Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000059.4 | MANE Select | 11954 nt | 200–10456 |
| NM_000059.2 | Alternative | 11386 nt | 228–10484 |
| NM_000059.3 | RefSeq Select | 11386 nt | 228–10484 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis sequence change replaces aspartic acid, which is acidic and polar, with alanine, which is neutral and non-polar, at codon 2723 of the BRCA2 protein (p.Asp2723Ala). This variant is present in population databases (rs41293513, gnomAD 0.0009%). This missense change has been observed in individual(s) with breast cancer and/or ovarian cancer (PMID: 12601471, 28888541). ClinVar contains an entry for this variant (Variation ID: 52516). Invitae Evidence Modeling incorporating data from in vitro experimental studies (PMID: 33609447) indicates that this missense variant is expected to disrupt BRCA2 function with a positive predictive value of 95%. Experimental studies have shown that this missense change affects BRCA2 function (PMID: 18451181, 24323938, 29394989, 29884841). This variant disrupts the p.Asp2723 amino acid residue in BRCA2. Other variant(s) that disrupt this residue have been determined to be pathogenic (PMID: 15290653, 15695382, 16489001, 17924331, 18607349, 21990134). This suggests that this residue is clinically significant, and that variants that disrupt this residue are likely to be disease-causing. In summary, the currently available evidence indicates that the variant is pathogenic, but additional data are needed to prove that conclusively. Therefore, this variant has been classified as Likely Pathogenic.
Data used in classification: The variant was observed in 3 independent UK families undergoing clinical diagnostic BRCA1/BRCA2 testing for HBOC according to diagnostic criteria, the denominator of which dataset of clinical testing was 16,600. Case control comparison against ethnically matched population data (3/16,600 in familial cases against 1/55,659 gnomAD NFE controls) 2-sided Fishers exact: pexact= 0.037 (PS4_strong). The frequency of this variant is 0/67,264 individuals (remainder of the GNOMAD population) (PM2_mod). This variant is predicted deleterious on AlignGVGD (class: C65), SIFT (Deleterious), Polyphen2 HumVar (probably damaging) and CADD (18.82 (PP3_sup). This variant is at same codon as an established pathogenic variant on ClinVar (c.8168A>G). (PM5_mod). The variant is in the DNA-binding domain of BRCA2 (PM1_sup). In the BRCA2 Homology-Directed Repair Activity assay DNA Binding Domain assay (Guidugli et al Cancer Res 2013;73:265-275,Couch Lab), the variant has a probability of pathogenicity of P=1.0. In the VarCall Bayesian statistical model for VUS classification using functional assay data (Guidugli et al Am J Hum Genet 2018; 102:233-248, Couch Lab), the variant has a probability of being deleterious of 0.994 and an overall classification of pathogenic. (PS3_strong). This variant is classified on ClinVar as likely pathogenic by multiple accredited USA diagnostic laboratories (Ambry Genetics 2015, Counsyl 2018 and GeneDx 2017) (PP5_sup). Data not used in classification: There are additional reports of this variant in DMuDB (11), BIC (2), and BRCA2 LOVD (4).
The p.Asp2723Ala variant in BRCA2 has been reported in at least 4 individuals with breast caner (Scott 2003, Gorringe 2008, BIC database). It has also been identified in 1/113396 European chromosomes by gnomAD (http://gnomad.broadinstitute.org); however, this frequency is low enough to be consistent with the frequency of hereditary breast and ovarian cancer (HBOC) in the general population. This variant has also been reported in Clinvar (Variation ID: 52516). Computational prediction tools and conservation analyseis suggest that this variant may impact the protein, though this information is not predictive enough to determine pathogenicity. In vitro assays provide some evidence that this variant impact protein function (Guidugli 2018); however, these types of assays may not accurately represent biological function. Another variant involving this codon (p.Asp2723His) has been identified in individuals with breast cancer and has been classified as Pathogenic by the ClinGen-approved ENIGMA expert panel. In summary, although additional studies are required to fully establish its clinical significance, this variant meets criteria to be classified as likely pathogenic for autosomal dominant hereditary breast and ovarian cancer syndrome (HBOC). ACMG/AMP Criteria applied: PM2, PS3_Moderate, PM5, PP3, PS4_Supporting.
The c.8168A>C variant in BRCA2 is a missense variant predicted to cause substitution of Aspartic acid by Alanine at amino acid 2723 (p.Asp2723Ala). This variant is present in gnomAD v2.1 (exomes only, non-cancer subset) or gnomAD v3.1 (non-cancer subset) but is below the ENIGMA BRCA1/2 VCEP threshold >0.00002 for BS1_Supporting (PM2_Supporting, BS1, and BA1 are not met). This BRCA2 missense variant is within a key functional domain and the computational predictor BayesDel (noAF) gives a score of 0.576, above the recommended threshold of 0.30 for prediction of impact on BRCA2 function via protein change. SpliceAI predictor score of 0.00 suggests that the variant has no impact on splicing (score threshold <0.10) (PP3 met). Reported by two calibrated studies to exhibit protein function similar to pathogenic control variants (PMIDs: 33609447, 33293522) (PS3 met). Multifactorial likelihood ratio analysis using clinically calibrated data produced a combined LR for this variant of 0.557 (based on Pathology LR=0.214; Co-occurrence LR=1.102; Family History LR=2.36), which is above the ENIGMA BRCA1/2 VCEP threshold for BP5 (>0.48) and below PP4 (<2.08) (BP5 and PP4 not met; 31853058, Internal lab contributors). Cosegregation analysis of family(ies) carrying this variant provided evidence towards pathogenicity, and has a Bayes Score of 74321.3, above the thresholds for Very strong pathogenic evidence (LR >350) (PP1_Very strong; Internal lab contributors). In summary, this variant meets the criteria to be classified as a Pathogenic variant variant for BRCA2-related cancer predisposition based on the ACMG/AMP criteria applied as specified by the ENIGMA BRCA1/2 VCEP (PP3, PS3, PP1_Very strong).
The p.D2723A pathogenic mutation (also known as c.8168A>C), located in coding exon 17 of the BRCA2 gene, results from an A to C substitution at nucleotide position 8168. The aspartic acid at codon 2723 is replaced by alanine, an amino acid with dissimilar properties. This alteration is non-functional in multiple protein functional assays including homology-directed DNA repair (HDR) assay (Guidugli L et al, Cancer Res. 2013 Jan; 73(1):265-75; Farrugia DJ et al., Cancer Res. 2008 May; 68(9):3523-31; Guidugli L et al. Am. J. Hum. Genet., 2018 Jan.). This variant occurs in a structural hotspot region of the protein and is predicted to destabilize the protein and disrupt the native protein-protein (BRCA2-DSS1) interaction (Ambry internal data; Yang et al. Science. 2002; 297(5588):1837-48). This amino acid position is highly conserved in available vertebrate species. In addition, this alteration is predicted to be deleterious by in silico analysis. Based on the majority of available evidence to date, this alteration is interpreted as a disease-causing mutation.
"This variant has been reported in ClinVar as Likely pathogenic (4 clinical laboratories) and as Uncertain significance (1 clinical laboratories) and as Likely Pathogenic (1 clinical laboratories) and as Pathogenic (1 clinical laboratories) and as Pathogenic by ClinGen ENIGMA BRCA1 and BRCA2 Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: BRCA2D2723ABRCA2D2723ASomaticNCBI Gene:675|Show additional gene information Variant OverviewBRCA2, a tumor suppressor involved in the DNA damage response, is mutated in various cancer types.The BRCA2 D2723A mutation is likely oncogenic.Hide mutation effect description The BRCA2 D2723A mutation is located in the C-terminal domain of the protein. In a study of 22 BRCA2 missense mutations identified in multiple breast cancer families using functional assays measuring the ability of mutated BRCA2 to repair DNA damage by homologous recombination and to control centriole amplification, BRCA2 D2723A was found to be inactivating (PMID: 18451181). Expression of this mutation in a BRCA-deficient cell line demonstrated that it is likely inactivating as measured by decreased homologous recombination (HR) DNA-repair activity in an in vitro homology-directed DNA repair (HDR) assay compared to wildtype BRCA2 and BRCA2-negative controls (PMID: 29394989). In vitro studies have demonstrated that this mutation is inactivating as measured by loss of HDR in an in vitro HDR assay (PMID: 35736817). Germline BRCA2 D2723A mutations are considered likely pathogenic by the ACMG framework classification (PMID: 35736817). JAX-CKB: No results found
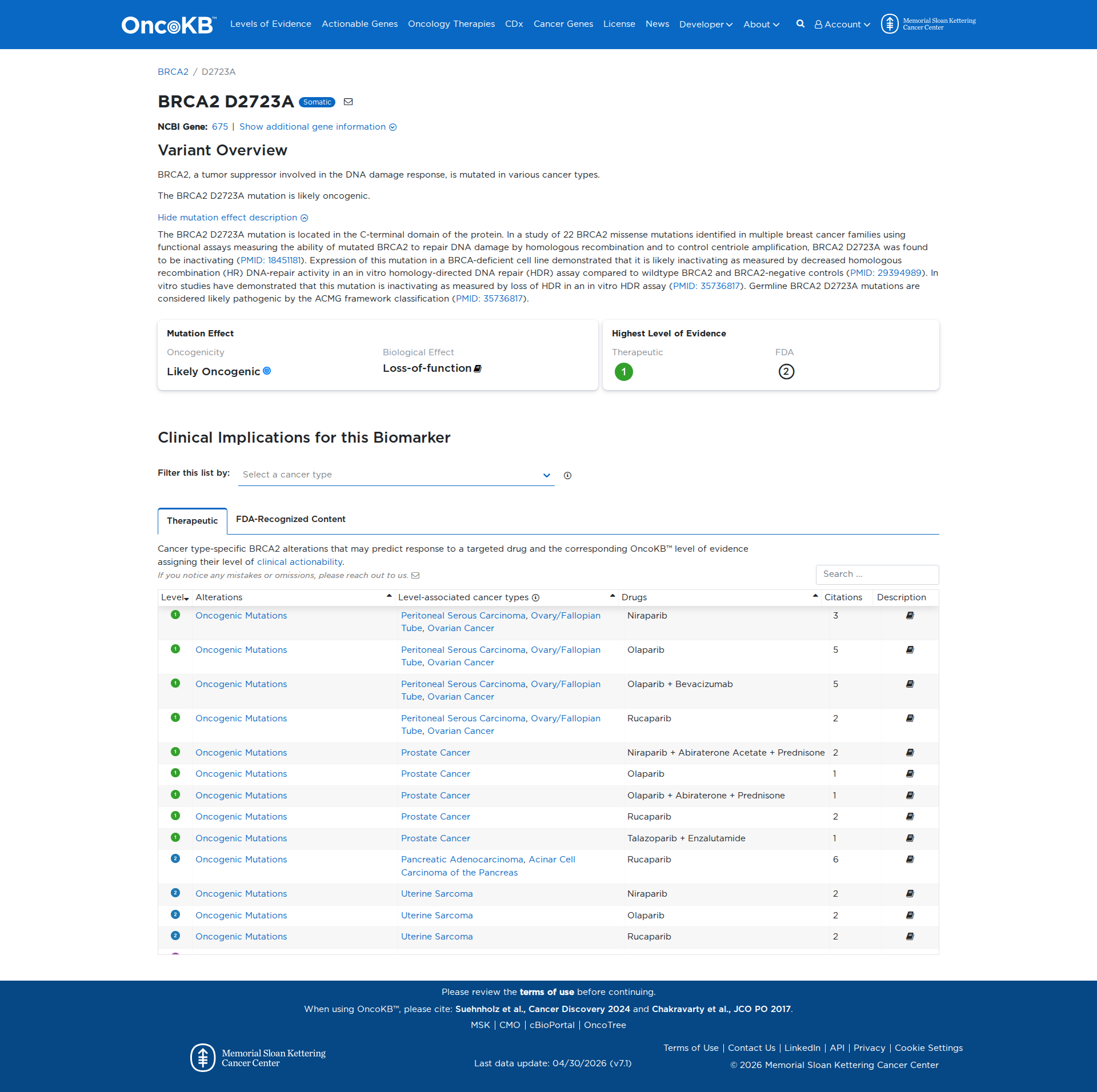

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 0 bp |
| Donor Loss (DL) | 0.0 | 6 bp |
| Acceptor Gain (AG) | 0.0 | 45 bp |
| Donor Gain (DG) | 0.0 | 163 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)