Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_012433.2 | Alternative | 4314 nt | 49–3963 |
| NM_012433.3 | RefSeq Select | 4338 nt | 95–4009 |
| NM_012433.4 | MANE Select | 6463 nt | 30–3944 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Open"Present in ClinVar, however no clinical evidence available for this variant."
COSMIC Somatic Evidence
Open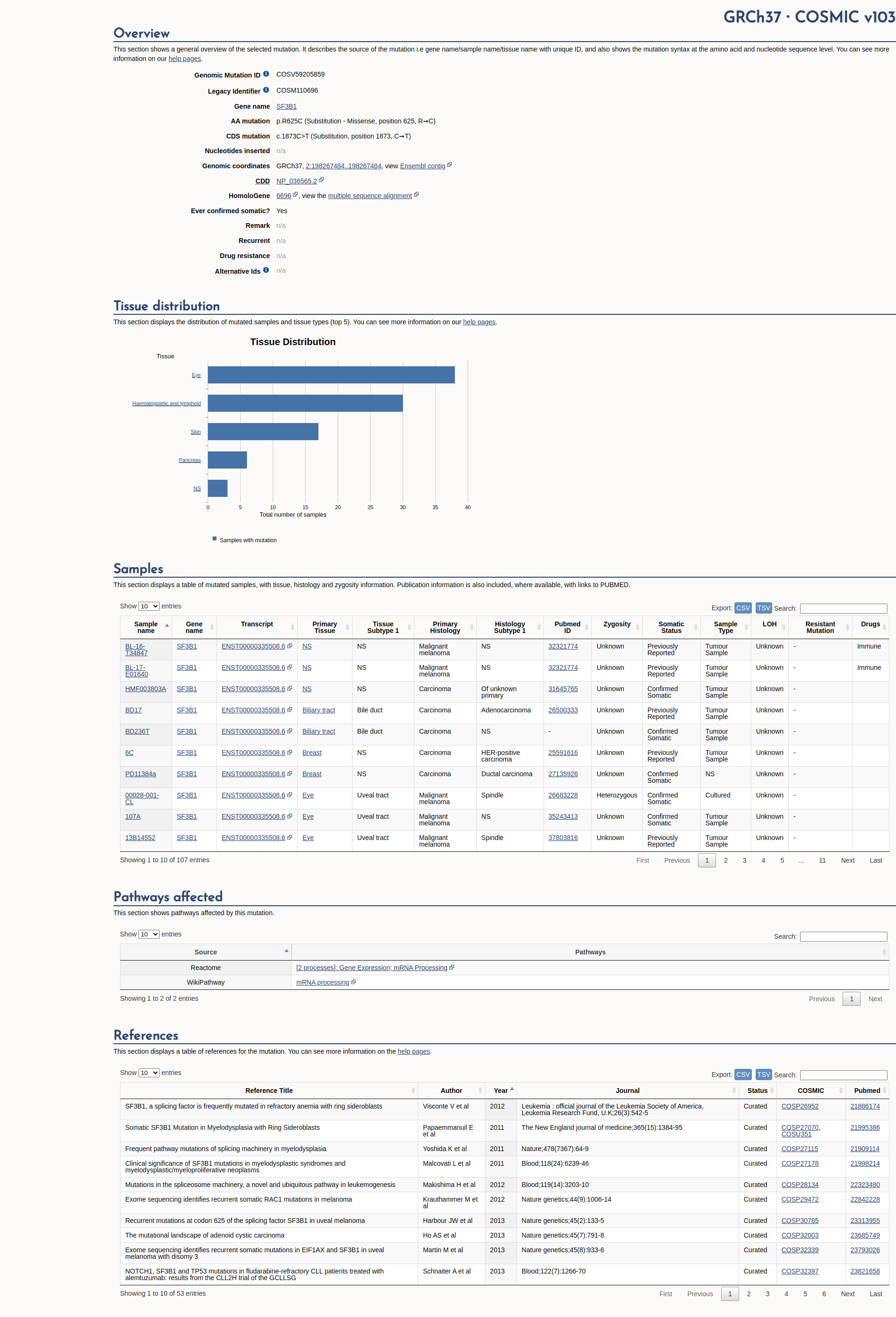
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: SF3B1R625CSF3B1R625CSomaticNCBI Gene:23451|Show additional gene information Variant OverviewSF3B1, a component of the spliceosome complex, is frequently mutated in hematologic malignancies.The SF3B1 R625C mutation is likely oncogenic.Hide mutation effect description The SF3B1 R625C mutation occurs in the protein’s HEAT domain, which functions as a scaffolding region for protein interactions. This mutation has been found in uveal melanoma (UVM) (PMID: 33031100). Mutations within this domain alter SF3B1-mediated splicing, resulting in the degradation of SF3B1-targeted transcripts and aberrant protein expression in the cell (PMID: 26565915). In patient-derived UVM and myelodysplastic syndrome samples, the R625C mutation has been found to alter the splicing and gene expression signatures (PMID: 23861464, 24434863). JAX-CKB: No results found


Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 66 bp |
| Donor Loss (DL) | 0.03 | -117 bp |
| Acceptor Gain (AG) | 0.01 | -247 bp |
| Donor Gain (DG) | 0.02 | 8 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)