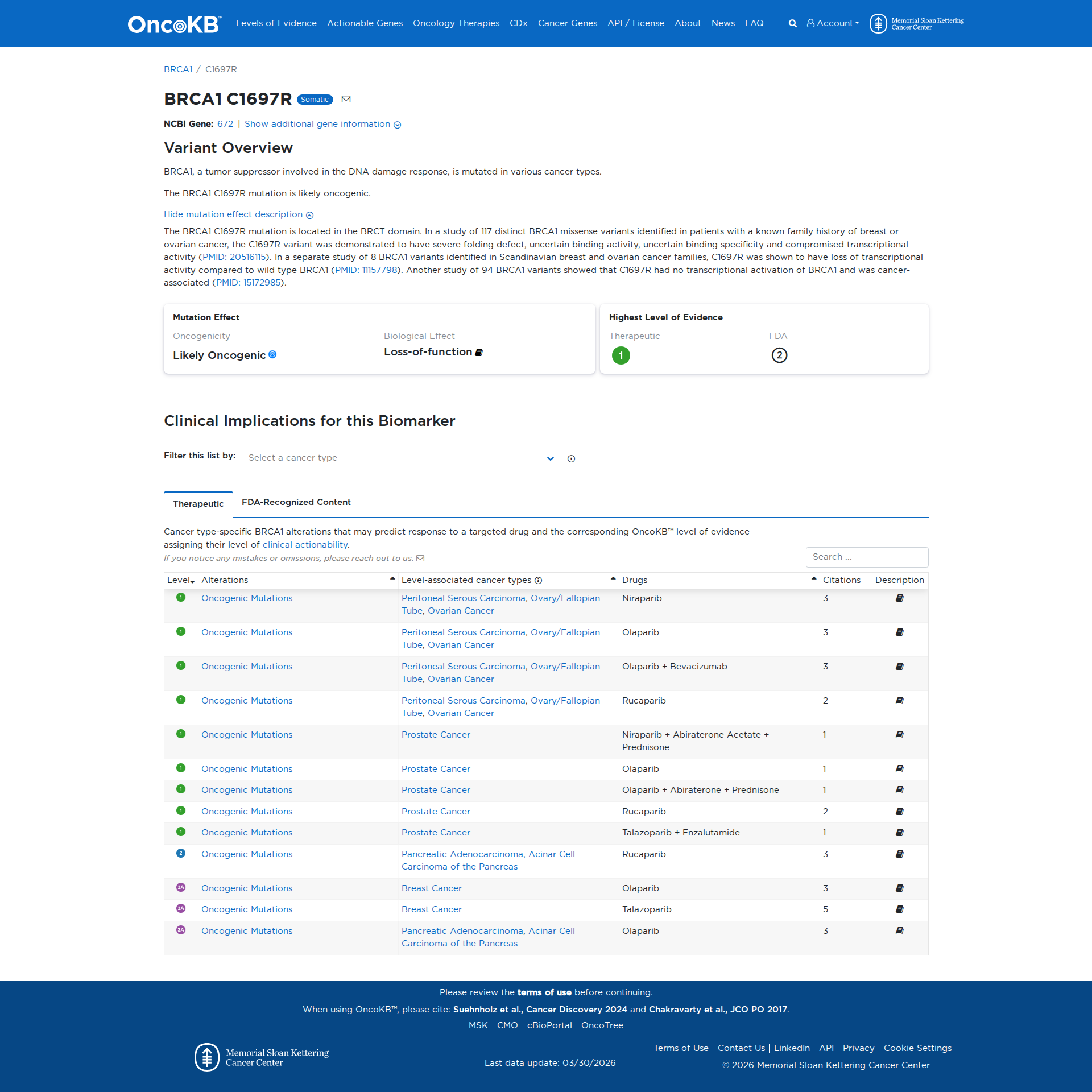

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.01 | -28 bp |
| Donor Loss (DL) | 0.0 | 14 bp |
| Acceptor Gain (AG) | 0.08 | 14 bp |
| Donor Gain (DG) | 0.05 | -63 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)