Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_001369786.1 | Alternative | 5417 nt | 178–747 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe c.15A>T variant in the KRAS gene is a missense variant predicted to cause substitution of lysine by asparagine at amino acid 5 (p.Lys5Asn). This variant is absent from gnomAD v2.1.1 (PM2_Supporting). The computational predictor REVEL gives a score of 0.764 supporting a deleterious impact to KRAS function (PP3). The variant is located in the KRAS gene, which has been defined by the ClinGen RASopathy Expert Panel as a gene with a low rate of benign missense variants and pathogenic missense variants are common (PP2). This variant has been reported in at least 5 individuals with clinical features of a RASopathy, in 2 confirmed de novo occurrences (PS4, PS2_VeryStrong; PMID: 17056636; Ambry, EGL Genetics, Invitae internal data, ClinVar SCV000742406.3, SCV000202928.7). In vitro functional studies showed that the p.Val14Ile variant enhanced MEK activation (PS3_Supporting; PMID: 20949621). In summary, this variant meets criteria to be classified as pathogenic for autosomal dominant RASopathies based on the ACMG/AMP criteria applied, as specified by the ClinGen RASopathy Variant Curation Expert Panel: PS2_VeryStrong, PS4, PS3_Supporting, PM2_Supporting, PP2, PP3 (Specification Version 2.3, 12/3/2024).
"This variant has been reported in ClinVar as Uncertain significance (1 clinical laboratories) and as Likely pathogenic (1 clinical laboratories) and as Pathogenic by ClinGen RASopathy Variant Curation Expert Panel expert panel."
COSMIC Somatic Evidence
Open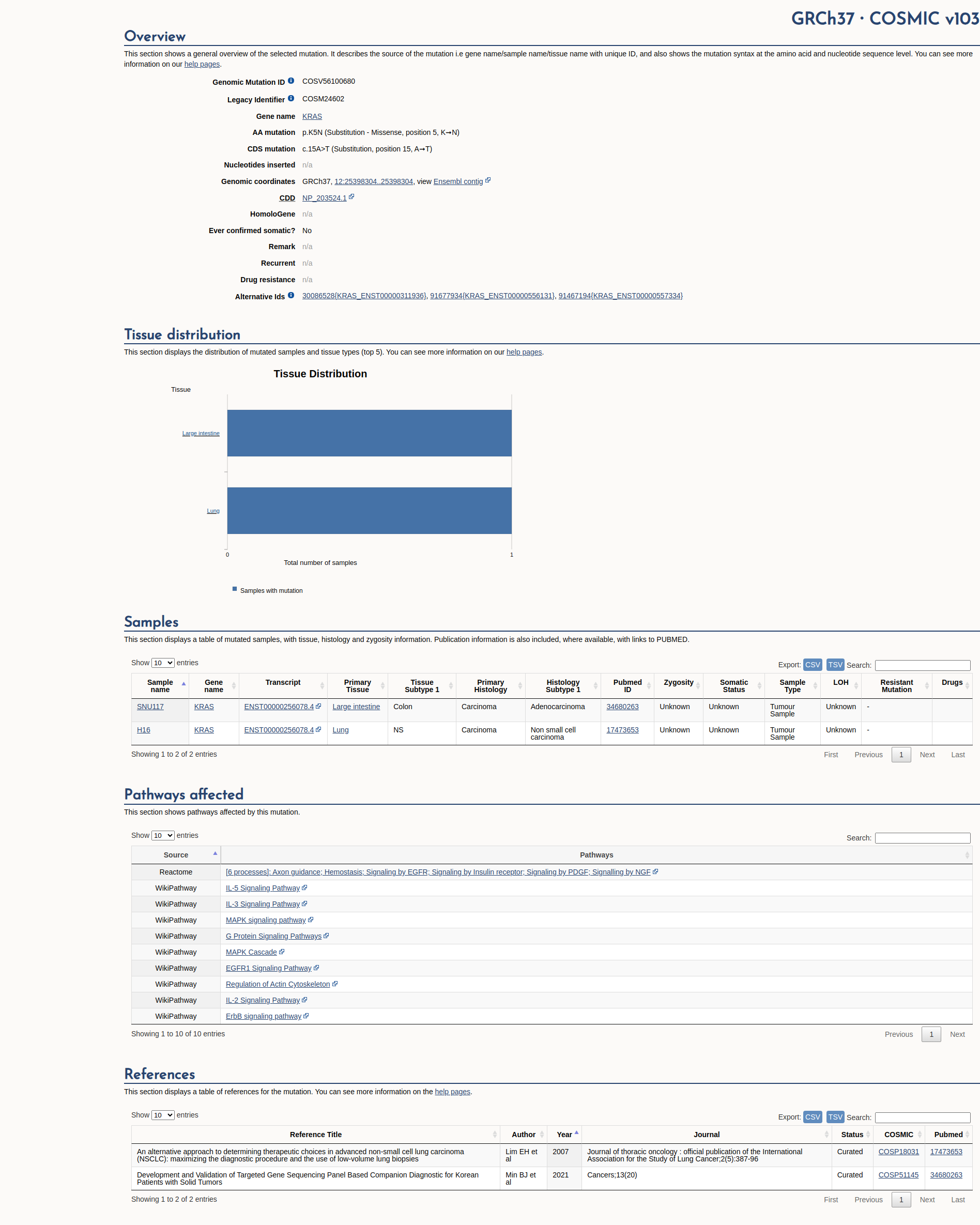
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: KRASK5NKRASK5NSomaticNCBI Gene:3845|Show additional gene information Variant OverviewKRAS, a GTPase which functions as an upstream regulator of the MAPK pathway, is frequently mutated in various cancer types including lung, colorectal and pancreatic cancers.The KRAS K5N mutation is likely oncogenic.Hide mutation effect description The KRAS K5N mutation is located in the catalytic G-domain of the protein. This mutation was found in the germline of a patient with Noonan syndrome (PMID: 17875937). Expression of this mutation in simian and human cell lines demonstrated that it is likely activating, as measured by a modest increase in pathway activation compared to wildtype (PMID: 20949621, 25941399). JAX-CKB: No results found
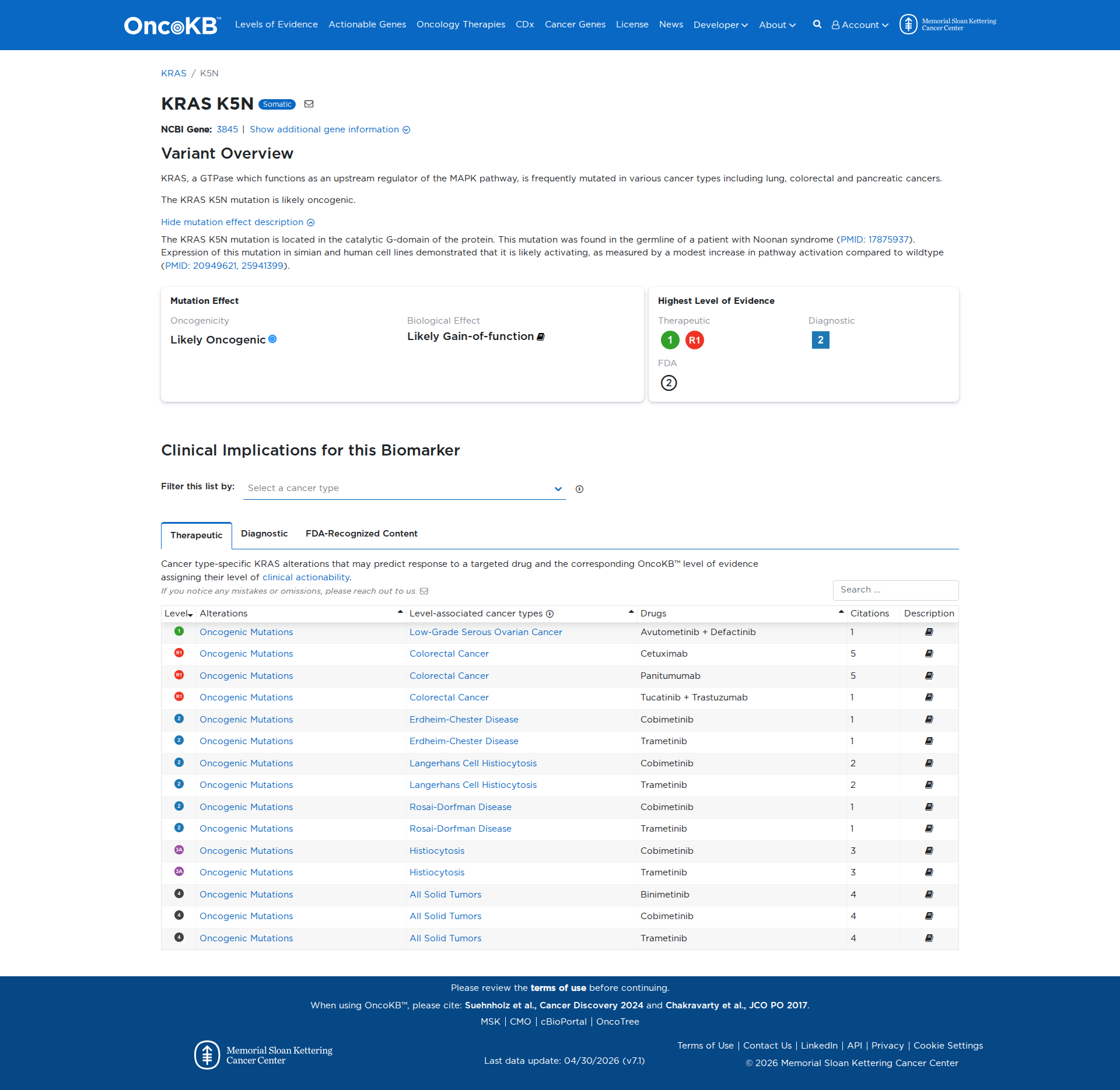
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | None bp |
| Donor Loss (DL) | 0.0 | None bp |
| Acceptor Gain (AG) | 0.0 | None bp |
| Donor Gain (DG) | 0.0 | None bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)