Genetic Information
Gene & Transcript Details
Gene
TP53
Transcript
NM_000546.5
MANE Select
Total Exons
—
Reference Sequence
NC_000017.10
Alternative Transcripts
| ID | Status | Details |
|---|---|---|
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
HGVS Notation
NM_000546.5:c.105G>T
Protein Change
L35F
Location
Exon 4
(Exon 4 of )
4
5'Exon Structure3'
Functional Consequence
Loss of Function
Alternate Identifiers
—
Clinical & Population Data
Population Frequency
gnomAD Global Frequency
0.0 in 100,000
Extremely Rare
ACMG Criteria Applied
PM2
This variant is absent or extremely rare in population databases (PM2 criteria applies).
ClinVar
OpenClassification
Uncertain Significance (VUS)
0 publications
Clinical Statement
"This variant has been reported in ClinVar as Uncertain Significance by ClinGen TP53 Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
OpenCOSMIC ID
COSM46160
Recurrence
2 occurrences
PM1 Criteria
Not Applied
COSMIC Database Preview
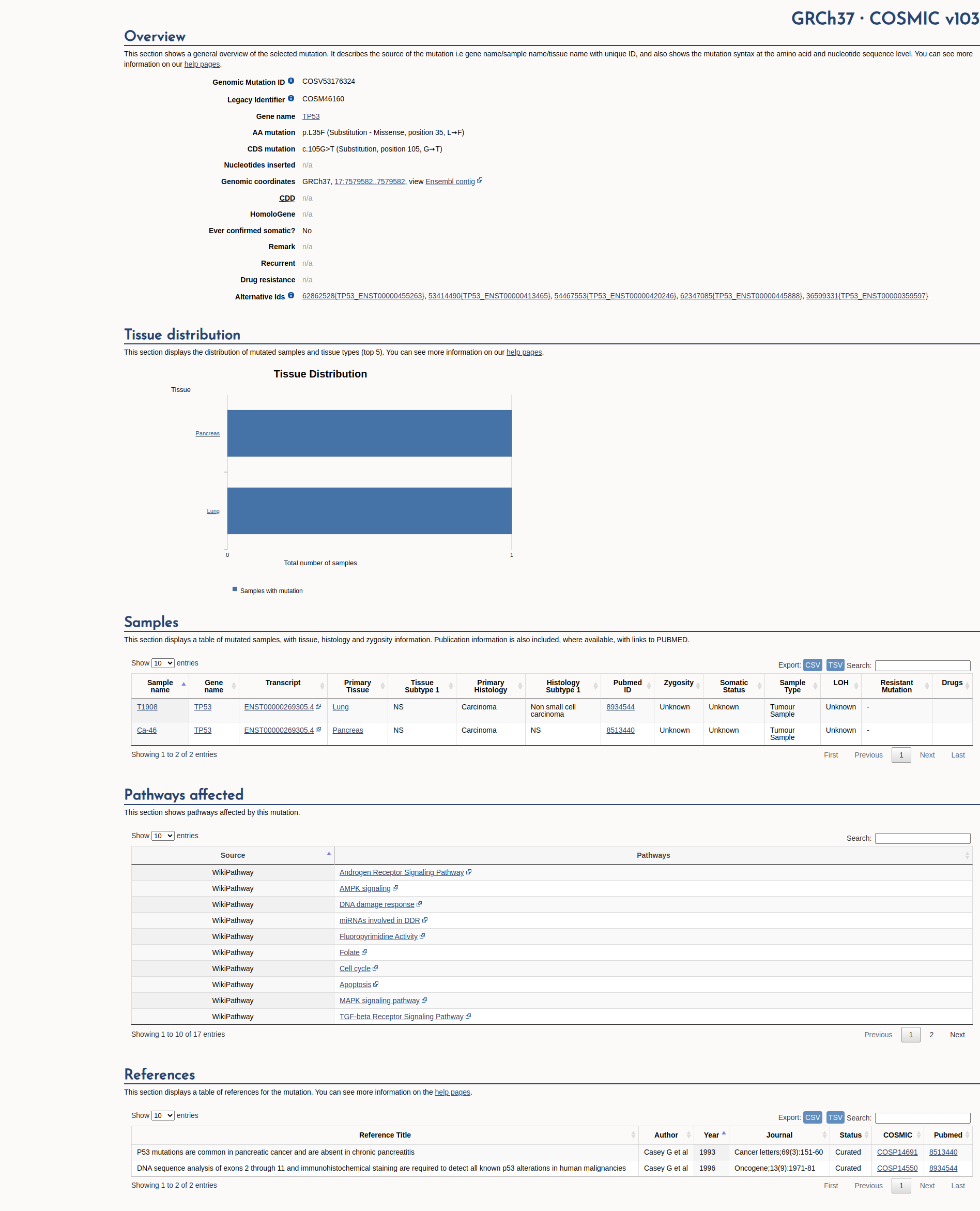
Accessing full COSMIC database details requires institutional login or subscription.
Functional Impact & Domains
Functional Domain
Hotspot Status
Not a hotspot
Domain Summary
This variant is not located in a mutational hotspot or critical domain.
Related Variants in This Domain
No evidence of other pathogenic variants at this position in gene TP53.
Functional Summary
Error in OpenAI Consolidation. OncoKB: TP53L35FTP53L35FSomaticNCBI Gene:7157|Show additional gene information Variant OverviewTP53, a tumor suppressor in the DNA damage pathway, is the most frequently mutated gene in cancer.The TP53 L35F mutation has not specifically been reviewed by the OncoKB team, and therefore its biological significance is unknown. JAX-CKB: No results found
Database Previews
OncoKB
JAX-CKB
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAIPredictor Consensus
Mixed/VUS
PP3 Applied
No
REVEL Score
0.0
Threshold: ≥0.75 = PP3 applied
SpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.01 | 355 bp |
| Donor Loss (DL) | 0.05 | -70 bp |
| Acceptor Gain (AG) | 0.35 | -11 bp |
| Donor Gain (DG) | 0.0 | 106 bp |
High impact (≥0.5)
Medium impact (0.2-0.49)
Low impact (<0.2)
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
Filter Criteria:
PM2
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)