Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_002524.4 | Alternative | 4454 nt | 255–824 |
| NM_002524.5 | MANE Select | 4326 nt | 132–701 |
| NM_002524.3 | Alternative | 4461 nt | 255–824 |
| NM_002524.2 | Alternative | 1963 nt | 254–823 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenVariant summary: NRAS c.173C>T (p.Thr58Ile) results in a non-conservative amino acid change located in the Small GTP-binding protein domain (IPR005225) of the encoded protein sequence. Five of five in-silico tools predict a damaging effect of the variant on protein function. The variant was absent in 251312 control chromosomes (gnomAD). c.173C>T has been reported in the literature in at least one individual affected with Noonan Syndrome who inherited the variant from an unaffected father with mosaicism (Altmuller_2017). However, these data do not allow any conclusion about variant significance. When assayed experimentally in HEK293 cells, the variant showed constitutively enhanced ERK/AKT activation, confirming it was a gain-of-function variant (Altmuller_2017). Five ClinVar submitters have assessed the variant since 2014: one classified the variant as likely pathogenic, and four as pathogenic. Based on the evidence outlined above, the variant was classified as likely pathogenic.
This sequence change replaces threonine, which is neutral and polar, with isoleucine, which is neutral and non-polar, at codon 58 of the NRAS protein (p.Thr58Ile). This variant is not present in population databases (gnomAD no frequency). This missense change has been observed in individual(s) with Noonan syndrome (PMID: 28594414). In at least one individual the variant was observed to be de novo. ClinVar contains an entry for this variant (Variation ID: 1070042). Invitae Evidence Modeling of protein sequence and biophysical properties (such as structural, functional, and spatial information, amino acid conservation, physicochemical variation, residue mobility, and thermodynamic stability) indicates that this missense variant is expected to disrupt NRAS protein function with a positive predictive value of 95%. Experimental studies have shown that this missense change affects NRAS function (PMID: 28594414). For these reasons, this variant has been classified as Pathogenic.
"This variant has been reported in ClinVar as Pathogenic (3 clinical laboratories) and as Likely pathogenic (2 clinical laboratories) and as Pathogenic by ClinGen RASopathy Variant Curation Expert Panel expert panel."
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: NRAST58INRAST58ISomaticNCBI Gene:4893|Show additional gene information Variant OverviewNRAS, a GTPase, is mutated in a diverse range of cancers, most frequently in melanoma and thyroid cancer.The NRAS T58I mutation is likely oncogenic.Hide mutation effect description The NRAS T58I mutation is located in the switch II region of the catalytic domain of the protein. This mutation has been found in melanoma. In vitro studies using a Novellus FACT assay with Ba/F3 cells expressing NRAS T58I demonstrate that the mutation is activating as measured by increased MAPK signaling compared to wildtype (PMID: 34117033). A patient with melanoma who was treated with the RAF inhibitors vemurafenib and dabrafenib acquired the NRAS Q61R and T58I mutations and developed drug resistance (PMID: 24265153). JAX-CKB: NRAS T58I lies within the GTP binding pocket of the Nras protein (UniProt.org). T58I has not been characterized, but can be predicted to confer a loss of function to Nras based on the effects of KRAS T58I, which results in defective intrinsic GTPase activity and decreased response to GTPase-activating proteins, leading to increased phosphorylation of of Mek and Akt, and transformation in cell culture (PMID: 16474405).
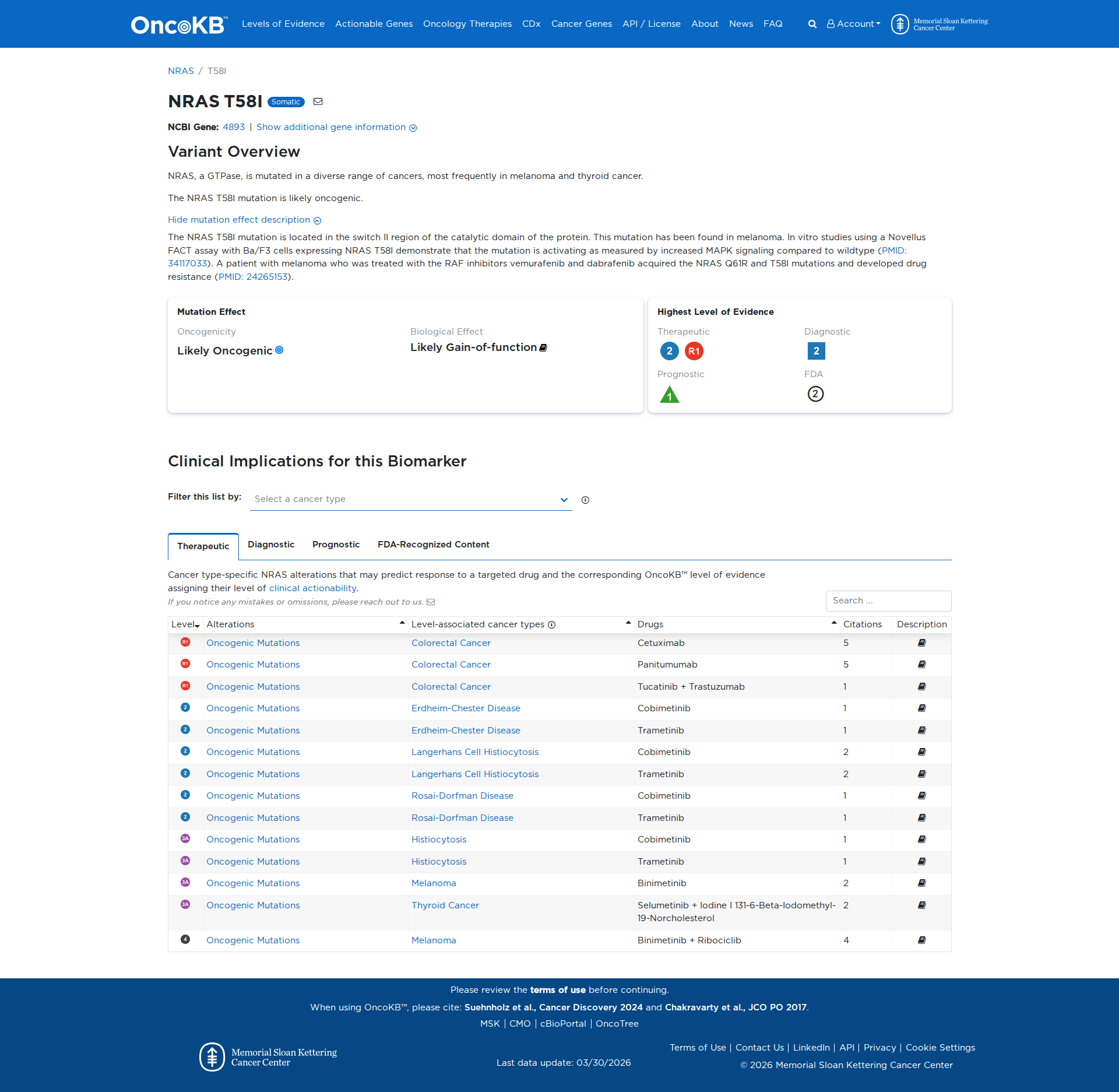

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.01 | 50 bp |
| Donor Loss (DL) | 0.0 | -117 bp |
| Acceptor Gain (AG) | 0.0 | 281 bp |
| Donor Gain (DG) | 0.0 | 268 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)