Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000249.4 | MANE Select | 2494 nt | 31–2301 |
| NM_000249.3 | RefSeq Select | 2662 nt | 199–2469 |
| NM_000249.2 | Alternative | 2524 nt | 61–2331 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis submission and the accompanying classification are no longer maintained by the submitter. For more information on current observations and classification, please contact variantquestions@myriad.com.
Variant summary: MLH1 c.290A>G (p.Tyr97Cys) results in a non-conservative amino acid change located in the DNA mismatch repair protein family, N-terminal domain (IPR002099) of the encoded protein sequence. Five of five in-silico tools predict a damaging effect of the variant on protein function. The variant allele was found at a frequency of 0.00024 in 251380 control chromosomes, predominantly at a frequency of 0.0019 within the South Asian subpopulation in the gnomAD database. The observed variant frequency within South Asian control individuals in the gnomAD database is approximately 2.67 fold of the estimated maximal expected allele frequency for a pathogenic variant in MLH1 causing Hereditary Nonpolyposis Colorectal Cancer phenotype (0.00071), strongly suggesting that the variant is a benign polymorphism found primarily in populations of South Asian origin. c.290A>G has been reported in the literature in individuals affected with colon cancer, breast cancer, low-grade glioma and liver hepatocellular carcinoma (Bartley_2012, Huang_2019, Dorling_2021, Lerner-Ellis_2021) but it has also been reported in unaffected controls (Dorling_2021). The individual affected with colon cancer was reported with high MSI, but positive MLH1, MSH2, MSH6, and PMS2 staining via IHC. This patient also did not meet the Amsterdam II criteria, nor did they have a family history of Lynch-syndrome associated cancers (Bartley_2012). These reports do not provide unequivocal conclusions about association of the variant with Hereditary Nonpolyposis Colorectal Cancer. To our knowledge, no experimental evidence demonstrating an impact on protein function has been reported. The following publications have been ascertained in the context of this evaluation (PMID: 22086678, 29625052, 33471991, 32885271). Seven clinical diagnostic laboratories have submitted clinical-significance assessments for this variant to ClinVar after 2014 Multiple laboratories reported the variant with conflicting assessments: VUS (n=5), Benign (n=1) and likely Benign (n=1). Based on the evidence outlined above, the variant was classified as likely benign.
This alteration is classified as benign based on a combination of the following: seen in unaffected individuals, population frequency, intact protein function, lack of segregation with disease, co-occurrence, RNA analysis, in silico models, amino acid conservation, lack of disease association in case-control studies, and/or the mechanism of disease or impacted region is inconsistent with a known cause of pathogenicity.
This variant is considered benign. This variant has been observed at a population frequency that is significantly greater than expected given the associated disease prevalence and penetrance. This variant is strongly associated with less severe personal and family histories of cancer, typical for individuals without pathogenic variants in this gene [PMID: 27363726].
"This variant has been reported in ClinVar as Uncertain significance (4 clinical laboratories) and as Likely benign (3 clinical laboratories) and as Benign (2 clinical laboratories) and as likely benign (1 clinical laboratories)."
COSMIC Somatic Evidence
Open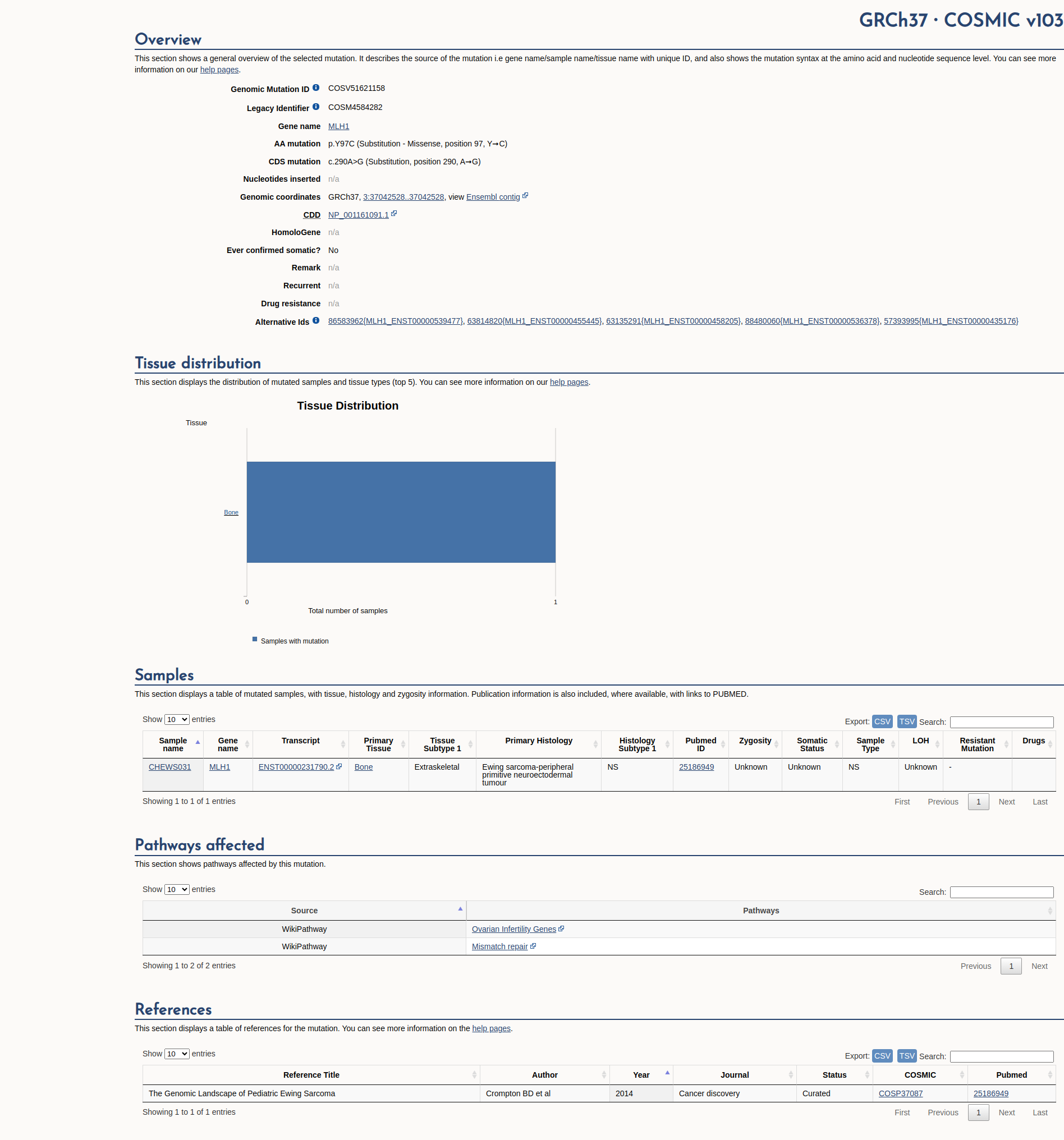
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: MLH1Y97CMLH1Y97CSomaticNCBI Gene:4292|Show additional gene information Variant OverviewMLH1, a DNA mismatch repair protein, is recurrently altered by deletion and mutation in various cancer types.The MLH1 Y97C mutation has not specifically been reviewed by the OncoKB team, and therefore its biological significance is unknown. JAX-CKB: No results found
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.01 | -82 bp |
| Donor Loss (DL) | 0.0 | 16 bp |
| Acceptor Gain (AG) | 0.0 | -78 bp |
| Donor Gain (DG) | 0.0 | 11 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)