Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_001354609.2 | Alternative | 9687 nt | 227–2530 |
| NM_001354609.1 | Alternative | 9702 nt | 226–2529 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenFor these reasons, this variant has been classified as Pathogenic. Experimental studies are conflicting or provide insufficient evidence to determine the effect of this variant on BRAF function (PMID: 16439621, 18413255, 19376813). Advanced modeling of protein sequence and biophysical properties (such as structural, functional, and spatial information, amino acid conservation, physicochemical variation, residue mobility, and thermodynamic stability) performed at Invitae indicates that this missense variant is expected to disrupt BRAF protein function. ClinVar contains an entry for this variant (Variation ID: 40387). This missense change has been observed in individual(s) with cardio-facio-cutaneous syndrome (PMID: 16439621, 25463315). In at least one individual the variant was observed to be de novo. This variant is not present in population databases (gnomAD no frequency). This sequence change replaces glycine, which is neutral and non-polar, with valine, which is neutral and non-polar, at codon 596 of the BRAF protein (p.Gly596Val).
Variant classified using ACMG guidelines
The Gly596Val variant has been identified in 7 individuals with clinical feature s of Cardio-facio-cutaneous syndrome (CFC) and 3 individuals with clinical featu res of Noonan syndrome (Rodriguez-Viciana 2006, Yoon 2007, Rodriguez-Viciana 200 8, LMM unpublished data). It was absent from large population studies. In-vitro functional studies provide some evidence that the Gly596Val variant may impact p rotein function (Rodrigues-Viciana 2006). Furthermore, animal models in zebrafis h have shown that this variant causes severe developmental abnormalities (Anasta saki 2009). In summary, this variant meets our criteria to be classified as path ogenic (http://personalizedmedicine.partners.org/Laboratory-For-Molecular-Medici ne/).
"This variant has been reported in ClinVar as Pathogenic (10 clinical laboratories) and as Likely pathogenic (1 clinical laboratories) and as Pathogenic by ClinGen RASopathy Variant Curation Expert Panel expert panel."
COSMIC Somatic Evidence
Open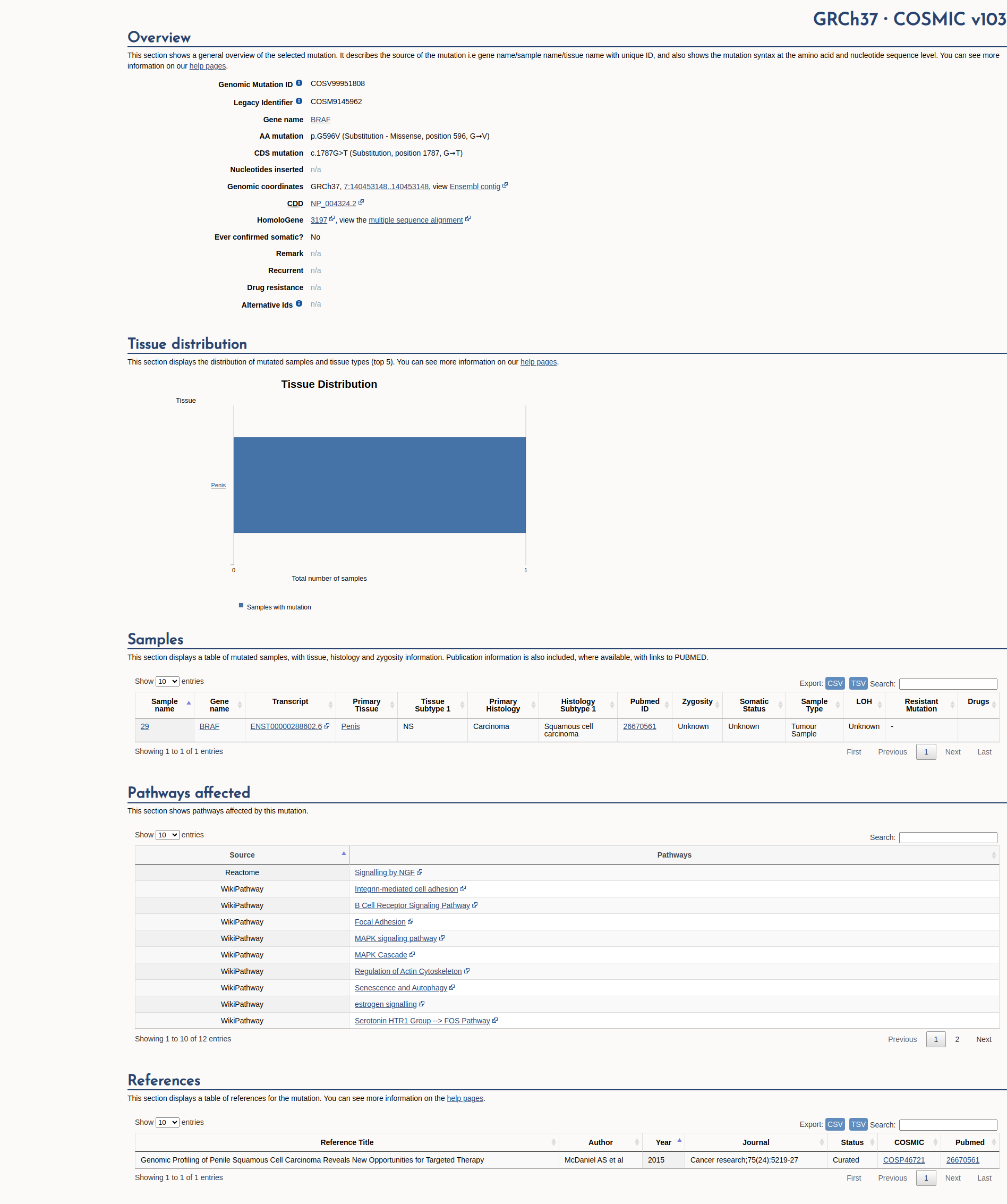
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: BRAFG596VBRAFG596VSomaticNCBI Gene:673|Show additional gene information Variant OverviewBRAF, an intracellular kinase, is frequently mutated in melanoma, thyroid and lung cancers among others.The BRAF G596V mutation is likely oncogenic.Hide mutation effect description The class III BRAF G596V mutation is located in the kinase domain in exon 15 of the protein (PMID: 33198372). This mutation has been found in cardio-facio-cutaneous syndrome (PMID: 16439621). Expression of this mutant in HEK293T cells demonstrated it has impaired kinase activity and is not able to induce downstream MEK and ERK activation in this context (PMID: 16439621). However, when expressed in zebrafish embryos, the G596V mutation causes the same aberrant elongation phenotype as BRAF V600E, suggesting that it might function in a manner similar to BRAF activating mutants during development in vivo (PMID: 19376813). In a Phase II trial, a patient with non-small cell lung cancer harboring BRAF G596V was treated with vemurafenib and demonstrated a partial response (PMID: 26200454). JAX-CKB: BRAF G596V lies within the protein kinase domain of the Braf protein (UniProt.org). G596V results in activation of Erk in zebrafish models (PMID: 19376813), but impaired Braf kinase activity and does not activate downstream Mek and Erk in cell culture (PMID: 16439621), and therefore, is predicted to lead to a loss of Braf protein function.
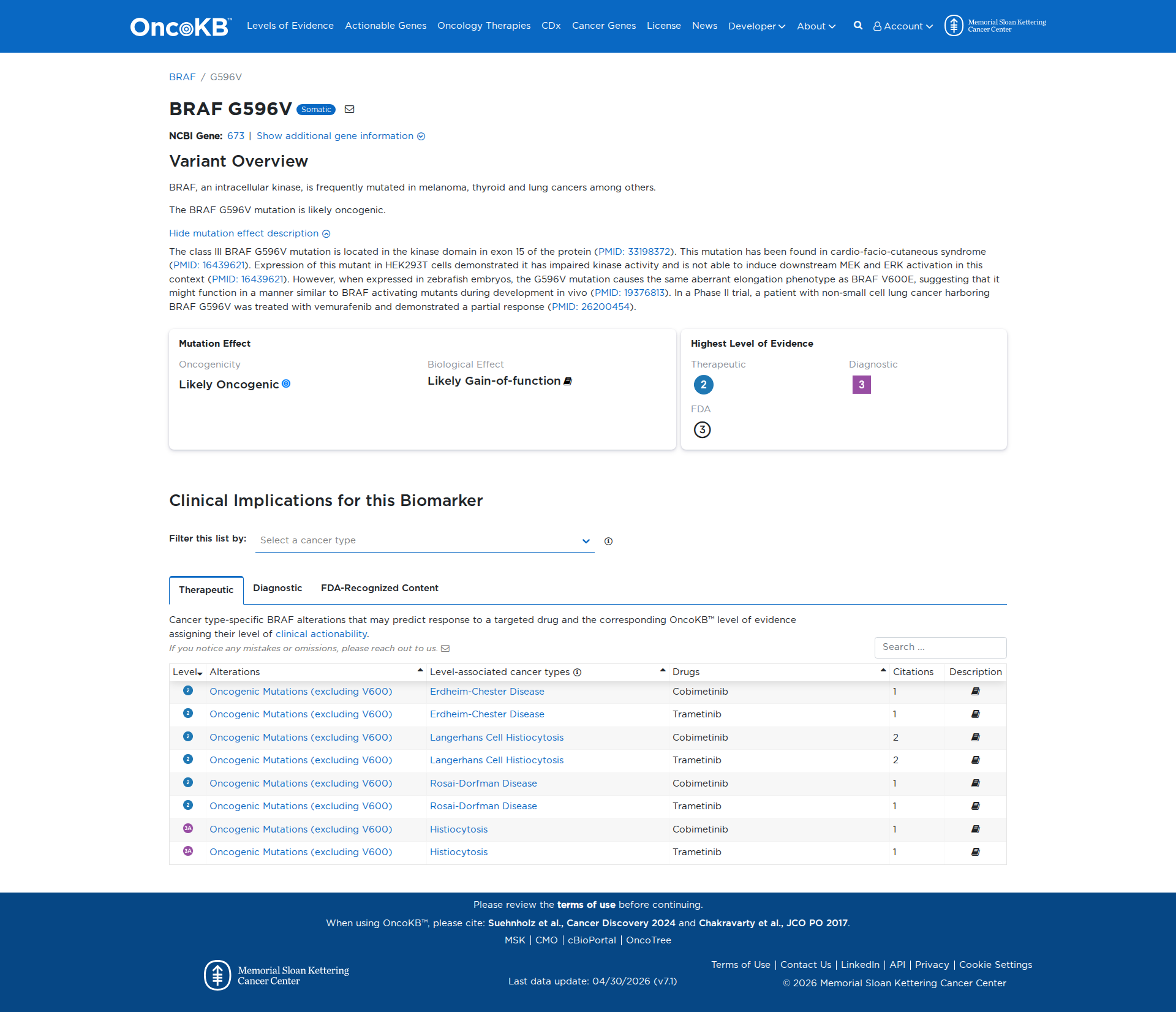

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.07 | 45 bp |
| Donor Loss (DL) | 0.0 | 45 bp |
| Acceptor Gain (AG) | 0.07 | -6 bp |
| Donor Gain (DG) | 0.04 | -73 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)