Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_001369786.1 | Alternative | 5417 nt | 178–747 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis sequence change replaces proline, which is neutral and non-polar, with leucine, which is neutral and non-polar, at codon 34 of the KRAS protein (p.Pro34Leu). This variant is not present in population databases (gnomAD no frequency). This missense change has been observed in individual(s) with Noonan syndrome (PMID: 17056636). In at least one individual the variant was observed to be de novo. ClinVar contains an entry for this variant (Variation ID: 40454). Invitae Evidence Modeling incorporating data from in vitro experimental studies (internal data) indicates that this missense variant is expected to disrupt KRAS function with a positive predictive value of 95%. Experimental studies have shown that this missense change affects KRAS function (PMID: 20949621). This variant disrupts the p.Pro34 amino acid residue in KRAS. Other variant(s) that disrupt this residue have been determined to be pathogenic (PMID: 16474405, 17056636). This suggests that this residue is clinically significant, and that variants that disrupt this residue are likely to be disease-causing. For these reasons, this variant has been classified as Pathogenic.
The Pro34Leu variant in KRAS has not been identified by our laboratory, but has been reported in one individual with Noonan syndrome where it was demonstrated t o have occurred de novo (Zenker 2007). Another variant at the same nucleotide p osition (c.101C>A, p.Pro34Gln) has also been reported in another patient with No onan syndrome where it too, was demonstrated to have occurred de novo (Zenker 20 07). Functional studies have shown that the Pro34Leu variant results in a GTPas e-Activating Protein (GAP)-resistant conformation and consequently, to a weak ga in-of-function, consistent with disease pathogenesis (Gremer 2010). This variant has not been reported in large population studies, consistent with a pathogenic role. Computational analyses (biochemical amino acid properties, conservation, AlignGVGD, PolyPhen2, and SIFT) suggest that the variant may impact the protein. In summary, this variant meets our criteria to be classified as pathogenic bas ed on the de novo occurrence and supportive functional studies (http://pcpgm.par tners.org/LMM).
The variant is not observed in the gnomAD v2.1.1 dataset. Missense changes are a common disease-causing mechanism. In silico tool predictions suggest damaging effect of the variant on gene or gene product (REVEL: 0.87; 3Cnet: 0.97). Same nucleotide change resulting in same amino acid change has been previously reported as pathogenic/likely pathogenic with strong evidence (ClinVar ID: VCV000040454). The variant has been previously reported as de novo in at least two similarly affected unrelated individuals (PMID: 17056636, 20949621). A different missense change at the same codon (p.Pro34Arg) has been reported as pathogenic/likely pathogenic with strong evidence (ClinVar ID: VCV000012590). Therefore, this variant is classified as Pathogenic according to the recommendation of ACMG/AMP guideline.
The c.101C>T (p.P34L) alteration is located in exon 2 (coding exon 1) of the KRAS gene. This alteration results from a C to T substitution at nucleotide position 101, causing the proline (P) at amino acid position 34 to be replaced by a leucine (L). This variant was not reported in population-based cohorts in the Genome Aggregation Database (gnomAD). This variant was reported in individual(s) with features consistent with KRAS-related RASopathy; in at least one individual, it was determined to be de novo (Zenker, 2007; Chen, 2019; Wang, 2023). Other variant(s) at the same codon, c.101C>A (p.P34Q), have been identified in individual(s) with features consistent with KRAS-related RASopathy (Zenker, 2007). This amino acid position is highly conserved in available vertebrate species. This missense alteration is located in a region that has a low rate of benign missense variation (Lek, 2016; Firth, 2009). This alteration is predicted to be deleterious by in silico analysis. Based on the available evidence, this alteration is classified as pathogenic.
"This variant has been reported in ClinVar as Pathogenic (7 clinical laboratories) and as Pathogenic by ClinGen RASopathy Variant Curation Expert Panel expert panel."
COSMIC Somatic Evidence
Open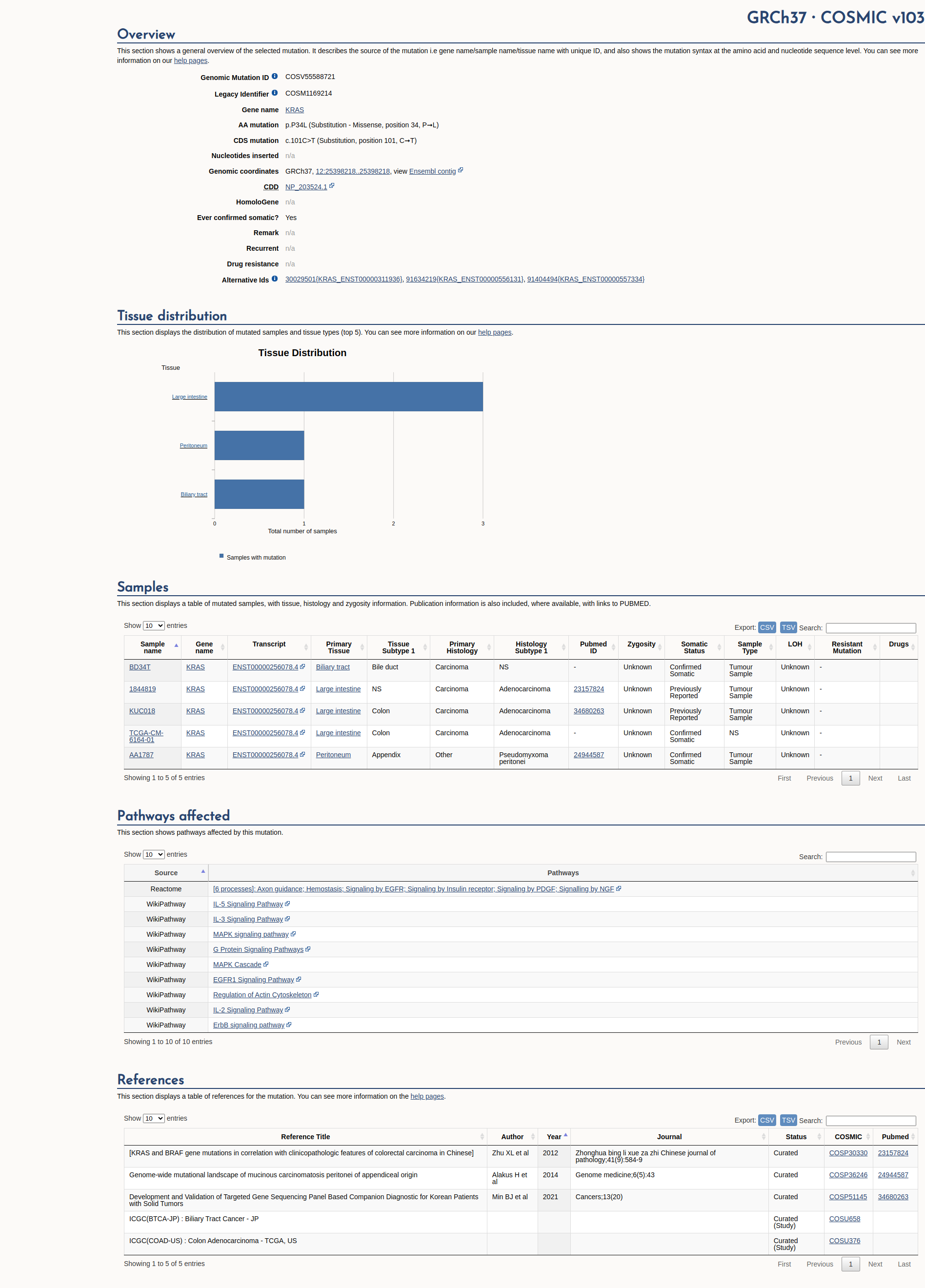
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: KRASP34LKRASP34LSomaticNCBI Gene:3845|Show additional gene information Variant OverviewKRAS, a GTPase which functions as an upstream regulator of the MAPK pathway, is frequently mutated in various cancer types including lung, colorectal and pancreatic cancers.The KRAS P34L mutation is known to be oncogenic.Hide mutation effect description The KRAS P34L mutation is located in the catalytic G-domain of the protein. This mutation was identified as a germline mutation in Noonan syndrome (PMID: 16474405). Expression of this mutation in vitro and in a simian cell line demonstrated that it is activating, as shown by decreased binding to regulatory proteins and increased pathway activation compared to wildtype (PMID: 20949621). JAX-CKB: No results found


Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | None bp |
| Donor Loss (DL) | 0.0 | None bp |
| Acceptor Gain (AG) | 0.0 | None bp |
| Donor Gain (DG) | 0.0 | None bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)