Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_024675.4 | MANE Select | 4008 nt | 154–3714 |
| NM_024675.3 | RefSeq Select | 4069 nt | 201–3761 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenPM5_supporting, PS4, PVS1
This variant is predicted to result in loss of function through nonsense-mediated decay of the encoded transcript or premature truncation of the encoded protein in a gene in which loss of function is a known mechanism of disease (ACMG/AMP: PVS1). Well-established functional studies have demonstrated this variant to have a damaging effect on protein function or splicing (ACMG/AMP: PS3; PMIDs:21285249, 23448497, 31757951). This variant has been reported at an elevated frequency in affected individuals/in multiple affected individuals in the literature (ACMG/AMP: PS4; PMIDs:17200668, 21182766, 21285249, 23471749, 27595995). This variant has been shown to segregate with disease in multiple affected family members (ACMG/AMP: PP1; PMIDs:17200668, 23471749).
This sequence change creates a premature translational stop signal (p.Trp1038*) in the PALB2 gene. RNA analysis indicates that this premature translational stop signal induces altered splicing and may result in an absent or altered protein product. This variant is present in population databases (rs180177132, gnomAD 0.01%). This premature translational stop signal has been observed in individual(s) with breast cancer (PMID: 17200668, 21182766, 21285249, 23471749). It has also been observed to segregate with disease in related individuals. ClinVar contains an entry for this variant (Variation ID: 126711). Studies have shown that this premature translational stop signal results in activation of a cryptic splice site, and produces a non-functional protein and/or introduces a premature termination codon (internal data). For these reasons, this variant has been classified as Pathogenic.
"This variant has been reported in ClinVar as Pathogenic (31 clinical laboratories) and as pathogenic (1 clinical laboratories) and as Pathogenic by ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: PALB2W1038*PALB2W1038*SomaticNCBI Gene:79728|Show additional gene information Variant OverviewPALB2, a scaffolding protein involved in DNA repair, is altered in various cancers.The PALB2 W1038* is a truncating mutation in a tumor suppressor gene, and therefore is likely oncogenic.Hide mutation effect description The mutation effect description for truncating mutations in PALB2 is: PALB2 truncating mutations can form several forms of C-terminally truncated PALB2 protein and are found in breast cancers. As such, PALB2 is considered a rare breast cancer susceptibility gene. In patient studies, PALB2 truncation mutations were shown to significantly increase the risk of breast cancer, with similar risks for estrogen receptor-positive and -negative breast cancer. PALB2 truncation mutations have also been found in cases of hereditary male breast cancer (PMID: 28858227, 18053174, 25529982, 29484706, 28279176, 28779002). JAX-CKB: PALB2 W1038* results in a premature truncation of the Palb2 protein at amino acid 1038 of 1186 (UniProt.org). W1038* confers a loss of function to the Palb2 protein as demonstrated by decreased protein stability (PMID: 31757951), aberrant cytosolic accumulation, decreased Brca2 binding and Rad51 foci formation (PMID: 28158555), and impaired homologous recombination in cultured cells (PMID: 28158555, PMID: 31757951).
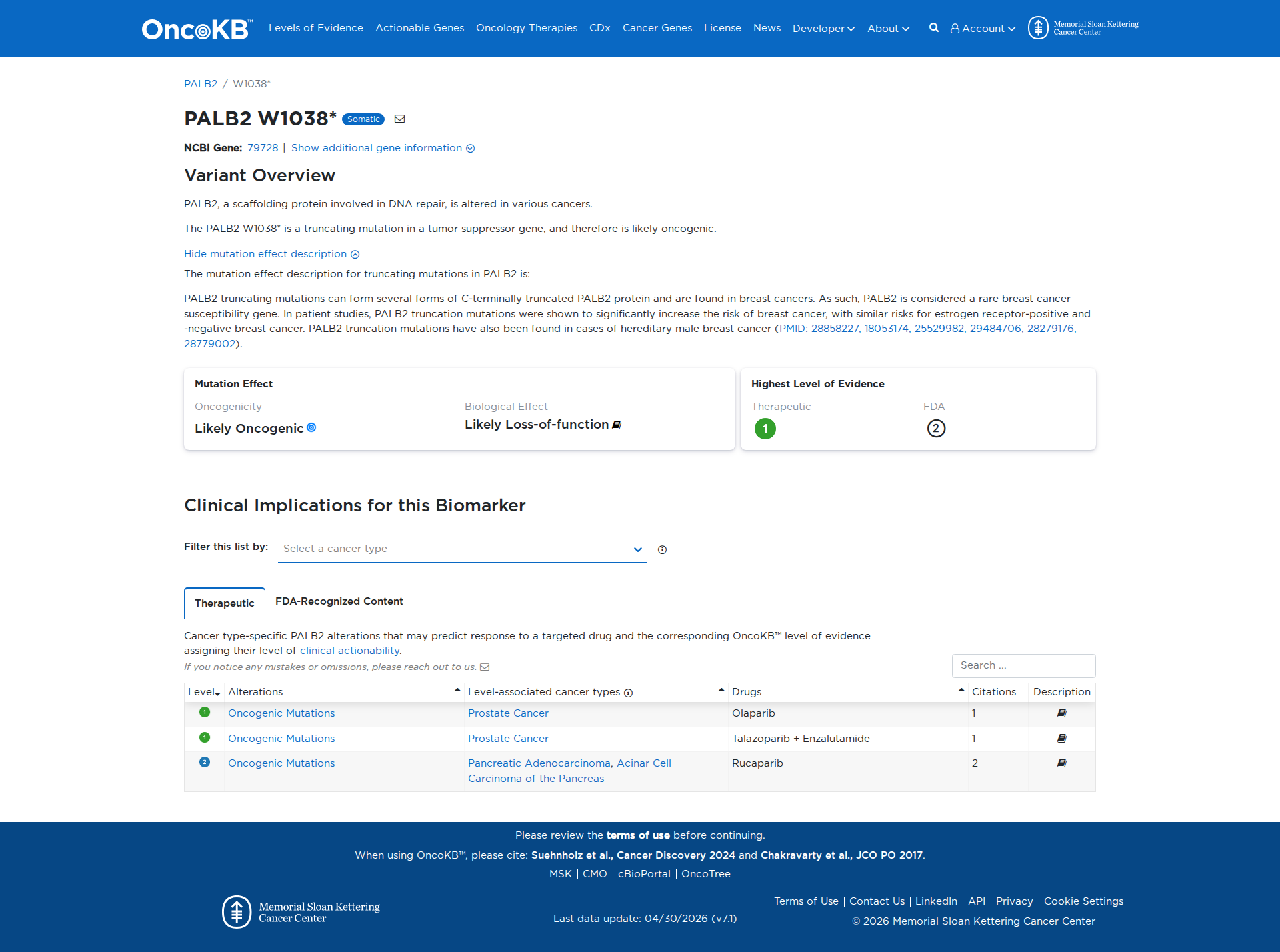

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.63 | 116 bp |
| Donor Loss (DL) | 0.74 | 0 bp |
| Acceptor Gain (AG) | 0.0 | 460 bp |
| Donor Gain (DG) | 0.17 | 31 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)