Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_002524.4 | Alternative | 4454 nt | 255–824 |
| NM_002524.5 | MANE Select | 4326 nt | 132–701 |
| NM_002524.3 | Alternative | 4461 nt | 255–824 |
| NM_002524.2 | Alternative | 1963 nt | 254–823 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis sequence change replaces alanine, which is neutral and non-polar, with threonine, which is neutral and polar, at codon 11 of the NRAS protein (p.Ala11Thr). This variant is present in population databases (no rsID available, gnomAD 0.003%). This variant has not been reported in the literature in individuals affected with NRAS-related conditions. ClinVar contains an entry for this variant (Variation ID: 561350). An algorithm developed to predict the effect of missense changes on protein structure and function (PolyPhen-2) suggests that this variant is likely to be tolerated. Experimental studies have shown that this missense change does not substantially affect NRAS function (PMID: 34117033). In summary, the available evidence is currently insufficient to determine the role of this variant in disease. Therefore, it has been classified as a Variant of Uncertain Significance.
"This variant has been reported in ClinVar as Likely benign (1 clinical laboratories) and as Uncertain significance (2 clinical laboratories) and as Uncertain Significance by ClinGen RASopathy Variant Curation Expert Panel expert panel."
COSMIC Somatic Evidence
Open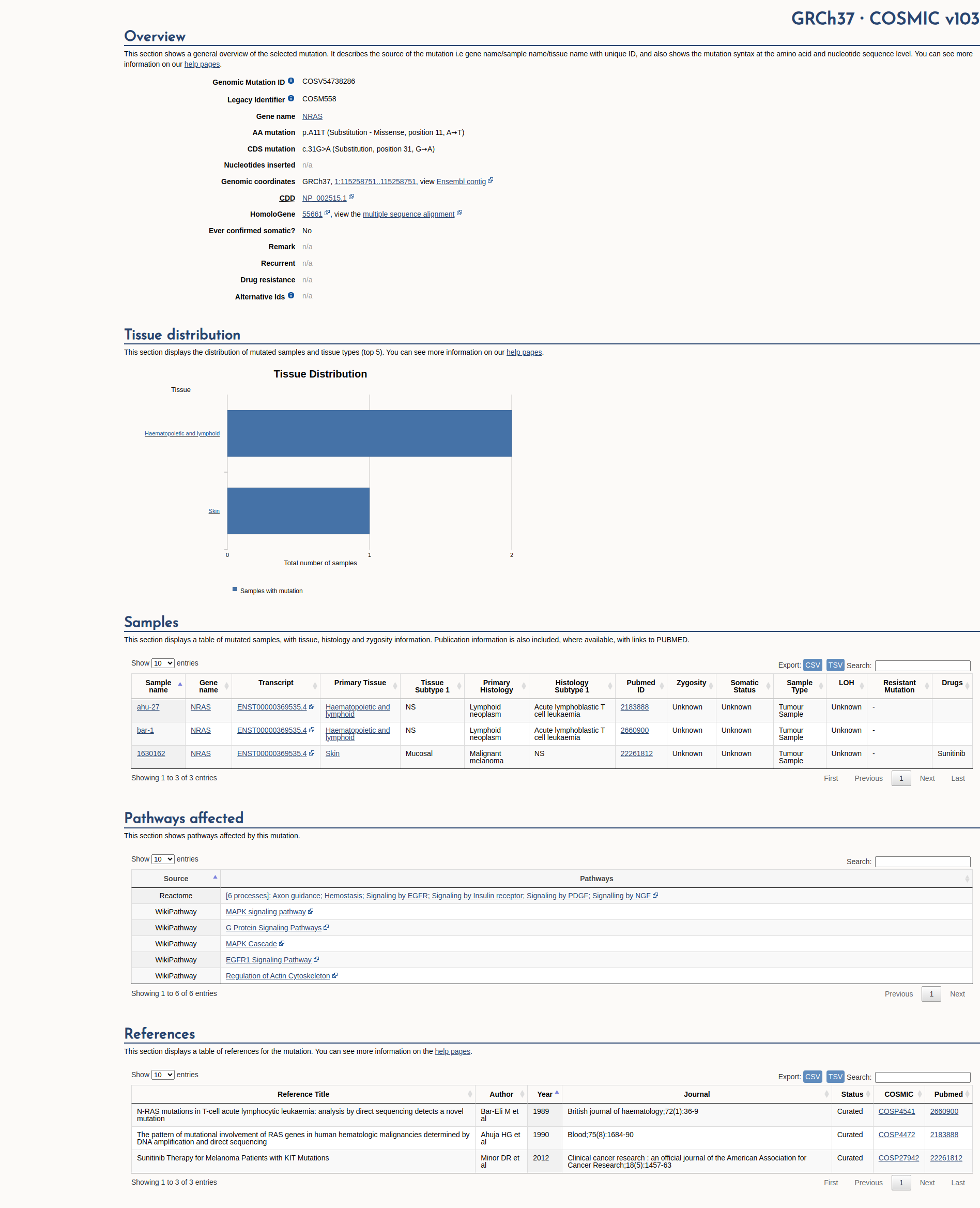
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: NRASA11TNRASA11TSomaticNCBI Gene:4893|Show additional gene information Variant OverviewNRAS, a GTPase, is mutated in a diverse range of cancers, most frequently in melanoma and thyroid cancer.The NRAS A11T mutation is likely neutral.Hide mutation effect description The NRAS A11T mutation is located in the catalytic GTP-binding domain of the protein. This mutation has been identified in acute lymphoblastic T-cell leukemia (PMID: 2660900). In vitro studies using a Novellus FACT assay with Ba/F3 cells expressing NRAS A11T demonstrate that the mutation is likely neutral as measured by MAPK signaling activity comparable to wildtype (PMID: 34117033). JAX-CKB: NRAS A11T lies within a GTP-binding region of the Nras protein (UniProt.org). A11T has been identified in the scientific literature (PMID: 2183888), but has not been biochemically characterized and therefore, its effect on Nras protein function is unknown (PubMed, Nov 2025).

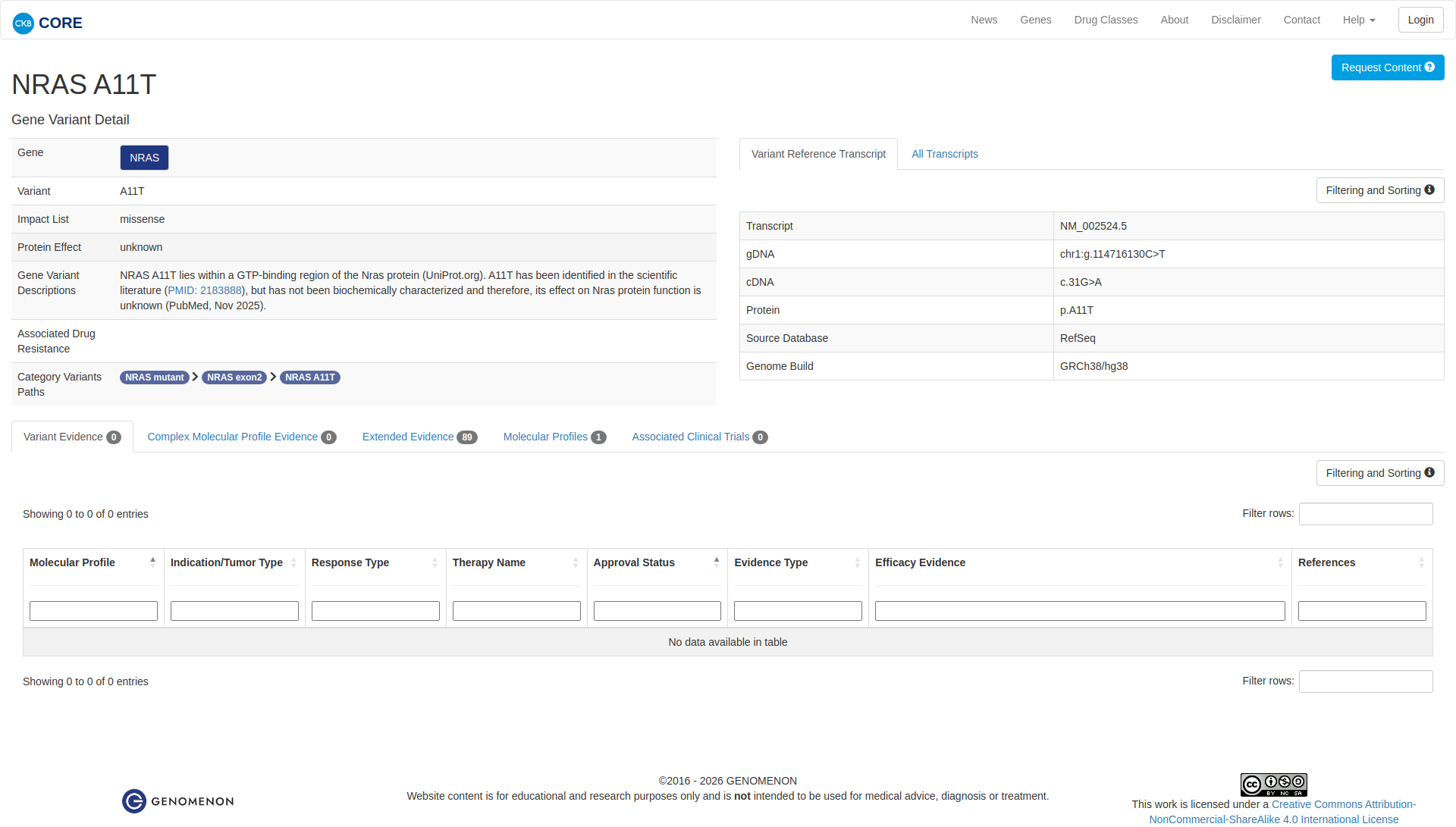
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 47 bp |
| Donor Loss (DL) | 0.0 | 143 bp |
| Acceptor Gain (AG) | 0.0 | -4 bp |
| Donor Gain (DG) | 0.0 | -3 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)